Validating Natural Product Biosynthesis: A Comprehensive LC-MS and Bioassay Framework for Drug Discovery
This article provides a comprehensive framework for researchers and drug development professionals on validating natural product biosynthesis.
Validating Natural Product Biosynthesis: A Comprehensive LC-MS and Bioassay Framework for Drug Discovery
Abstract
This article provides a comprehensive framework for researchers and drug development professionals on validating natural product biosynthesis. It explores the foundational role of natural products as privileged structures in drug discovery and details advanced LC-MS methodologies for their analysis. The content covers practical strategies for troubleshooting complex mixtures and optimizing biosynthetic pathways through synthetic biology. Furthermore, it establishes rigorous protocols for the orthogonal validation of both chemical identity and biological activity, integrating LC-MS with functional bioassays to bridge the gap from compound discovery to therapeutic application.
Natural Products as Privileged Structures: Foundations for Biosynthetic Validation
The Historical and Ongoing Role of Natural Products in Drug Discovery
Natural products (NPs) and their structural analogues have been a cornerstone of pharmacotherapy for centuries, making a major contribution to the treatment of diseases, particularly in the realms of cancer and infectious diseases [1]. These secondary metabolites, produced by terrestrial and marine plants, microorganisms, fungi, and other organisms, represent an immense reservoir of chemical diversity evolved for specific biological functions, including defense mechanisms and competition with other organisms [2] [1]. Historically, records of natural product use date back to 2600 B.C. from Mesopotamia, documenting oils from Cupressus sempervirens (Cypress) and Commiphora species (myrrh) for treating coughs, colds, and inflammation [2]. The Ebers Papyrus (2900 B.C.), an Egyptian pharmaceutical record, documents over 700 plant-based drugs, while Chinese Materia Medica (1100 B.C.) and the Tang Herbal (659 A.D.) provide extensive documentation of natural product uses [2].
Despite a decline in pursuit by the pharmaceutical industry from the 1990s onwards, recent technological and scientific developments—including improved analytical tools like liquid chromatography-mass spectrometry (LC-MS), genome mining, and advanced microbial culturing—are revitalizing interest in NP-based drug discovery [1]. Between 2000 and 2020, approximately 30 percent of newly introduced small molecule drugs were derived from natural products, underscoring their continued relevance [3]. This guide objectively compares the performance of natural products against other drug discovery approaches, providing supporting experimental data and detailing the methodologies that validate their biosynthesis and bioactivity.
Comparative Analysis of Natural Products in Drug Discovery
The following table summarizes key quantitative data comparing natural products with synthetic compounds and combinatorial chemistry libraries, highlighting the distinct advantages and challenges of each approach.
Table 1: Performance Comparison of Natural Products vs. Alternative Drug Discovery Approaches
| Parameter | Natural Products | Synthetic Compounds/Combinatorial Chemistry | Supporting Data and Evidence |
|---|---|---|---|
| Chemical Diversity & Structural Complexity | High scaffold diversity, structural complexity, higher molecular rigidity [1]. | Lower structural diversity, less complex, more planar structures [1]. | NPs have higher molecular mass, more sp³ carbon & oxygen atoms, greater H-bond acceptors/donors, and lower cLogP values [1]. |
| Clinical Success Rate | Higher translatability and progression through clinical trials [3]. | Lower historical success rate for new chemical entities. | About one-third of FDA-approved drugs over the past 20 years are based on NPs or their derivatives [4]. |
| Therapeutic Areas | Dominant in cancer and infectious diseases; also successful in cardiovascular, multiple sclerosis, and immunological disorders [1] [4]. | Broad, but less dominant in anti-infectives and anticancer. | Drugs like Artemisinin (malaria), Taxol (cancer), and Dimethyl fumarate (multiple sclerosis) are NP-derived [1] [4]. |
| Bioactivity & Target Engagement | "Bioactive" compounds covering wider chemical space; often identified by phenotypic assays [1]. | Typically identified via target-based high-throughput screening (HTS). | NP pools are enriched with bioactive compounds optimized by evolution for biological interactions [1]. |
| Major Challenges | Technical barriers to screening, isolation, characterization, optimization, and supply; intellectual property issues [1] [4]. | Limited chemical space; can struggle with complex targets like protein-protein interactions [1]. | Complexity of NP mixtures can complicate isolation; dereplication is essential to avoid rediscovery [1]. |
Modern Workflows for Validating Natural Product Biosynthesis and Bioactivity
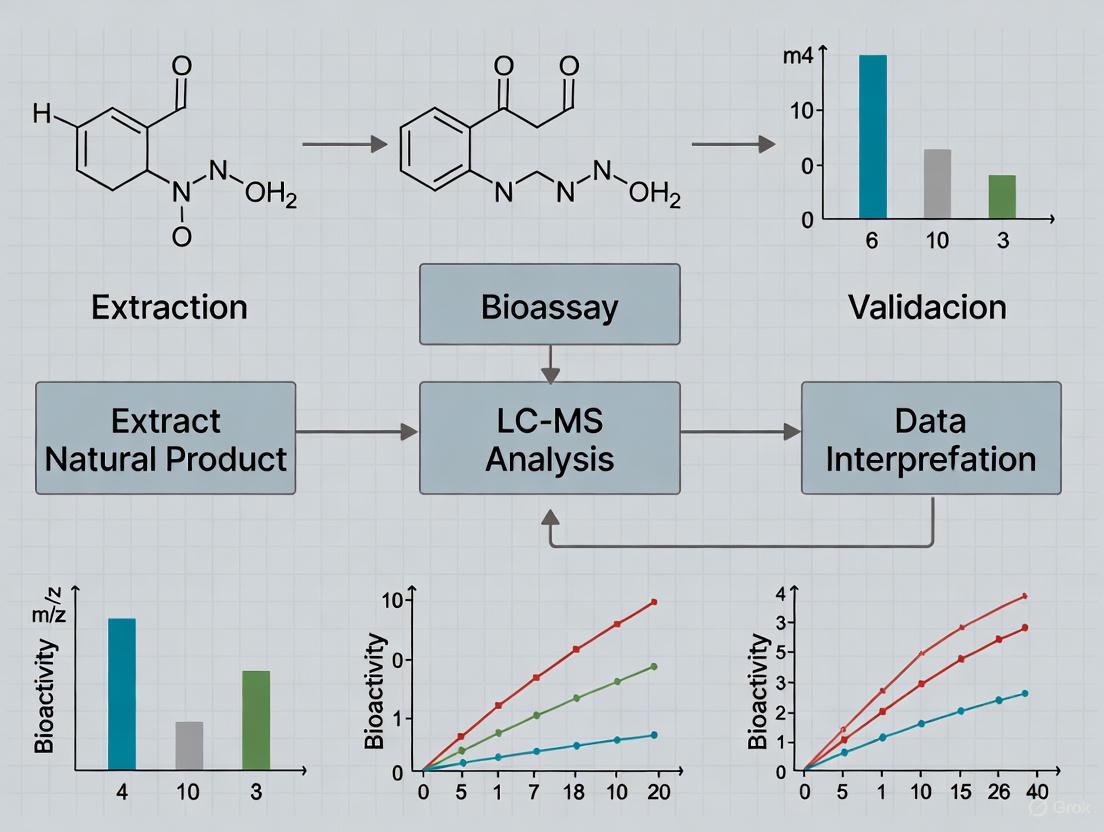
Modern drug discovery from natural products relies on an integrated workflow that couples advanced analytical chemistry with robust biological testing to identify and validate active compounds. The following diagram illustrates this multi-step process, from initial extraction to final compound identification.
Detailed Experimental Protocols for Key Workflow Steps
1. Protocol for LC-MS/MS Analysis of Short-Chain Fatty Acids (SCFAs) This protocol, adapted from a validated method for quantifying plasmatic SCFAs, highlights the role of LC-MS in analyzing NP-derived metabolites [5].
- Sample Preparation and Derivatization:
- Protein Precipitation: Add 300 µL of ice-cold acetonitrile to 100 µL of plasma. Vortex mix for 30 seconds and centrifuge at 14,000 × g for 10 minutes [5].
- Chemical Derivatization: Transfer the supernatant and add an internal standard (e.g., deuterated caproic acid). Derivatize with 3-nitrophenylhydrazine (3-NPH) and 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) in the presence of pyridine. Incubate at 40°C for 30 minutes [5].
- LC-MS/MS Analysis:
- Chromatography: Use a reversed-phase C18 column (e.g., Gemini NX-C18, 3.0 × 100 mm, 3 µm). Employ an isocratic mobile phase of 0.2% (v/v) formic acid and acetonitrile (70:30, v/v) at a flow rate of 0.5 mL/min. The total run time is 18 minutes [5].
- Mass Spectrometry: Operate the mass spectrometer in negative electrospray ionization (ESI-) mode. Use high-resolution detection (e.g., QTOF). Key source parameters: spray voltage = -4500 V, vaporizer temperature = 500°C. Monitor specific multiple reaction monitoring (MRM) transitions for derivatized SCFAs (e.g., acetic, propionic, butyric, caproic acids) [5].
2. Protocol for Bioactivity-Guided Fractionation Using TIMS-MS/MS This protocol leverages advanced instrumentation to deconvolute complex NP mixtures [3].
- High-Throughput Fractionation:
- Liquid Chromatography: Separate complex NP extracts using liquid chromatography (LC). Collect fractions automatically.
- Parallel Analysis: Split each LC fraction into two streams.
- Stream A (Chemical Profiling): Analyze by trapped ion mobility spectrometry tandem mass spectrometry (TIMS-MS/MS). The TIMS cell separates ions by size and shape, providing collisional cross section (CCS) values, an additional molecular descriptor. Use parallel accumulation-serial fragmentation (PASEF) to acquire clean, mobility-separated MS/MS spectra for thousands of molecules [3].
- Stream B (Bioactivity Screening): Test in various biological assays (e.g., antimicrobial, anticancer) to determine bioactivity.
- Data Integration and Prioritization: Correlate the chemical structures predicted from MS/MS fragmentation patterns (using machine learning models like MS2Mol) with the bioactivity data. Prioritize molecules that show both desirable bioactivity and drug-like structural properties for isolation [3].
Key Signaling Pathways Modulated by Natural Products
Natural products often exert their therapeutic effects by modulating key cellular defense and homeostasis pathways. The KEAP1/NRF2 pathway is a prime example, regulated by diverse NPs and relevant for conditions like multiple sclerosis, cancer, and neurodegenerative diseases [1]. The following diagram details this pathway and the natural products that target it.
Experimental Validation of Pathway Modulation
- Example: Validation of NRF2 Pathway Activation by Sulforaphane
- Assay: Use a cell line (e.g., HEK293) stably transfected with a luciferase reporter gene under the control of an Antioxidant Response Element (ARE) [1].
- Procedure: Treat cells with increasing concentrations of sulforaphane (isolated from Brassica oleracea) for 6-24 hours. Measure luciferase activity as a direct indicator of NRF2-mediated transcriptional activation.
- Downstream Analysis: Confirm pathway activation by quantifying the increased expression of downstream target proteins (e.g., NQO1, HO-1) using Western blotting or quantitative PCR. Sulforaphane is one of the most potent naturally occurring inducers of this pathway and has shown protective effects in animal models of multiple sclerosis and neurodegenerative diseases [1].
The Scientist's Toolkit: Essential Research Reagent Solutions
The following table details key reagents, materials, and instrumentation essential for conducting research in natural product drug discovery, particularly within the context of LC-MS and bioassay validation.
Table 2: Essential Research Reagents and Materials for NP Discovery
| Tool/Reagent | Function/Application | Example Use-Case in NP Research |
|---|---|---|
| High-Resolution Mass Spectrometer (HRMS) | Accurately determines the mass of molecules and their fragments; essential for structural elucidation [5] [1]. | Used in LC-HRMS and TIMS-MS/MS workflows to identify unknown NPs in complex extracts with high confidence [3]. |
| 3-Nitrophenylhydrazine (3-NPH) | Derivatization reagent for carboxylic acids (e.g., SCFAs) to improve their chromatographic retention and mass spectrometric detection [5]. | Derivatization of short-chain fatty acids from microbial metabolism prior to LC-MS/MS analysis in plasma samples [5]. |
| Liquid Chromatography (LC) Columns | Separate complex mixtures of compounds before they enter the mass spectrometer. | Reversed-phase C18 columns are standard for separating NP extracts. Poly(vinyl alcohol)-based columns can be used for steric exclusion chromatography of underivatized SCFAs [5]. |
| Bioassay Kits & Reagents | Determine the biological activity of fractions or pure compounds (e.g., cytotoxicity, antimicrobial activity). | Used in parallel with chemical profiling to link a specific biological effect to one or more compounds in a mixture [3]. |
| Dereplication Databases | Computational tools containing spectral and biological data of known compounds to avoid rediscovery. | Screening HRMS and NMR data against databases to quickly identify known compounds in a bioactive hit extract [1]. |
The historical legacy of natural products in drug discovery is undeniable, providing some of the most important therapeutic agents for a range of debilitating diseases. The ongoing role of NPs is being secured by technological revolutions in analytical chemistry, particularly LC-MS/MS and TIMS-MS/MS, coupled with advanced bioassay techniques and machine learning. These tools are overcoming previous challenges by enabling the high-throughput deconvolution of complex natural extracts, the rapid identification of novel bioactive scaffolds, and the validation of their biosynthesis and mechanism of action. As these technologies continue to evolve, they will further unlock the immense, untapped potential of natural chemical diversity, ensuring that natural products remain a vital source of innovative lead compounds for the drug discovery pipeline for the foreseeable future.
Natural products (NPs) and their structural analogues have historically been major contributors to pharmacotherapy, particularly for cancer and infectious diseases, accounting for nearly 70% of new small molecule drugs approved over the past 40 years [6] [7]. Their profound success stems from evolutionary pre-optimization; NPs are "privileged structures" refined by nature to interact with biological macromolecules, resulting in superior biocompatibility, structural novelty, and functional diversity compared to purely synthetic compounds [8]. This review examines the experimental validation of NPs as drug leads through the integrated lens of LC-MS-based biosynthetic analysis and bioassay-guided research, providing a comparative assessment of these methodologies.
The Privileged Status of Natural Products in Drug Discovery
Evolutionary Optimization and Structural Superiority
Natural products possess distinct chemical properties resulting from prolonged evolutionary selection. Their scaffolds often exhibit greater stereochemical complexity and molecular rigidity than synthetic compounds, enabling highly specific interactions with protein targets [7]. This "pre-validated" biological relevance makes them ideal starting points for drug development, as they are inherently equipped to navigate complex biological systems [9].
Historical and Contemporary Impact
From 1981 to 2019, approximately 32% of newly introduced small molecule drugs were natural products or their direct derivatives, rising to nearly 70% in certain therapeutic areas like antimicrobials and anticancer agents [10] [7]. Notable examples include artemisinin (antimalarial), paclitaxel (anticancer), and resveratrol (investigated for Alzheimer's disease) [11]. This track record underscores their continued relevance in modern medicine.
Broad Therapeutic Target Engagement
The privileged status of NPs is further evidenced by their ability to interact with multiple protein targets, a polypharmacology that underpins their efficacy in treating complex diseases [8]. For instance, berberine directly binds to PKM2 to inhibit colorectal cancer progression, while curcumin exhibits multi-target anti-inflammatory activity [11] [8].
Analytical Frameworks for Validating Natural Product Biosynthesis and Bioactivity
Two primary methodological approaches—biosynthetic analysis via LC-MS and bioassay-guided isolation—enable researchers to decrypt the privileged structures of natural products.
LC-MS-Based Proteomic Investigation of Biosynthesis
Liquid Chromatography-Mass Spectrometry (LC-MS) has revolutionized the study of natural product biosynthesis by enabling direct detection of enzymatic intermediates and pathway mapping [12] [13].
Table 1: LC-MS Proteomic Platforms for Natural Product Biosynthesis Analysis
| Platform Type | Key Characteristics | Applications in NP Research | Key Insights Provided |
|---|---|---|---|
| Bottom-Up Proteomics | Analysis of protease-digested peptide fragments | High-throughput protein identification; mapping NRPS/PKS carrier domains [11] | Identifies expressed biosynthetic gene clusters; detects phosphopantetheinylation [12] |
| Top-Down Proteomics | Analysis of intact proteins and their post-translational modifications | Characterization of functional NRPS/PKS mega-enzymes [11] | Direct detection of acyl-/peptidyl-intermediates tethered to biosynthetic enzymes [13] |
| Data-Independent Acquisition (DIA) | Parallel fragmentation of all eluting ions | Comprehensive, unbiased detection of biosynthetic intermediates [11] | Provides systematic view of pathway dynamics and enzyme loading [12] |
| PrISM (Proteomic Investigation of Secondary Metabolism) | Selective detection of phosphopantetheinylated carrier proteins | Discovery of new natural products from environmental isolates without prior genome sequencing [12] | Links expressed NRPS/PKS enzymes to new natural products through carrier domain detection [12] |
The following workflow illustrates the PrISM methodology for discovering natural products through proteomic analysis:
Bioassay-Guided Isolation (BGI) and Metabolomic Approaches
Traditional bioassay-guided isolation remains a powerful method for identifying bioactive natural products through iterative fractionation and activity testing [10]. However, modern implementations increasingly integrate metabolomics to enhance efficiency and accuracy.
Table 2: Comparison of Natural Product Discovery Approaches
| Methodological Aspect | Bioassay-Guided Isolation (BGI) | Metabolomics-Based Discovery | Hybrid Strategies |
|---|---|---|---|
| Primary Focus | Biological activity-driven compound purification | Comprehensive chemical profiling coupled with statistical analysis | Integrates activity testing with chemical annotation [10] |
| Key Strengths | Direct linkage to bioactivity; historically proven success (e.g., artemisinin, paclitaxel) [10] | Broad chemical coverage; high sensitivity and throughput; reduces rediscovery [10] | Leverages strengths of both approaches; accelerates discovery timeline [10] |
| Common Limitations | Susceptible to masking effects; can miss minor active constituents; labor-intensive [10] | Indirect connection to bioactivity; requires sophisticated data analysis [10] | Requires multidisciplinary expertise; more complex workflow design [10] |
| Target Identification | Typically follows compound isolation | Can correlate chemical features with activity before isolation [6] | Provides both activity confirmation and comprehensive chemical data [10] |
Integrated Workflows: Leveraging Complementary Strengths
The most effective natural product discovery pipelines combine LC-MS biosynthetic analysis with bioassay validation in hybrid workflows.
Rational Library Minimization Strategy
A key advancement involves using LC-MS/MS and molecular networking to rationally reduce natural product library size while maximizing structural diversity and retaining bioactivity. This approach achieved an 84.9% reduction in library size needed to reach maximal scaffold diversity, while increasing bioassay hit rates from 11.3% to 22% in anti-Plasmodium assays [6].
The following diagram illustrates this library minimization process:
Chemical Proteomics for Target Identification
Chemical proteomics integrates synthetic chemistry, cellular biology, and mass spectrometry to comprehensively identify protein targets of natural products [8]. This approach uses designed probes that retain the pharmacological activity of parent natural compounds, enabling systematic target fishing from complex proteomes [8].
Experimental Protocols for Key Methodologies
Protocol 1: LC-MS-Based Proteomic Analysis of NRPS/PKS Systems (PrISM)
This protocol enables detection of phosphopantetheinylated carrier proteins in microbial proteomes [12]:
- Sample Preparation: Culture microbial isolates under conditions promoting secondary metabolism. Harvest cells during active production phase (typically early stationary phase).
- Protein Extraction: Lyse cells and separate proteins by SDS-PAGE. Excise high molecular weight bands (>200 kDa) for targeted analysis.
- In-Gel Digestion: Subject gel bands to in-gel tryptic digestion using standard protocols.
- LC-MS Analysis: Analyze peptides by nanoLC-MS/MS using high-resolution mass spectrometry (e.g., Linear Ion Trap-FTMS).
- Detection of Ppant-Modified Peptides: Monitor for phosphopantetheine ejection ions at m/z 261.1267 and 359.1036 with mass accuracy <2 ppm.
- Data Analysis: Perform de novo sequencing of modified peptides and database searching to identify NRPS/PKS biosynthetic machinery.
Protocol 2: Rational Natural Product Library Minimization
This protocol uses MS-based metabolomics to create focused screening libraries [6]:
- LC-MS/MS Data Acquisition: Perform untargeted LC-MS/MS on all extracts in the natural product library.
- Molecular Networking: Process MS/MS data through GNPS (Global Natural Products Social Molecular Networking) to group spectra into structural scaffolds based on fragmentation similarity.
- Scaffold Diversity Analysis: Calculate scaffold diversity across the library.
- Rational Selection: Apply iterative selection algorithm (custom R code) that prioritizes extracts with the greatest number of unique scaffolds not already represented.
- Library Generation: Continue selection until desired scaffold diversity threshold is reached (typically 80-100% of maximal diversity).
- Bioactivity Validation: Screen minimized library against biological targets and compare hit rates with full library.
The Scientist's Toolkit: Essential Research Reagents and Solutions
Table 3: Key Research Reagents and Platforms for Natural Product Drug Discovery
| Reagent/Platform | Function in NP Research | Application Examples |
|---|---|---|
| High-Resolution Mass Spectrometer (e.g., FTMS) | Enables accurate mass measurement (<2 ppm) for detection of Ppant ejection ions and metabolite identification [12] | Identification of NRPS/PKS carrier domain peptides; structural elucidation of new natural products [12] [13] |
| Activity-Based Probes (ABPP) | Chemical probes that retain parent compound activity while enabling target enrichment and identification [8] | Target fishing for natural products like celiptium and retapamulin; mapping compound-protein interactions [8] |
| Molecular Networking Platforms (e.g., GNPS) | Groups MS/MS spectra based on fragmentation similarity to identify structurally related compounds [6] | Scaffold-based library minimization; dereplication of known compounds from complex extracts [6] |
| Affinity Chromatography Matrices | Solid supports for immobilizing natural products to capture interacting proteins [8] | Identification of FKBP12 as FK506 target; discovery of histone deacetylase targets of trapoxin [8] |
| Standardized Cell Line Panels | Provides biologically relevant systems for evaluating NP efficacy and mechanism [11] | Testing anticancer effects in MCF-7, A549, HCT-116 lines; mechanism studies in HepG2 cells [11] |
| Bioinformatic Software (e.g., Skyline, Proteome Discoverer) | Processes LC-MS raw data for peptide/protein identification and quantification [11] | Analysis of proteomic changes in response to NP treatment; quantification of protein expression [11] |
Natural products rightfully hold their status as "privileged structures" in drug discovery, a designation strongly supported by experimental evidence from LC-MS-based biosynthetic analysis and bioassay research. The integration of these approaches provides a powerful framework for validating the unique biosynthetic origins and polypharmacology of natural products. As drug discovery evolves, the continued synergy of advanced analytical technologies with functional biological validation will ensure natural products remain indispensable sources of privileged scaffolds for addressing unmet medical needs.
Natural products, specialized metabolites produced by various organisms, remain an indispensable source of pharmaceutical agents, with approximately 32% of newly introduced small molecule drugs between 1981 and 2019 originating from these compounds [10]. Among the most pharmacologically significant classes are polyketides, nonribosomal peptides, and terpenoids, which are biosynthesized by complex enzymatic machinery [14]. These compounds exhibit remarkable structural diversity and potent biological activities, serving as antibiotics (e.g., erythromycin, tetracycline), immunosuppressants (e.g., cyclosporine), anticancer agents (e.g., doxorubicin), and insecticides [15] [16] [17].
The biosynthesis of these natural products is governed by specific enzyme assemblies encoded by biosynthetic gene clusters (BGCs) in microbial genomes [18] [17]. Advances in genome sequencing and bioinformatics have revealed that the number of predicted BGCs far exceeds the number of known compounds, suggesting vast untapped chemical diversity awaits discovery [18]. This guide focuses on comparing the biosynthetic pathways of polyketides, nonribosomal peptides, and terpenoids, with particular emphasis on methodologies for validating their production through LC-MS and bioassay techniques, providing researchers with essential tools for natural product discovery.
Comparative Analysis of Biosynthetic Pathways
The following table summarizes the core characteristics, enzymatic machinery, and key products of the three major classes of natural products.
Table 1: Comparative Overview of Major Biosynthetic Pathways
| Feature | Polyketides | Nonribosomal Peptides | Terpenoids |
|---|---|---|---|
| Core Biosynthetic Machinery | Polyketide Synthases (PKSs) [14] | Nonribosomal Peptide Synthetases (NRPSs) [14] | Terpene Cyclases/Synthases (TC/TS) [14] |
| Key Domains/Components | KS, AT, ACP, KR, DH, ER [16] | A, C, T [18] | N/A (Single or multi-domain enzymes) |
| Building Blocks | Acetyl-CoA, Malonyl-CoA, and other acyl-CoA derivatives [16] | Proteinogenic and non-proteinogenic amino acids [14] | Isopentenyl pyrophosphate (IPP), Dimethylallyl pyrophosphate (DMAPP) [14] |
| Assembly Mechanism | Sequential condensation and modification [16] | Template-directed, modular assembly [18] | Condensation of C5 units and cyclization [14] |
| Representative Products | Erythromycin, Tetracycline [16] [17] | Cyclosporine, Penicillin precursors [14] | Gibberellins, Carotenoids, Rhizovarins [14] |
| Bioinformatic Tool | AntiSMASH [14] [17] | AntiSMASH, RINPEP [18] | AntiSMASH [14] |
Polyketide Synthase (PKS) Pathways
Polyketide synthases are multidomain enzymes that assemble polyketides through the sequential condensation of acyl-CoA precursors [16]. They are categorized into three types. Type I PKSs are large, modular proteins where each module is responsible for one round of chain elongation; they can be further subdivided into cis-AT PKSs (where the acyltransferase domain is integrated within each module) and trans-AT PKSs (where the AT domain is a separate protein) [16]. Type II PKSs are complexes of discrete, monofunctional enzymes that work iteratively to produce aromatic polyketides [17]. Type III PKSs (chalcone synthase-like) are simpler, homodimeric enzymes that also operate iteratively [14].
The synthesis process mediated by cis-AT PKSs involves three stages: initiation, elongation, and termination [16]. During initiation, the AT domain selects a starter unit and loads it onto the corresponding Acyl Carrier Protein (ACP). In the elongation stage, the Ketosynthase (KS) domain catalyzes a condensation reaction between the growing polyketide chain and an ACP-bound extender unit. Subsequent processing by optional domains like Ketoreductase (KR), Dehydratase (DH), and Enoylreductase (ER) introduces functional groups. Finally, the Thioesterase (TE) domain catalyzes termination through cyclization or hydrolysis, releasing the final polyketide product [16].
Nonribosomal Peptide Synthetase (NRPS) Pathways
Nonribosomal peptide synthetases are modular assembly lines that synthesize peptides without a mRNA template [14] [18]. Each NRPS module is responsible for incorporating one monomeric building block into the growing peptide chain and typically contains three core domains [18]. The Adenylation (A) domain recognizes and activates a specific amino acid substrate. The Condensation (C) domain catalyzes the formation of a peptide bond between the growing chain and the new amino acid. The Thioesterification (T) domain (often synonymous with the Peptidyl Carrier Protein, PCP) shuttles the intermediates between domains. The final module often contains a Termination (Te) domain that releases the mature peptide, often through cyclization [18].
A remarkable feature of NRPSs is their substrate promiscuity, allowing for the incorporation of hundreds of different proteinogenic and non-proteinogenic amino acids, leading to immense structural diversity [18]. The resulting peptides often undergo further post-assembly modifications, such as cyclization, glycosylation, or methylation, which enhance their structural complexity and biological stability [14].
Terpenoid Biosynthetic Pathways
Terpenoids, also known as isoprenoids, represent one of the largest and most structurally diverse families of natural products [14]. Their biosynthesis proceeds via two primary pathways: the mevalonate (MVA) pathway in eukaryotes and some bacteria, and the non-mevalonate (MEP) pathway in prokaryotes and plant plastids. Both pathways produce the universal five-carbon building blocks, isopentenyl pyrophosphate (IPP) and dimethylallyl pyrophosphate (DMAPP) [14].
The pathway begins with the condensation of IPP and DMAPP to form geranyl pyrophosphate (G10), which can be further elongated to farnesyl pyrophosphate (F15) and geranylgeranyl pyrophosphate (G20). Terpene Cyclases or Synthases then catalyze the conversion of these linear prenyl diphosphates into the parent carbon skeletons of mono-, sesqui-, and diterpenes, respectively [14]. These hydrocarbon skeletons are often subsequently modified by various tailoring enzymes (e.g., oxidoreductases, methyltransferases) to produce the vast array of known terpenoid structures, which perform essential ecological functions as phytohormones, pigments, and defense compounds [14].
Experimental Validation: LC-MS and Bioassay Methodologies
The identification and validation of natural products require a combination of sophisticated analytical and biological techniques. The following sections detail key experimental protocols.
LC-MS/MS-Based Proteomics for Enzyme Detection
Liquid chromatography-mass spectrometry (LC-MS)-based proteomics is a powerful high-throughput technique for profiling protein expression in cells, and can be used to screen for expressed NRPSs and PKSs from bacterial strains [19].
Table 2: Key Steps in LC-MS/MS Proteomics for PKS/NRPS Detection
| Step | Procedure | Purpose | Key Reagents/Equipment |
|---|---|---|---|
| 1. Protein Extraction | Lyse bacterial cells from a given strain and growth condition. | To release the full complement of cellular proteins. | Lysis buffer, Protease inhibitors, Centrifuge [19] |
| 2. Size-Based Separation | Separate proteins by SDS-PAGE (Sodium Dodecyl Sulfate-Polyacrylamide Gel Electrophoresis). | To enrich for large, modular NRPSs and PKSs (often >200 kDa). | SDS-PAGE apparatus, Molecular weight markers [19] |
| 3. Tryptic Digestion | Excise gel bands and digest proteins enzymatically (e.g., with trypsin). | To break down proteins into smaller peptides for MS analysis. | Trypsin, Digestion buffer [19] |
| 4. LC-MS/MS Analysis | Separate peptides by liquid chromatography and analyze via tandem mass spectrometry. | To acquire fragmentation spectra (MS/MS) for peptide identification. | Nano-LC system, High-resolution mass spectrometer [19] |
| 5. Data Analysis | Search MS/MS spectra against protein databases using specialized software. | To identify proteins, pinpoint expressed NRPS/PKS gene clusters. | Search algorithms (e.g., Mascot, Sequest), Genomic databases [19] |
Bioassay-Guided Isolation (BGI)
Bioassay-guided isolation is a classical approach where a crude natural extract is fractionated, and each fraction is tested for a desired biological activity (e.g., antimicrobial, anticancer). The active fractions are subsequently subjected to further purification steps, guided by the bioassay results at each stage, until the active compound(s) are isolated [20] [10]. A key, occasionally neglected aspect of BGI is the careful design of the bioassay to ensure it is specific, reproducible, and relevant to the intended therapeutic target [20].
Representative Protocol for Antimicrobial Bioassay:
- Extract Preparation: Plant or microbial material is extracted with solvents of varying polarity (e.g., hexane, ethyl acetate, methanol, water) to obtain a range of extracts [21].
- Agar Well Diffusion Assay:
- Test bacterial strains (e.g., Staphylococcus aureus, Pseudomonas aeruginosa) are spread onto Mueller-Hinton agar plates.
- Wells are punched into the agar and filled with the test extracts and fractions.
- Plates are incubated, and the zones of inhibition around the wells are measured to determine antimicrobial activity [21].
- Fractionation: Active extracts are fractionated using techniques like vacuum liquid chromatography (VLC) or solid-phase extraction (SPE).
- Iterative Testing: All resulting fractions are tested again in the antimicrobial assay. The active fractions are subjected to further purification (e.g., using preparative HPLC) until pure active compounds are obtained [10] [21].
LC-MS Metabolomics for Compound Discovery and Dereplication
Liquid Chromatography-Mass Spectrometry (LC-MS) metabolomics provides a broad, high-throughput platform for characterizing the chemical profile of natural extracts. This approach is crucial for dereplication—the early identification of known compounds to avoid rediscovery—and for prioritizing novel metabolites for isolation [10] [21].
Protocol for LC-HRMS² Analysis of Plant Extracts:
- Sample Preparation: Extract powdered plant material with solvents (e.g., methanol, aqueous) and concentrate under reduced pressure or freeze-drying [21].
- LC-HRMS Analysis:
- Analyte separation is performed using a UHPLC system with a reverse-phase C18 column.
- The LC system is coupled to a high-resolution mass spectrometer (e.g., Q-TOF) equipped with an Electrospray Ionization (ESI) source.
- MS scans are operated in full-scan mode (e.g., m/z 100-1700) [21].
- Data-Dependent MS/MS:
- Following the MS1 scan, the most intense ions are automatically selected for fragmentation (MS/MS).
- This generates structural information based on the fragmentation patterns [21].
- Data Processing and Metabolite Identification:
- Raw data is processed using software (e.g., MassHunter).
- Accurate mass and isotopic patterns are used to predict molecular formulae.
- Metabolites are identified by comparing acquired MS2 spectra and retention times with those of reference standards or databases. Confidence levels are assigned as proposed by the Metabolomics Standards Initiative [21].
The workflow below illustrates the hybrid strategy that combines BGI and metabolomics for efficient natural product discovery.
Diagram 1: Hybrid discovery workflow integrating LC-MS and bioassay.
Advanced Engineering and Screening Strategies
Heterologous Expression and Engineering of Biosynthetic Pathways
Many BGCs are "cryptic" and not expressed under laboratory conditions. Heterologous expression is a key strategy to activate these silent clusters by transferring them into a well-characterized host organism [22]. Aspergillus oryzae is a frequently used host for expressing fungal BGCs due to its clean metabolic background, available genetic tools, and robust precursor supply [22].
Recent innovations include the development of plug-and-play vectors for A. oryzae. These vectors contain multiple, different promoter-terminator expression cassettes with unique restriction sites, facilitating the simultaneous reconstruction of entire biosynthetic pathways comprising multiple genes. This system, combined with LC-MS screening of transformants grown on simple CD agar plates, can save over ten days compared to traditional methods that rely on PCR screening and fermentation in rich media [22].
Engineering of the biosynthetic machinery itself is a powerful approach to improve yields or generate novel analogs. A 2025 study on the butenyl-spinosyn modular PKS (mPKS) revealed that a majority (>93%) of PKS mRNAs are truncated, leading to non-functional polypeptide fragments. Splitting the large 13-kb busA gene (encoding a 456-kDa PKS) into three smaller, separately translated genes encoding single modules rescued the translation of truncated mRNAs and increased the biosynthetic efficiency by 13-fold. This strategy has also been successfully applied to other megasynthases, such as those for avermectin and epothilone [15].
Table 3: Rational Engineering Strategies for cis-AT Polyketide Synthases
| Engineering Strategy | Description | Key Consideration | Outcome/Example |
|---|---|---|---|
| Module/Domain Swapping | Exchanging entire modules or specific catalytic domains between PKSs to create chimeric systems. | Requires compatible protein-protein interactions and docking interfaces to maintain function. | Synthesis of chimeric polyketides with altered backbones or functional groups [16]. |
| Active-Site Engineering | Using site-directed mutagenesis to alter the specificity of a domain, most commonly the AT domain. | Requires high-resolution structural knowledge of the target domain. | Production of polyketides with non-natural extender units or altered stereochemistry [16]. |
| mRNA Truncation Rescue | Splitting large, multi-module PKS genes into smaller, separately translated genes. | Requires addition of heterologous docking domains (NDD/CDD) to maintain module communication. | 13-fold yield improvement for butenyl-spinosyn; broader application to other mPKSs [15]. |
The Scientist's Toolkit: Essential Research Reagents and Solutions
Table 4: Key Reagent Solutions for Natural Product Biosynthesis Research
| Reagent/Solution | Function/Application | Example Use Case |
|---|---|---|
| AntiSMASH Software | A bioinformatic tool for the genome-wide identification, annotation, and analysis of BGCs. | Predicting core structures of nonribosomal peptides and identifying hybrid NRPS-PKS clusters [14] [18]. |
| Heterologous Expression Vectors (e.g., pUARA2, pUSA2) | Plasmids designed for the reconstruction and expression of multiple genes from a BGC in a host like A. oryzae. | Expressing the rugulosin biosynthetic gene cluster in Aspergillus oryzae [22]. |
| Docking Domain Sequences | Genetic sequences encoding N- and C-terminal docking domains that facilitate interaction between PKS subunits. | Enabling communication between split PKS modules after rescuing mRNA truncation [15]. |
| LC-MS Grade Solvents | High-purity solvents for liquid chromatography and mass spectrometry to minimize background noise and ion suppression. | Preparing samples for LC-HRMS² analysis of plant extracts for metabolomic profiling [21]. |
| Bioassay Reagents | Materials for biological activity testing, such as culture media, bacterial strains, and indicator compounds. | Conducting antimicrobial agar well diffusion assays against Staphylococcus aureus [21]. |
Polyketides, nonribosomal peptides, and terpenoids represent three pillars of natural product discovery, each with distinct biosynthetic logic and engineering potential. The future of the field lies in the intelligent integration of complementary methodologies. While bioassay-guided isolation provides direct evidence of bioactivity, LC-MS metabolomics offers unparalleled breadth in chemical coverage and dereplication power [10]. Combining these with heterologous expression and rational protein engineering creates a powerful, hybrid strategy for accelerating the discovery and development of novel therapeutic agents from nature's vast chemical repertoire [22] [15] [10].
In the pursuit of engineering organisms to produce valuable natural products, validation stands as the critical gatekeeper ensuring that metabolic interventions yield the intended results. The fidelity of biosynthetic engineering—whether for pharmaceutical development, nutritional enhancement, or bio-based chemical production—hinges on robust analytical verification. Without rigorous validation, engineered pathways may produce unexpected metabolites, accumulate toxic intermediates, or fail to achieve target yields, compromising both scientific integrity and practical applications. This guide examines the complementary roles of liquid chromatography-tandem mass spectrometry (LC-MS/MS) and bioassay methodologies in providing this essential validation, offering researchers a framework for selecting appropriate techniques based on their specific project requirements, constraints, and objectives.
The challenges in biosynthetic engineering are substantial: introduced pathways may encounter flux imbalances, enzyme incompatibilities, or unexpected regulatory interactions within host organisms [23]. As engineering strategies grow more ambitious—shifting from single-gene insertions to complex pathway implementations—the potential for deviation from predicted outcomes increases accordingly. Consequently, validation technologies must evolve beyond simple confirmation of product presence to provide comprehensive metabolic profiling, quantitative accuracy, and functional assessment of biosynthetic output.
Analytical Technique Comparison: LC-MS/MS Versus Bioassays
Core Principles and Applications
LC-MS/MS combines the separation power of liquid chromatography with the detection specificity and sensitivity of tandem mass spectrometry, enabling precise identification and quantification of target compounds and their biosynthetic intermediates in complex biological matrices [24] [25]. This technology excels at detecting structural analogues, phosphorylation states, and pathway intermediates with high specificity, making it indispensable for detailed metabolic characterization.
Bioassays, particularly microbiological assays using engineered microorganisms, leverage biological responsiveness to determine metabolite levels through growth-based or turbidimetric measurements [24]. These assays employ strains with specific auxotrophies or biosynthetic deficiencies that are complemented by the compound of interest, providing functional readouts of metabolic activity.
Performance Comparison Table
The following table summarizes the key characteristics of each validation methodology:
Table 1: Comparative Analysis of Biosynthetic Validation Techniques
| Parameter | LC-MS/MS | Microbiological Bioassays |
|---|---|---|
| Sensitivity | High (capable of detecting compounds at nanogram-per-milliliter levels) [25] | Moderate (sufficient for many metabolic engineering applications) [24] |
| Specificity | Excellent (discriminates between closely related structures and phosphorylation states) [24] | Variable (may respond to multiple related metabolites unless carefully designed) [24] |
| Quantitative Accuracy | High (with proper internal standardization; precision of 3.23–14.26% RSD) [25] | Semi-quantitative (may show discrepancies compared to reference methods) [24] |
| Throughput | Moderate (extensive sample preparation required) [24] | High (amenable to parallel processing and rapid screening) [24] |
| Equipment Requirements | Specialized, expensive instrumentation requiring technical expertise [24] | Standard laboratory equipment, minimal specialized instrumentation [24] |
| Intermediate Detection | Comprehensive (can detect and quantify biosynthetic intermediates) [24] | Limited (requires specialized panel of mutant strains) [24] |
| Functional Assessment | No (provides chemical information only) | Yes (demonstrates biological activity and bioavailability) [24] |
| Cost per Sample | High (reagents, instrumentation, maintenance) | Low (minimal reagent costs, no specialized equipment) [24] |
Technique Selection Guidelines
The choice between these methodologies depends on project goals, resources, and development stage:
- Early-stage pathway screening benefits from bioassay throughput and cost-efficiency, enabling rapid evaluation of multiple engineered variants [24].
- Detailed metabolic characterization requires LC-MS/MS specificity to identify bottlenecks, intermediate accumulation, or unexpected side products [24] [23].
- Functional validation of bioactive compounds gains from bioassay confirmation that the produced metabolite exhibits expected biological activity [24].
- Regulatory submissions typically demand LC-MS/MS validation for precise quantification and comprehensive metabolic profiling [26] [25].
Experimental Protocols for Biosynthetic Validation
LC-MS/MS Method for Metabolite Quantification
Protocol Overview: This method enables precise quantification of target metabolites and their biosynthetic intermediates in biological samples, using the example of thiamin vitamers from Arabidopsis thaliana [24] and LXT-101 from beagle plasma [25] as representative applications.
Figure 1: LC-MS/MS Experimental Workflow
Materials and Reagents:
- Extraction solvents: Methanol, acetonitrile, acidified aqueous solutions [24] [25]
- Internal standards: Stable isotope-labeled analogues (e.g., cortisol-d4 for cortisol analysis) [26]
- LC columns: Reversed-phase C18 or C8 columns (e.g., Hypersil GOLD C18, 50 mm × 2.1 mm, 5 μm) [25]
- Mobile phases: Acetonitrile/water with modifiers (e.g., 0.1% formic acid) [25]
- Calibration standards: Authentic reference compounds for quantification [25]
Detailed Procedure:
- Sample Preparation: Homogenize biological material (plant tissue, microbial cells) in extraction solvent. For tissue analysis, use approximately 100 mg fresh weight extracted with 1 mL methanol:water (70:30, v/v) with 0.1% formic acid [24].
- Extraction: Subject samples to vortex mixing followed by centrifugation at 12,000-15,000 × g for 10-15 minutes at 4°C [25].
- Concentration: Transfer supernatant and evaporate under nitrogen stream at 37°C. Reconstitute in initial mobile phase composition [25].
- Chromatographic Separation:
- Mass Spectrometric Detection:
- Ionization: Electrospray ionization (ESI) in positive or negative mode
- Scan Mode: Selected reaction monitoring (SRM) or multiple reaction monitoring (MRM)
- Parameters: Optimized collision energies, source temperatures, and gas flows for target compounds
- Data Analysis: Quantify using internal standard method with calibration curves (typically 2-600 ng/mL range) [25].
Validation Parameters:
- Linearity: R² ≥ 0.997 over relevant concentration range [25]
- Precision: Intra- and inter-batch precision ≤15% RSD [25]
- Accuracy: 85-115% of nominal values [25]
- Recovery: Consistent extraction efficiency (75-126%) [25]
Microbiological Bioassay for Metabolic Screening
Protocol Overview: This panel-based yeast assay enables functional assessment of vitamin B1 and its biosynthetic intermediates in plant materials, using Saccharomyces cerevisiae mutants with specific auxotrophies [24].
Figure 2: Bioassay Experimental Workflow
Materials and Reagents:
- Yeast Strains: Panel of Saccharomyces cerevisiae mutants (e.g., thi4 thiazole auxotroph, thi5 pyrimidine auxotroph) [24]
- Growth Media: Minimal media lacking specific metabolites [24]
- Reference Standards: Pure thiamin, phosphorylated derivatives, and biosynthetic intermediates [24]
- Extraction Solvents: Appropriate for releasing metabolites from biological matrices [24]
Detailed Procedure:
- Strain Preparation: Maintain yeast strains on complete media, then transfer to minimal media to create metabolite starvation prior to assay.
- Sample Extraction: Prepare tissue extracts using methods that preserve metabolite integrity (e.g., mild acid extraction for thiamin vitamers) [24].
- Assay Setup:
- Dilute sample extracts in minimal media
- Inoculate with standardized yeast culture
- Include calibration standards and negative controls
- Dispense into multi-well plates for high-throughput processing
- Incubation: Grow cultures with shaking at 30°C for 24-48 hours [24].
- Growth Measurement: Quantify turbidity at 600 nm using plate reader [24].
- Data Analysis: Compare sample growth to standard curve to determine metabolite concentration.
Key Considerations:
- Strain Specificity: Different mutants respond to different metabolic precursors, enabling pathway intermediate profiling [24].
- Matrix Effects: Biological extracts may contain compounds that inhibit or stimulate yeast growth, requiring appropriate controls.
- Quantification Limits: The dynamic range should be established for each metabolite-strain combination.
Research Reagent Solutions: Essential Materials for Biosynthetic Validation
Table 2: Essential Research Reagents for Biosynthetic Validation Studies
| Reagent Category | Specific Examples | Function/Application | Technical Notes |
|---|---|---|---|
| Chromatography Columns | Hypersil GOLD C18 (50 mm × 2.1 mm, 5 μm) [25] | Separation of metabolites prior to mass spectrometric detection | Reversed-phase chemistry suitable for diverse metabolite classes |
| Mass Spectrometry Internal Standards | Cortisol-d4 [26], stable isotope-labeled analogues | Normalization of extraction efficiency and ionization variability | Should be structurally analogous to target analytes |
| Reference Standards | Thiamin, TMP, TPP, HMP, HET [24] | Method calibration and quantification | High-purity characterized compounds essential for accurate quantification |
| Bioassay Organisms | S. cerevisiae thi4 mutant [24] | Functional assessment of specific metabolites through growth response | Specific auxotrophies determine metabolite responsiveness |
| Extraction Solvents | Acidified methanol, acetonitrile with formic acid [24] [25] | Metabolite extraction from biological matrices | Solvent composition optimized for target metabolite stability and solubility |
| Mobile Phase Additives | Formic acid (0.1%) [25] | Enhance ionization efficiency in mass spectrometry | Concentration critical for optimal signal intensity |
Integrated Validation Strategies: Case Studies in Natural Product Biosynthesis
Thiamin Biofortification in Plants
In metabolic engineering of thiamin biosynthesis in Arabidopsis thaliana, researchers implemented a dual validation approach using both LC-MS/MS and yeast bioassays [24]. The LC-MS/MS method provided absolute quantification of thiamin, its phosphorylated derivatives (TMP, TPP), and biosynthetic intermediates (HMP, HET) with high specificity, enabling precise assessment of metabolic engineering outcomes [24]. Concurrently, a panel of yeast assays using strains auxotrophic for different thiamin pathway intermediates offered functional validation and the ability to screen large numbers of engineered lines rapidly [24].
This integrated approach revealed that while both methods correctly identified high-thiamin lines, the bioassay results showed discrepancies in absolute values compared to LC-MS/MS, confirming its utility as a semi-quantitative screening tool rather than a definitive quantification method [24]. The combination allowed efficient screening of numerous engineered lines followed by detailed characterization of promising candidates.
Natural Product Discovery and Pathway Elucidation
In the investigation of γ-lactone biosynthesis in Sextonia rubra wood, TOF-SIMS MS/MS imaging enabled in situ localization and characterization of biosynthetic intermediates at subcellular resolution (~400 nm) [27]. This spatial information proved crucial for proposing a revised biosynthetic pathway involving the reaction between 2-hydroxysuccinic acid and 3-oxotetradecanoic acid, contrary to previous hypotheses suggesting a single polyketide precursor [27]. The methodology combined the structural characterization power of MS/MS with spatial resolution sufficient to localize metabolites to specific cell types (ray parenchyma cells and oil cells) [27].
Pharmaceutical Development and Preclinical Validation
In the development of LXT-101 sustained-release suspension for prostate cancer treatment, a validated LC-MS/MS method provided critical pharmacokinetic data in beagle dog models [25]. The method demonstrated appropriate linearity (2-600 ng/mL, R²=0.9977), precision (intra-batch RSD 3.23-14.26%), and accuracy (93.36-99.27%) to support regulatory submissions [25]. This application highlights the role of robust validation methodologies in translating biosynthetic engineering achievements into clinically relevant therapeutics.
The fidelity of biosynthetic engineering depends fundamentally on appropriate validation strategies that match methodological capabilities to project requirements. LC-MS/MS provides the specificity, sensitivity, and quantitative rigor necessary for definitive characterization of engineered metabolic pathways, particularly when precise quantification of multiple metabolites and intermediates is required [24] [25]. Bioassays offer complementary strengths in functional assessment, throughput, and cost-effectiveness, making them invaluable for screening applications and initial pathway validation [24].
The most effective biosynthetic engineering initiatives implement these technologies as complementary rather than competing approaches, leveraging their respective strengths at appropriate stages of project development. As synthetic biology continues to expand its capabilities toward increasingly complex natural products, robust validation methodologies will remain essential for bridging the gap between genetic design and functional metabolic outcomes, ensuring that engineered biological systems deliver on their theoretical promise.
Advanced LC-MS Methodologies and Bioassay Integration for Biosynthetic Analysis
The validation of natural product biosynthesis represents a complex analytical challenge, requiring the precise identification and quantification of target metabolites within intricate biological matrices. Modern liquid chromatography-mass spectrometry (LC-MS) technologies have become indispensable in this field, providing the separation power, mass accuracy, and structural elucidation capabilities necessary to decipher biosynthetic pathways. The combination of ultra-high-performance liquid chromatography (UHPLC) with high-resolution mass spectrometry (HRMS) has emerged as a particularly powerful platform, enabling researchers to achieve unprecedented levels of analytical performance [28]. This technological synergy has transformed natural product research by facilitating comprehensive metabolite profiling with enhanced speed, sensitivity, and selectivity.
Recent advancements have further expanded this analytical toolbox with the introduction of high-resolution ion mobility (HRIM) separation, which adds a rapid separation dimension based on the size, charge, and shape of ionized molecules [29]. This review provides a systematic comparison of current state-of-the-art LC-MS instrumentation, with a specific focus on applications within natural product biosynthesis validation and bioassay research. By examining the complementary strengths of UHPLC, HRMS, and ion mobility technologies, we aim to provide researchers with a practical framework for selecting appropriate instrumentation for their specific analytical challenges in drug discovery and development.
UHPLC Technology: Enhanced Separation for Complex Samples
Fundamental Principles and Performance Gains
UHPLC technology represents a significant advancement over conventional HPLC, primarily through the utilization of sub-2-µm particle columns coupled with instrumentation capable of operating at significantly higher pressures (typically up to 1000-1300 bar) [28]. This fundamental improvement has yielded substantial gains in separation efficiency, analysis speed, and detection sensitivity. The reduced particle size increases the surface area for interaction, resulting in superior chromatographic resolution, while the higher pressure capabilities enable optimal mobile phase linear velocities for these particles. The commercial introduction of UHPLC systems raised the long-held 400-bar pressure limit of traditional LC pumps to 1000 bar, simultaneously reducing system dead volumes throughout the instrumentation [28].
The practical benefits of UHPLC are particularly valuable in natural product research, where analysts frequently encounter complex samples containing compounds with widely varying concentrations and chemical properties [28]. The enhanced resolution allows for the separation of structurally similar metabolites, including isomers that may play distinct roles in biosynthetic pathways. Furthermore, the improved peak sharpness associated with UHPLC separations directly translates to lower detection limits, enabling researchers to identify and quantify trace-level metabolites that might function as pathway intermediates or regulatory molecules.
Current UHPLC Instrumentation Landscape
The market for UHPLC instrumentation has expanded significantly, with all major chromatography vendors now offering sophisticated systems. Recent product introductions from 2024-2025 demonstrate continued innovation in this field, as highlighted in Table 1.
Table 1: Recent UHPLC System Introductions (2024-2025)
| Vendor | System Model | Maximum Pressure (bar) | Key Features | Target Applications |
|---|---|---|---|---|
| Agilent | Infinity III 1290 | 1300 | Binary or quaternary pump, flow rates up to 5 mL/min | High-resolution separations, method development |
| Waters | Alliance iS Bio HPLC | 830 (12,000 psi) | Bio-inert design with MaxPeak HPS technology, pH range 1-13 | Biopharmaceutical QC, biomolecule analysis |
| Shimadzu | i-Series HPLC/UHPLC | 1015 (70 MPa) | Compact, integrated design, eco-friendly operation | General LC applications supporting various detectors |
| Thermo Fisher Scientific | Vanquish Neo | Not specified | Tandem direct injection workflow for parallel column operation | High-throughput analysis, reduced carryover |
| Knauer | Azura HTQC UHPLC | 1240 | High-throughput configuration, flow rates up to 10 mL/min | Quality control applications |
These recent systems incorporate features such as bio-inert flow paths for analyzing corrosive mobile phases, advanced automation for improved reproducibility, and specialized workflows for specific application needs [30]. The trend toward more compact, energy-efficient designs with reduced operational costs is also evident, making UHPLC technology increasingly accessible to routine laboratories.
High-Resolution Mass Spectrometry: Structural Elucidation Power
Mass Analyzer Technologies and Performance Characteristics
High-resolution mass spectrometry has undergone revolutionary advancements, primarily driven by the improved performance and accessibility of time-of-flight (TOF) and Orbitrap mass analyzers [28]. These technologies have addressed previous limitations of high-resolution instruments concerning speed, dynamic range, and operational complexity, making them viable for routine applications in natural product research. The fundamental advantage of HRMS lies in its ability to provide accurate mass measurements with errors typically less than 5 ppm, enabling the determination of elemental compositions with high confidence—a critical capability for identifying unknown metabolites in biosynthetic pathway elucidation.
The performance comparison of modern HRMS technologies reveals distinct strengths for different applications. TOF instruments offer high acquisition speeds (up to 1000 spectra/second) and mass resolutions typically ranging from 40,000 to 100,000, making them well-suited for coupling with UHPLC where fast detection is essential to capture narrow chromatographic peaks [28]. Orbitrap technology provides even higher resolution capabilities (ranging from 100,000 to 500,000+), with improved sensitivity for targeted applications, though at generally lower acquisition rates than TOF systems. Recent introductions in the HRMS market include systems like the Sciex ZenoTOF 7600+, which incorporates Zeno Trap Technology and Electron Activated Dissociation (EAD) for advanced structural characterization, particularly beneficial for proteomics and biomarker research [30].
Tandem and Multi-stage MS for Confident Identification
The combination of high-resolution capabilities with tandem mass spectrometry (MS/MS) has proven particularly powerful for natural product identification. MS/MS provides fragmentation data that reveals structural details beyond what can be determined from mass measurement alone. Recent research has further explored the benefits of MS3 capabilities, where a second generation of product ions is generated from primary fragments, providing even deeper structural information [31].
A systematic comparison of LC-HR-MS2 and LC-HR-MS3 for screening toxic natural products demonstrated that while both approaches provided identical identification results for most analytes (96% in serum, 92% in urine), the MS2-MS3 data analysis showed better performance for a small subset of compounds at lower concentrations [31]. This enhanced performance comes at the cost of increased method complexity and potentially reduced number of compounds that can be analyzed in a single run, as the instrument must spend more time performing sequential fragmentation events.
Table 2: Comparison of Mass Analyzer Technologies for Natural Product Research
| Mass Analyzer Type | Mass Resolution | Mass Accuracy (ppm) | Acquisition Speed | Key Strengths | Natural Product Applications |
|---|---|---|---|---|---|
| Q-TOF | 40,000-100,000 | <5 | Very High | Fast data acquisition, good dynamic range | Untargeted metabolomics, metabolite profiling |
| Orbitrap | 100,000-500,000+ | <3 | Moderate to High | Very high resolution and mass accuracy | Structural elucidation, targeted analysis |
| TQ-MS (QqQ) | Unit Resolution | N/A | High | Excellent sensitivity, quantitative precision | Targeted quantification of known metabolites |
| MALDI-TOF/TOF | 20,000-40,000 | <10 | Moderate | Spatial imaging, solid samples | Tissue imaging in plant research |
Ion Mobility Spectrometry: Adding a Separation Dimension
High-Resolution Ion Mobility Fundamentals
High-resolution ion mobility represents a significant advancement in separation science, operating on fundamentally different principles than liquid chromatography. While LC separates molecules based on their chemical interactions with stationary and mobile phases, ion mobility separates ionized molecules based on their collision cross section (CCS), size, charge, and overall shape in the gas phase [29]. This separation occurs in milliseconds rather than minutes, providing an additional orthogonal separation dimension that can be coupled with LC-MS analysis.
The distinguishing feature of HRIM technology based on Structures for Lossless Ion Manipulation (SLIM) is the implementation of exceptionally long separation pathlengths (commercially available systems feature a 40-foot path) packed into a device approximately the size of a laptop through serpentine electrode patterns on printed circuit board technology [29]. This design enables separation resolutions unattainable with conventional ion mobility techniques, while essentially eliminating ion losses that have historically limited the sensitivity of mobility-based separations.
Application Advantages in Natural Product Research
The unique separation mechanism of HRIM offers particular advantages for challenging separations in natural product research, especially for isomeric compounds that are difficult to distinguish by mass or chromatography alone. This capability is invaluable for studying biosynthetic pathways where multiple isomers may be present as intermediates or related products. HRIM has demonstrated exceptional performance in areas that have been notoriously challenging with conventional LC-MS, particularly lipid and glycan analysis [29]. These biomolecular classes exhibit extensive isomeric diversity and structural heterogeneity that complicate their analysis by traditional methods.
A key practical advantage of HRIM is its analyte-agnostic nature—unlike LC, which often requires matching column chemistry to specific separations, the same HRIM instrument can resolve multiple classes of analytes (glycans, peptides, proteins, small molecules) without hardware changes [29]. This flexibility significantly increases laboratory productivity when working with diverse sample types, a common scenario in natural product research where researchers may analyze various compound classes from the same biological source.
Integrated Experimental Workflows in Natural Product Research
LC-MS/MS with Molecular Networking for Metabolite Profiling
The integration of UHPLC separation with tandem mass spectrometry has enabled sophisticated analytical workflows for natural product discovery and biosynthesis validation. One particularly powerful approach combines LC-MS/MS analysis with molecular networking through platforms such as the Global Natural Products Social Molecular Networking (GNPS) website [32]. This workflow enables untargeted metabolite profiling where metabolites present in extracts and chromatography fractions can be annotated based on their MS/MS fragmentation patterns, with structurally related molecules clustered together in visual networks.
This methodology was successfully implemented in an undergraduate laboratory course focused on identifying metabolites from medicinal plants, demonstrating its practical accessibility [32]. Students first extracted plant specimens such as rosemary, aloe, echinacea, and ashwagandha, then performed bioactivity assessments using antioxidant (DPPH) assays. Active extracts were fractionated using solid-phase extraction, followed by LC-DAD-MS/MS analysis on a Thermo Fisher Scientific LTQ XL mass spectrometer. The resulting MS/MS spectra were processed through the GNPS platform to create molecular networks and compared against MS/MS spectral libraries for metabolite identification, introducing students to cutting-edge dereplication techniques essential for modern natural product research.
Diagram 1: LC-MS/MS and Molecular Networking Workflow for Natural Product Research. This workflow integrates biological screening with advanced mass spectrometry and computational analysis for comprehensive metabolite profiling.
Bioassay-Guided Fractionation with LC-MS Detection
Another established approach in natural product research combines bioassay-guided fractionation with LC-MS detection to rapidly identify bioactive constituents. This methodology was exemplified in research on Picria fel-terrae, a traditional Chinese medicine, where investigators sought to identify acetylcholinesterase (AChE) inhibitors [33]. Following primary extraction, the ethyl acetate fraction showed strong AChE inhibitory activity and was selected for further investigation.
The analytical workflow involved separation by HPLC with the eluate collected in 96-well plates using a fraction collector. After solvent removal, the residues in each well were tested for AChE inhibitory activity. Positive wells were subsequently analyzed by LC-ESI-MS for compound identification. This integrated approach detected six active compounds, identified as various picfeltarraenins, which showed stronger AChE inhibition than the known inhibitor Tacrine [33]. The combination of biological screening with chromatographic separation and mass spectrometric detection provides a powerful strategy for pinpointing bioactive natural products without the need for extensive isolation of inactive constituents.
Comprehensive Two-Dimensional Liquid Chromatography (LC×LC)
For exceptionally complex samples, comprehensive two-dimensional liquid chromatography (LC×LC) coupled to mass spectrometry offers enhanced separation capabilities beyond what can be achieved with one-dimensional separations. This approach has been successfully applied to food and natural product samples, providing unparalleled selectivity and sensitivity for detecting minor bioactive components [34].
Advanced LC×LC–MS techniques employ different separation mechanisms in each dimension (e.g., reversed-phase × reversed-phase or HILIC × reversed-phase) to maximize orthogonality, along with focusing modulation strategies to achieve precise separations and accurate quantification [34]. The incorporation of microLC in the first-dimension separation improves reliability and consistency of retention times, while the comprehensive nature of the separation enables detection and identification of minor components that are challenging to isolate using conventional LC methods. This approach has been validated through satisfactory limits of detection, limits of quantification, and high intraday and interday precision, establishing it as a powerful tool for the qualitative and quantitative assessment of complex natural product mixtures.
Comparative Performance Data and Experimental Protocols
Quantitative Comparison of LC-MS Platforms
The selection of an appropriate LC-MS platform depends heavily on the specific analytical requirements and sample characteristics. Different configurations offer distinct advantages for targeted versus untargeted analyses, qualitative versus quantitative applications, and throughput versus depth of analysis. Table 3 provides a comparative overview of key performance characteristics across major LC-MS platforms relevant to natural product research.
Table 3: Performance Comparison of LC-MS Platforms for Natural Product Analysis
| Platform Configuration | Separation Dimensions | Analysis Speed | Sensitivity | Structural Information | Ideal Application Context |
|---|---|---|---|---|---|
| UHPLC-Q-TOF | Chromatography + Mass | Fast to Moderate | High | MS and MS/MS with accurate mass | Untargeted metabolomics, metabolite profiling |
| UHPLC-Orbitrap | Chromatography + Mass | Moderate | High to Very High | MS and MS/MS with high resolution | Targeted and untargeted analysis requiring high mass accuracy |
| LC×LC-MS | 2D Chromatography + Mass | Slow | Moderate to High | MS and MS/MS | Extremely complex mixtures, isomer separation |
| UHPLC-TQ-MS | Chromatography + Mass | Very Fast | Very High | MRM transitions | High-sensitivity quantification of known compounds |
| LC-HRIM-MS | Chromatography + Ion Mobility + Mass | Very Fast | High | CCS values + MS and MS/MS | Isomer separation, structural characterization |
Detailed Experimental Protocol: LC-HR-MS3 for Natural Products
For laboratories seeking to implement advanced MS3 capabilities for natural product identification, the following experimental protocol adapted from published methodology provides a robust foundation [31]:
Sample Preparation:
- Prepare natural product standards by dissolving in 1:1 acetonitrile:dimethyl sulfoxide to 0.50 mg/mL
- Dilute in sample diluent (1:1:2 mixture of MeOH, ACN, and 5.0 mM ammonium formate in water, added with 0.05% formic acid) to 1.0 μg/mL for spectral library construction
- For biological samples (serum or urine), precipitate proteins with 3 volumes of acetonitrile relative to sample volume
- Centrifuge at 13,000 rpm for 10 minutes, collect supernatant, and dry under nitrogen flow at 37°C
- Reconstitute in appropriate volume of sample diluent for analysis
LC-HR-MS3 Method Parameters:
- Column: Accucore C18 (2.1 mm × 100 mm, 2.6 µm particle)
- Mobile Phase: A: 5 mM ammonium formate in water with 0.05% formic acid; B: MeOH:ACN (1:1) with 0.05% formic acid
- Gradient: Optimized for natural product separation (typically 10-90% B over 20-30 minutes)
- Column Temperature: 35°C
- Injection Volume: 5 µL
- Autosampler Temperature: 10°C
- Ionization: ESI positive mode
- Spray Voltage: 3.4 kV
- Capillary Temperature: 300°C
- Sheath Gas Flow Rate: 40
- Aux Gas Flow Rate: 10
- Aux Gas Heater Temperature: 375°C
Data-Dependent Acquisition Settings:
- Full-scan: m/z 100-1000 at 120K resolution
- MS2: Top 10 abundant precursor ions, isolation window 1.5 m/z, normalized HCD energy (20, 35, 45, 55, 65) auto-optimized, 30K resolution
- MS3: Top 3 MS2 product ions, isolation window 2 m/z, normalized HCD energy 30, 7.5K resolution
- Use inclusion list with mass-to-charge ratios of target analytes
- Employ internal mass calibration (e.g., EASY-IC) for high mass accuracy
Diagram 2: LC-HR-MS3 Data Acquisition Workflow. This multi-stage fragmentation process provides detailed structural information for confident compound identification.
Essential Research Reagents and Materials
Successful implementation of LC-MS methods for natural product biosynthesis validation requires specific reagents, standards, and materials. Table 4 outlines key components of the "research toolkit" for these applications.
Table 4: Essential Research Reagents and Materials for LC-MS Analysis of Natural Products
| Item Category | Specific Examples | Function/Purpose | Application Notes |
|---|---|---|---|
| Chromatography Columns | C18 reversed-phase (sub-2µm particles), HILIC, phenyl-hexyl | Compound separation based on chemical properties | Column chemistry should match analyte characteristics; sub-2µm particles for UHPLC |
| Mobile Phase Additives | Formic acid, ammonium formate, ammonium acetate | Modulate pH and improve ionization efficiency | Concentration typically 0.05-0.1%; volatile salts compatible with MS detection |
| Mass Calibration Standards | Sodium formate, Pierce LTQ Velos ESI Positive Ion Calibration Solution | Instrument mass accuracy calibration | Required before each analysis session for high mass accuracy |
| Natural Product Standards | Commercially available compounds (e.g., alkaloids, terpenoids, flavonoids) | Method development, quantification, identification | Critical for creating in-house spectral libraries |
| Sample Preparation Materials | Solid-phase extraction cartridges, protein precipitation reagents, filtration devices | Sample clean-up and concentration | Reduces matrix effects and instrument contamination |
| Data Analysis Software | Vendor-specific software, GNPS, XCMS, MZmine | Data processing, metabolite identification, statistical analysis | Open-source platforms facilitate reproducible research |
The ongoing evolution of LC-MS instrumentation continues to transform natural product research, providing increasingly powerful tools for elucidating complex biosynthetic pathways. The integration of UHPLC separation with high-resolution mass spectrometry and emerging technologies such as high-resolution ion mobility offers researchers unprecedented capabilities for comprehensive metabolite profiling and structural characterization. Each technological approach brings distinct advantages—UHPLC delivers exceptional chromatographic resolution, HRMS provides confident compound identification, and HRIM adds rapid separation based on molecular shape and size.
Looking forward, several trends are likely to shape the future of LC-MS in natural product biosynthesis validation. The continued development of integrated multi-dimensional separation platforms (LC×LC, LC-IM-MS) will address increasingly complex analytical challenges, particularly for isomeric compounds. Advances in computational tools and data processing algorithms will enhance our ability to extract biological insights from complex datasets, with packages like TARDIS demonstrating the value of open-source solutions for targeted data analysis [35]. Additionally, the growing emphasis on reproducibility and method transferability across laboratories will drive instrument development toward more robust and standardized platforms.
For researchers focused on validating natural product biosynthesis, the optimal instrumental configuration will ultimately depend on their specific analytical requirements—balancing needs for separation power, identification confidence, quantification sensitivity, and analytical throughput. By understanding the complementary strengths of available technologies and implementing appropriate experimental workflows, scientists can effectively address the complex challenges inherent in natural product research and drug development.
The validation of natural product biosynthesis relies heavily on advanced chromatographic techniques to separate and identify complex mixtures of bioactive compounds. Comprehensive two-dimensional liquid chromatography (2D-LC) and supercritical fluid chromatography (SFC) have emerged as powerful solutions that address the limitations of conventional one-dimensional separations. These techniques provide the resolution, sensitivity, and throughput necessary to unravel complex natural product matrices, thereby accelerating the discovery of novel therapeutic compounds through integrated LC-MS and bioassay research.
Within natural product research, a significant challenge lies in the efficient dereplication of known compounds to focus resources on novel chemical entities. Advanced chromatographic techniques coupled with mass spectrometry enable researchers to address this challenge by providing superior separation power and complementary orthogonality for complex sample analysis.
Technique Comparison: Comprehensive 2D-LC versus SFC
The selection of appropriate chromatographic techniques is pivotal for successful natural product analysis. The table below provides a systematic comparison of comprehensive 2D-LC and SFC based on critical performance parameters.
Table 1: Technical comparison of Comprehensive 2D-LC and SFC for natural product analysis
| Parameter | Comprehensive 2D-LC | Supercritical Fluid Chromatography (SFC) |
|---|---|---|
| Separation Mechanism | Two orthogonal separation mechanisms (e.g., RPLC x HILIC) [36] | Normal-phase separation using supercritical CO₂ with modifiers [36] |
| Peak Capacity | Very high (>1000) due to multiplicative effect of two dimensions [36] | High, with efficient separations for lipid classes and non-polar metabolites [36] |
| Analysis Speed | Typically longer run times due to sequential separations | Generally faster analysis than conventional LC |
| Loading Capacity | High, especially with semi-preparative first dimension [36] | Compatible with high sample loading for preparative applications |
| Ion Suppression Reduction | Significant reduction through separation of co-eluting compounds [36] | Moderate, dependent on mobile phase composition |
| MS Compatibility | Excellent with ESI-MS; may require flow splitting | Excellent with ESI and APCI interfaces |
| Ideal Application | Complex metabolite mixtures (e.g., fecal metabolome) [36] | Lipid class separations [36]; chiral separations |
Orthogonality and Peak Capacity
The principal advantage of comprehensive 2D-LC lies in its dramatically increased peak capacity, achieved through the combination of two independent separation mechanisms. Research demonstrates that offline 2D-LC methods more than doubled the number of unique database matches (from 1,513 to 3,414) compared to conventional one-dimensional separations when applied to the human fecal metabolome [36]. This enhanced separation power is particularly valuable for detecting low-abundance metabolites in complex natural product extracts.
Complementary Applications in Natural Products Research
SFC provides complementary capabilities, particularly for the separation of non-polar to moderately polar compounds. Its utility has been demonstrated in lipidomics, where SFC-based fractionation enabled identification of 404 lipids compared to 150 with a 1D RPLC-MS approach [36]. This makes SFC particularly suitable for analyzing certain classes of natural products, including terpenes, carotenoids, and fatty acid conjugates.
Experimental Protocols and Workflows
Offline 2D-LC-MS/MS for Metabolite Identification
A detailed experimental protocol for offline 2D-LC-MS/MS analysis of complex biological samples provides a robust framework for natural product research:
Sample Preparation: Fecal samples or natural product extracts are homogenized in chilled 1:1:1 methanol:acetonitrile:acetone solvent containing stable isotope-labeled internal standards. After centrifugation, supernatants are dried under nitrogen and reconstituted in water:methanol (9:1) [36].
First Dimension Separation: Semi-preparative RPLC is performed on a Waters Atlantis T3 OBD prep column (10 × 150 mm; 5 μm) at 55°C. Mobile phases consist of (A) water with 0.1% formic acid and (B) methanol with 0.025% formic acid. The gradient runs from 0% to 100% B over 20 minutes, maintained for 20 minutes, with a flow rate of 3 mL/min [36].
Fraction Collection: Eluent from the first dimension is collected into time-based fractions (e.g., 30-second intervals), which are subsequently concentrated before second-dimension analysis [36].
Second Dimension Separation: Concentrated fractions are analyzed using an orthogonal separation, typically HILIC or RPLC with different selectivity, coupled to a high-resolution tandem mass spectrometer [36].
Data Acquisition and Processing: MS/MS data are acquired using data-dependent acquisition methods. The resulting spectra are searched against commercial, public, and local spectral libraries, with annotations validated using retention time alignment and prediction [36].
Figure 1: Experimental workflow for offline 2D-LC-MS/MS analysis of complex natural product mixtures.
SFC-MS Workflow for Lipid and Natural Product Analysis
While detailed SFC protocols in the provided literature are limited, a generalized workflow for SFC-MS analysis includes:
Sample Preparation: Extraction optimized for target compound polarity, often similar to LC-MS protocols.
SFC Separation: Utilizes supercritical CO₂ as the primary mobile phase with methanol or ethanol modifiers containing additive compounds (e.g., ammonium acetate or formate) to enhance ionization and separation. Columns typically include packed silica or specialized bonded phases.
MS Analysis: Coupling to mass spectrometry via specialized interfaces that maintain back-pressure and compatibility with SFC mobile phases.
Automated LC-MS/MS Data Analysis Workflow
Recent advancements in data processing have led to the development of automated workflows for natural product annotation:
Collision Energy Optimization: The AutoAnnotatoR package incorporates a function to optimize collision energy (CE) values for each target ion, as CE significantly impacts fragment ion abundance and quality of structural information [37].
Diagnostic Ion Screening: Users can import tables of diagnostic fragment ions to screen for target components and identify potential novel compounds based on characteristic fragmentation patterns [37].
Database Matching: The workflow enables simultaneous matching of MS¹ and MS² spectral data against specialized databases, significantly improving identification accuracy compared to MS¹-only approaches [37].
Customization: The R-based package allows researchers to import specialized databases and diagnostic ion information tailored to their specific natural products of interest [37].
Figure 2: Automated data analysis workflow for natural product identification using LC-MS/MS data.
Research Reagent Solutions and Materials
Successful implementation of comprehensive 2D-LC and SFC methodologies requires specific reagents, materials, and instrumentation. The following table details essential components for establishing these analytical workflows.
Table 2: Essential research reagents and materials for comprehensive 2D-LC and SFC analyses
| Category | Specific Examples | Function/Application |
|---|---|---|
| Chromatography Columns | Atlantis T3 OBD prep column (10 × 150 mm; 5 μm) [36] | First dimension semi-preparative RPLC separation |
| Mass Spectrometers | Thermo Fisher Scientific LTQ XL [32] | Tandem MS capability for metabolite identification |
| Mobile Phase Additives | Formic acid (0.025-0.1%) [36] | Modifies pH and improves ionization efficiency |
| Extraction Solvents | Methanol:acetonitrile:acetone (1:1:1) [36] | Comprehensive metabolite extraction from biological matrices |
| Internal Standards | Stable isotope-labeled compounds (D₃-creatine, D₁₀-isoleucine, etc.) [36] | Quality control and quantification reference |
| Software Platforms | GNPS (Global Natural Products Social Molecular Networking) [32] | MS/MS spectral library searching and molecular networking |
| Data Analysis Tools | AutoAnnotatoR R package [37] | Automated compound annotation for botanical natural products |
Integration with Bioassay Research and Biosynthesis Validation
Advanced chromatographic techniques provide critical support for bioassay-guided fractionation and biosynthesis validation in natural product research.
Enhanced Dereplication Strategies
The improved separation power of comprehensive 2D-LC directly addresses a fundamental challenge in natural product discovery: efficient dereplication. By combining orthogonal separation mechanisms with high-resolution mass spectrometry, researchers can rapidly identify known compounds in complex mixtures, focusing resources on novel chemical entities [32]. This approach is particularly valuable when analyzing medicinal plant extracts, where multiple bioactive compounds may contribute to observed biological effects [32].
Correlation of Metabolic Features with Bioactivity
The increased metabolite identification capacity of comprehensive 2D-LC enables more robust correlation between chemical features and observed bioactivities. In a study of fecal metabolome changes following microbiota transplantation, the enhanced identification capability of 2D-LC revealed 72 additional significantly differentiated metabolites between pre- and post-transplant samples compared to conventional 1D-LC [36]. This improved descriptive power provides deeper insight into complex biological systems relevant to natural product research.
Validation of Biosynthetic Pathways
Comprehensive chromatographic techniques contribute significantly to the validation of natural product biosynthesis through:
Enhanced Detection of Biosynthetic Intermediates: The superior resolution of 2D-LC enables detection of low-abundance intermediates in biosynthetic pathways, facilitating pathway elucidation.
Isomer Separation: The orthogonal separation mechanisms in 2D-LC provide powerful capability to separate and identify stereoisomers that may be involved in biosynthetic pathways.
Comprehensive Metabolic Profiling: The expanded coverage of the metabolome enables more complete mapping of biosynthetic relationships between natural products within an organism.
Comprehensive 2D-LC and SFC represent significant advancements in chromatographic solutions for complex mixture analysis in natural product research. The dramatically improved peak capacity and orthogonality of 2D-LC enable identification of previously undetectable metabolites in complex natural product extracts, while SFC provides complementary capabilities for specific compound classes. When integrated with advanced MS detection and automated data analysis workflows, these techniques powerfully accelerate the discovery and validation of bioactive natural products. As these technologies continue to evolve with improvements in instrumentation, column chemistries, and data processing algorithms, their role in validating natural product biosynthesis and supporting drug development will undoubtedly expand.
Metabolite Profiling and Target Analysis in Engineered Biosynthetic Systems
Metabolite profiling has become an indispensable tool for validating and optimizing engineered biosynthetic systems in natural product research. By providing a comprehensive view of small molecule composition, these analytical approaches enable researchers to confirm successful pathway engineering, identify bottlenecks in biosynthetic flux, and discover new natural products with pharmaceutical potential. The integration of liquid chromatography-mass spectrometry (LC-MS) with robust bioassay methods creates a powerful framework for linking chemical structures to biological activity, thereby accelerating drug discovery and development. This guide objectively compares the performance of current metabolite profiling technologies and methodologies, providing experimental data and protocols that support their application in validating natural product biosynthesis.
Analytical Platform Comparison: LC-MS Technologies for Metabolite Profiling
Table 1: Comparison of LC-MS Instrumentation for Metabolite Profiling Applications
| Instrument Type | Mass Accuracy | Analysis Scope | Specialty | Optimal Application | Sample Throughput |
|---|---|---|---|---|---|
| Q-TOF-MS | < 5 ppm [38] | Quant./Quali. | High-speed mass scan | Untargeted metabolomics, unknown ID | Medium (10-100 samples) |
| Triple Quadrupole (QQQ) | Unit resolution | Quantitative | High sensitivity | Targeted analysis, biomarker validation | High (10-1000 samples) [39] |
| FT-MS/Orbitrap | < 2 ppm [12] | Qualitative | High mass resolution | Unknown identification, structural elucidation | Low (1-10 samples) [39] |
| Q-TOF with Ion Mobility | >20,000 FWHM [38] | Quant./Quali. with separation | Isomer separation | Complex mixtures, structural isomers | Medium |
| MALDI-TOF/TOF | Unit resolution | Qualitative | Imaging capability | Spatial distribution in tissues | Low to medium |
The selection of appropriate LC-MS instrumentation depends heavily on the research objectives. Untargeted metabolomics aims to monitor as many metabolites as possible in the entire metabolome to identify molecules that are up- or down-regulated, typically utilizing HPLC/MS or GC/MS instrumentation [38]. This approach is ideal for discovery-phase research, such as comparing wild-type versus transgenic systems or healthy versus diseased states. In contrast, targeted analysis focuses on predetermined analytes in complex biological matrices and requires rigorous method validation for specificity, linearity, precision, and accuracy [38]. Targeted approaches using triple quadrupole systems offer superior sensitivity and are better suited for validation studies where specific metabolic pathways are being engineered.
High-resolution accurate mass (HRAM) instruments like Q-TOF and Orbitrap systems have revolutionized untargeted metabolomics by enabling comprehensive metabolite detection without dependence on authentic standards [39]. The mass accuracy of less than 5 ppm provides confident elemental composition assignment, while MS/MS capabilities yield structural information for compound identification [38]. For engineered biosynthetic systems, this allows researchers to detect both expected products and unexpected side products or shunt metabolites that may arise from pathway manipulations.
Experimental Workflows: From Sample Preparation to Data Analysis
Sample Preparation and Extraction Protocols
Proper sample preparation is critical for obtaining reliable metabolomic data. An optimized protocol for microbial or plant cells involves quenching metabolic activity, extracting metabolites, and preparing samples for LC-MS analysis:
Cell Harvesting: Rapidly collect cells by filtration or centrifugation at specified growth phases [40]. For time-series experiments, sample multiple time points throughout the fermentation or growth cycle to capture metabolic dynamics.
Metabolite Extraction: Use a methanol-based extraction protocol for comprehensive metabolite coverage. For monocyte cells, researchers have developed an effective method involving ice-cold 80% ACS reagent-grade methanol, vortexing for 30 seconds, sonication in an ice bath for 1 minute, and subsequent vortexing for another 30 seconds [41]. Centrifuge at 16,000×g for 10 minutes at 4°C and collect the metabolite fraction (supernatant) for LC-MS analysis.
Sample Cleanup and Concentration: Employ solid-phase extraction (SPE) for fractionation when analyzing complex mixtures. C18 cartridges effectively separate metabolites based on polarity, allowing enrichment of target compound classes [32].
Quality Control: Prepare pooled quality control (QC) samples by combining aliquots from all samples to monitor instrument performance throughout the analysis [41]. Include extraction blanks as negative controls to identify contamination or background signals.
Integrated LC-MS/Bioassay Workflow for Natural Product Validation
The following diagram illustrates the comprehensive workflow for validating natural product biosynthesis using integrated LC-MS and bioassay approaches:
This integrated approach enables simultaneous assessment of metabolic changes and biological activity, providing comprehensive validation of engineered biosynthetic systems. The combination of chemical profiling and bioactivity data offers stronger evidence of successful pathway engineering than either method alone.
Data Analysis and Metabolite Identification Strategies
Advanced data analysis approaches transform raw LC-MS data into biological insights:
Multivariate Analysis: Use principal component analysis (PCA) and partial least squares-discriminant analysis (PLS-DA) to identify metabolites that differentiate experimental groups [32]. These methods reduce data dimensionality and highlight significant changes in metabolite abundance.
Metabolic Pathway Enrichment Analysis (MPEA): Apply pathway enrichment analysis to untargeted metabolomics data to identify significantly modulated pathways. This approach successfully revealed the pentose phosphate pathway, pantothenate and CoA biosynthesis, and ascorbate and aldarate metabolism as key modulated pathways in E. coli succinate production [40].
Dereplication Strategies: Employ the Global Natural Products Social Molecular Networking (GNPS) platform for efficient dereplication to limit compound rediscovery [32]. This web-based platform compares MS/MS fragmentation patterns of unknown analytes to reference spectra in curated libraries.
Molecular Networking: Create molecular networks using MS/MS fragmentation data to cluster related metabolites and identify structural analogs [32]. This approach visualizes chemical relationships within complex metabolite mixtures.
Case Studies: Applications in Engineered Biosynthetic Systems
Metabolic Engineering of Escherichia coli for Succinate Production
Metabolic pathway enrichment analysis of an E. coli succinate production bioprocess identified three significantly modulated pathways during the product formation phase: the pentose phosphate pathway, pantothenate and CoA biosynthesis, and ascorbate and aldarate metabolism [40]. The former two pathways align with previous engineering targets for improving succinate production, while ascorbate and aldarate metabolism represents a novel target not previously explored for strain improvement. This case demonstrates how untargeted metabolomics combined with pathway analysis can reveal both expected and unexpected engineering targets.
Proteomic Investigation of Secondary Metabolism (PrISM) in Bacillus Species
The PrISM approach uses proteomics to detect expressed nonribosomal peptide synthetase (NRPS) and polyketide synthase (PKS) gene clusters through identification of phosphopantetheinylated carrier proteins [12]. This method enabled discovery of new natural products from environmental isolates without prior genome sequencing information. When applied to 22 Bacillus isolates, PrISM identified five strains expressing high molecular weight NRPS/PKS proteins, leading to discovery of a new 7-residue lipopeptide [12]. This case highlights how protein-level detection of biosynthetic machinery can guide natural product discovery.
Optimization of Monocyte Metabolomics Using Improved Normalization Methods
In monocyte metabolomics, researchers evaluated over 40 different data normalization techniques to account for technical and biological variation [41]. The most efficient and consistent method was measurement of residual protein in the metabolite fraction, which was validated and optimized using a commercial kit. This careful attention to normalization enabled detection of broad and profound changes in monocyte metabolism in response to LPS stimulation, including alterations in amino acids, Krebs cycle metabolites, and previously unreported decreases in aspartate and β-alanine [41]. This case emphasizes the importance of proper normalization in obtaining reliable metabolomic data.
Table 2: Key Research Reagent Solutions for Metabolite Profiling Studies
| Reagent/Resource | Function | Application Example | Validation Requirements |
|---|---|---|---|
| CD14+ Microbeads | Immune cell isolation | Primary human monocyte isolation for immunometabolism studies [41] | Cell viability >95%, purity >90% |
| LPS (Lipopolysaccharide) | Immune stimulation | Monocyte activation model for studying metabolic reprogramming [41] | Endotoxin activity verification |
| CBR-5884 | Metabolic inhibitor | Investigating metabolic pathway contributions to cytokine production [41] | Dose-response validation |
| LC-MS Grade Solvents | Mobile phase preparation | Ensuring minimal background interference in LC-MS analysis [41] | Purity certification, batch testing |
| Authentic Standards | Metabolite identification | Confirming retention time and fragmentation patterns | Purity >95%, stability assessment |
| GNPS Platform | Metabolite database searching | Dereplication and annotation of natural products [32] | MS/MS spectrum matching algorithms |
| ELISA Kits | Cytokine quantification | Validating functional outcomes of metabolic changes [41] | Standard curve R² >0.99, spike recovery |
| Cell Viability Assays | Cytotoxicity assessment | Ensuring metabolic changes not due to cell death [41] | Linear range determination |
Pathway Visualization: Engineering Targets in Natural Product Biosynthesis
The following diagram illustrates key metabolic pathways frequently targeted in engineering natural product biosynthetic systems:
This pathway diagram highlights how central carbon metabolism intersects with specialized natural product biosynthesis. Engineering targets typically include precursor supply pathways (pentose phosphate pathway, TCA cycle, amino acid metabolism) and cofactor biosynthesis pathways (pantothenate and CoA biosynthesis) that support the enzymatic assembly lines for natural product formation.
The validation of engineered biosynthetic systems requires careful selection of metabolite profiling approaches matched to research objectives. Untargeted LC-MS methods using high-resolution instruments provide comprehensive discovery capabilities, while targeted approaches using triple quadrupole systems offer superior sensitivity for quantitative validation. Integration with bioassay data creates a powerful framework for linking chemical structures to biological function. As metabolomics technologies continue to advance, with improvements in mass accuracy, sensitivity, and computational tools, their application in optimizing engineered biosystems will become increasingly sophisticated and essential for natural product-based drug development.
Functional bioassays are indispensable procedures in chemical biology and drug discovery, allowing researchers to quantify the biological potency or effect of a substance by observing its impact on living cells, tissues, or whole organisms [42] [43]. In the context of validating natural product biosynthesis, these assays provide the critical link between the chemical structures identified via analytical techniques like Liquid Chromatography-Mass Spectrometry (LC-MS) and their resulting biological activity profiles [44] [7]. The primary objective of designing robust functional bioassays is to establish clear structure-activity relationships (SARs), which elucidate how specific chemical features or substructures in a compound correlate with specific biological responses [45]. This guide provides a comparative analysis of mainstream bioassay methodologies, supported by experimental data and protocols, to aid researchers in selecting and optimizing the most appropriate systems for their natural product research.
Comparative Analysis of Bioassay Platforms
The choice of bioassay platform depends heavily on the research question, desired throughput, and the nature of the biological activity being investigated. The table below summarizes the key characteristics of prevalent bioassay types used in linking chemical structure to biological function.
Table 1: Comparison of Functional Bioassay Platforms for SAR Studies
| Bioassay Type | Core Principle | Key Readouts | Typical Applications in SAR | Pros | Cons |
|---|---|---|---|---|---|
| Cell Viability & Cytotoxicity | Measures compound-induced loss of cellular structure or function [46]. | Metabolic activity (e.g., resazurin reduction, MTT/WST-1 conversion), membrane integrity (e.g., propidium iodide uptake, LDH release) [46]. | Initial screening for general toxicity; identifying cytotoxic natural products [43]. | Simple, high-throughput, low cost. | Low specificity; does not reveal mechanism of action [46]. |
| Reporter Gene Assays | Engineered cells produce a detectable reporter protein (e.g., luciferase) in response to a specific receptor or pathway activation [46]. | Luminescence or fluorescence intensity from the reporter gene product [46]. | Profiling compounds against specific biological pathways (e.g., nuclear receptor signaling); quantitative SAR [45] [46]. | High specificity and sensitivity; direct link to a molecular target/pathway; highly multiplexable. | Requires genetic engineering of cell lines; potential for artificial system artifacts. |
| Multiplexed Cytological Profiling (High-Content Screening) | Uses automated microscopy and image analysis to quantify multiple morphological features in stained cells [45]. | Measurements of hundreds of morphological descriptors (e.g., organelle shape, cytoskeleton organization, cell size) [45]. | Generating high-dimensional biological activity profiles for deep SAR analysis; identifying mechanism of action [45]. | Provides rich, multi-parametric data; captures complex phenotypes; can reveal unexpected activities. | Lower throughput; complex data analysis; expensive instrumentation. |
| Calcium Signaling Measurements | Monitors rapid changes in intracellular calcium levels using fluorescent dyes or photoproteins [46]. | Fluorescence or bioluminescence intensity fluctuations corresponding to calcium transients [46]. | Interrogating GPCR signaling and ion channel activity; real-time kinetic studies [46]. | Real-time, kinetic data; highly sensitive to rapid signaling events. | Can be susceptible to interference from non-specific calcium modulators. |
Establishing Structure-Activity Relationships (SARs)
Computational Mining of High-Dimensional Bioassay Data
Advanced computational methods are required to extract meaningful SARs from complex, high-dimensional bioassay data. Frequent Pattern Mining (FPM) and Association Rule Mining (ARM), originally developed for market-basket analysis, have been successfully adapted for this purpose [45]. These methods automatically identify combinations of chemical substructures (chemical attributes) that are statistically associated with specific patterns in biological activity profiles [45]. An SAR rule takes the form {Chemical Substructure A, Chemical Substructure B} → {Biological Activity Profile X}, allowing researchers to prioritize compound groups for further study based on their chemical features and predicted bioactivity [45].
Experimental Protocol: A Workflow for SAR Analysis
The following workflow outlines a standardized protocol for connecting chemical structure to biological activity using a combination of LC-MS, bioassays, and computational analysis, particularly in the context of natural products.
- Sample Preparation (Natural Product Extract): Begin with a crude natural product extract. Perform a preliminary fractionation using solid-phase extraction or preparative LC to reduce complexity. Key is to maintain a log of fractions for tracking [44].
- Chemical Profiling via LC-MS: Analyze each fraction using High-Resolution LC-MS. This step helps in dereplication—the early identification of known compounds to avoid rediscovery [7]. Acquire accurate mass and MS/MS fragmentation data for tentative structural annotation [44] [47].
- Bioassay Profiling: In parallel, subject all fractions to a panel of relevant functional bioassays (e.g., from Table 1). The goal is to generate a biological activity profile for each fraction [45].
- Data Integration and SAR Rule Mining: Integrate the chemical data (e.g., presence of specific molecular fragments or LC-MS features) and biological activity profiles. Apply FPM and ARM algorithms to this integrated dataset to identify significant associations between chemical features and biological effects [45].
- Validation and Isolation: Based on the SAR rules, prioritize fractions containing novel or structurally interesting compounds with desired bioactivity. Proceed with targeted isolation and purification of the active compound(s) using guided bioassay testing [7].
- Confirmation: Finally, confirm the structure of the pure active compound using NMR and other spectroscopic techniques, and validate its activity and proposed SAR in a dose-response manner [44].
Diagram 1: Integrated workflow for linking chemical structure to biological activity, combining LC-MS, bioassays, and computational SAR mining.
The Scientist's Toolkit: Essential Reagents and Materials
Successful execution of functional bioassays requires specific reagents and tools. The following table details key solutions for setting up a robust bioassay platform.
Table 2: Research Reagent Solutions for Functional Bioassays
| Reagent/Material | Function in Bioassay | Key Considerations |
|---|---|---|
| Viability/Cytotoxicity Kits (e.g., MTT, Resazurin) | Quantify overall metabolic activity of cells as a proxy for cell health and number [46]. | Choose assays compatible with your detection platform (colorimetric/fluorometric). Can be influenced by compounds that directly interact with metabolic enzymes [46]. |
| Engineered Cell Lines with Reporter Genes | Serve as sensors for specific pathway activation (e.g., estrogen receptor, Nrf2 antioxidant pathway) [46]. | Select cell lines with high relevance to your target biology (e.g., HepG2 for liver toxicity). Ensure stable expression of the reporter construct [45]. |
| Fluorescent Dyes & Probes | Enable visualization and quantification of specific cellular events (e.g., Ca²⁺ flux, mitochondrial potential, apoptosis) [45] [46]. | Check for spectral overlap if multiplexing. Validate that the natural product does not autofluoresce at the same wavelengths. |
| LC-MS Grade Solvents & Columns | Essential for the reproducible separation and analysis of natural products prior to or after bioassay [44] [47]. | Use high-purity solvents to minimize background noise. Column chemistry (C18, HILIC, etc.) should be selected based on the polarity of the target natural products [44]. |
| Design of Experiments (DoE) Software | A statistical approach for optimizing multiple bioassay parameters simultaneously, saving resources and time [48]. | Moves beyond inefficient "One Factor at a Time" approaches. Identifies complex interactions between factors (e.g., cell density, serum concentration, compound exposure time) [48]. |
Validation and Quality Control in Bioassay Development
Ensuring that a bioassay is reproducible, reliable, and biologically relevant is paramount [42]. Key sources of variability must be identified and controlled. These typically include analyst-to-analyst variation, day-to-day variation, and critical reagent lot variation [49]. Statistical approaches, such as Variance Component Analysis (VCA), are recommended to quantify the contribution of each source to the total variability [49]. This involves conducting a variability study where the bioassay is performed by multiple analysts over several days, with multiple replicates. The data, often log-transformed for potency assays, is then analyzed to estimate the variance components, helping to focus improvement efforts on the largest sources of error [49].
Diagram 2: A systematic approach to managing bioassay variability using Variance Component Analysis.
The strategic design of functional bioassays is a cornerstone of modern efforts to link the chemical structure of natural products to their biological activity. As detailed in this guide, no single bioassay platform is superior in all aspects; the choice hinges on the specific goals of the SAR study, balancing throughput, specificity, and data richness. The convergence of advanced analytical techniques like LC-MS for structural elucidation, a diverse panel of biologically relevant bioassays, and robust computational methods for data mining creates a powerful framework for natural product-based drug discovery. By adhering to rigorous validation practices and leveraging integrated experimental workflows, researchers can effectively navigate the complex chemical space of natural products to identify novel therapeutic leads with validated mechanisms of action.
The modernization of Traditional Medicine (TM), particularly Traditional Chinese Medicine (TCM), hinges on the ability to scientifically validate the efficacy, safety, and mechanistic pathways of complex natural products [50] [51]. For researchers and drug development professionals, this presents a unique challenge: how to systematically analyze multi-component therapies that operate via multi-target, multi-pathway mechanisms, a stark contrast to the conventional "one-target, one-drug" paradigm [51]. This case study objectively compares three predominant analytical frameworks—Chinmedomics, Network Pharmacology, and Conventional Bioassay-Guided Fractionation—in their application to TM analysis and pathway characterization. The evaluation is framed within a critical thesis on validating natural product biosynthesis, where Liquid Chromatography-Mass Spectrometry (LC-MS) provides the analytical backbone and bioassays deliver the functional context [32] [52].
Comparative Analysis of Analytical Approaches
The table below summarizes the core characteristics, strengths, and limitations of the three primary research strategies used in TM analysis.
Table 1: Comparison of Analytical Approaches in Traditional Medicine Research
| Feature | Chinmedomics | Network Pharmacology | Bioassay-Guided Fractionation |
|---|---|---|---|
| Core Philosophy | Holistic evaluation by correlating in vivo absorbed components with biomarker reversal [53]. | "Network-target, multiple-component-therapeutics" mode based on database mining [51]. | Reductionist approach to isolate active compounds through iterative testing [52]. |
| Key Methodology | Integrates metabolomics, serum pharmacochemistry, and bioinformatics [53]. | Constructs "compound-protein/gene-disease" networks using computational algorithms and databases [51]. | Step-wise separation (e.g., extraction, fractionation) guided by bioactivity results [32] [52]. |
| Role of LC-MS | Central. Used for metabolite profiling and identifying absorbed herbal components from serum [53] [54]. | Supplemental. Often used for validation; primary reliance is on database predictions [51]. | Central. Coupled with bioassays for the dereplication and identification of active compounds [32]. |
| Role of Bioassay | Confirms efficacy and links metabolic biomarker changes to therapeutic effect [53]. | Limited; used for experimental validation of computationally predicted targets [51]. | The primary driver of the isolation process [52]. |
| Pathway Characterization | Strong. Identifies actual in vivo metabolic pathways and connects them to drug action [53]. | Predictive. Infers pathways and mechanisms from network models and prior knowledge [51]. | Indirect. Mechanism is often elucidated after a single active compound is isolated [52]. |
| Throughput | Medium to High (automated omics platforms) [53]. | Very High (in silico) [51]. | Low (iterative and labor-intensive) [52]. |
| Key Advantage | Directly reveals the in vivo pharmacodynamic material basis and its mechanism under efficacious conditions [53]. | Rapid, cost-effective for generating testable hypotheses on a large scale [51]. | Directly links a specific compound to a measurable biological activity [52]. |
| Primary Limitation | Complex data integration requires sophisticated bioinformatics [53]. | Predictive nature; results require rigorous experimental validation [51]. | High risk of missing synergistic effects; can be slow [52]. |
Experimental Protocols for Key Methodologies
Chinmedomics Workflow
The Chinmedomics approach is an integrated, systems-level strategy for evaluating TM efficacy and identifying active components [53].
- Disease/Syndrome Model Establishment and Grouping: Animal or human subjects are classified into groups: control, disease/syndrome model, and TM-treated groups. The model is validated through behavioral, biochemical, and histopathological analyses [53].
- Sample Collection: Bio-samples (e.g., blood (serum/plasma), urine, tissues) are collected from all groups. Serum is critical for two parallel analyses [53] [54].
- Metabolomic Profiling (From Model Bio-samples):
- Analysis: Samples from control and model groups are analyzed using LC-MS/MS or GC-MS to profile endogenous metabolites [53] [54].
- Data Processing: Multivariate statistical analysis (e.g., PCA, PLS-DA) is applied to identify differentially expressed metabolites serving as potential biomarkers for the disease/syndrome [53].
- Biomarker Identification: The chemical structures of these biomarker metabolites are identified by matching MS/MS spectra and retention times against standard databases like HMDB or Metlin [53].
- Serum Pharmacochemical Analysis (From Treated Group Serum):
- Analysis: Serum from the TM-treated group is analyzed using LC-MS/MS to characterize the compounds absorbed into the bloodstream (prototypes and metabolites) [53].
- Compound Identification: Absorbed herbal components are identified by comparing their MS data with reference compounds and databases [53].
- Efficacy Evaluation and Correlation Analysis:
- The TM's efficacy is evaluated by monitoring the reversal of the identified disease biomarkers towards the normal state after treatment [53].
- A correlation network model is constructed using bioinformatics tools (e.g., WGCNA, Cytoscape) to link the absorbed herbal components with the recalled biomarkers. Highly correlated components are considered potential active ingredients [53].
- Validation: The predicted active ingredients and their targets are validated through in vitro or in vivo biological experiments [53].
LC-MS/MS and Dereplication for Natural Product Discovery
This protocol is central for identifying known compounds early in the discovery process, avoiding re-isolation [32].
- Sample Preparation:
- LC-MS/MS Analysis:
- The extract or fraction is analyzed using LC coupled to a tandem mass spectrometer (e.g., Q-TOF, Orbitrap, LTQ XL) [32] [54].
- Chromatography: UPLC or HPLC is used to separate compounds.
- Mass Spectrometry: The mass spectrometer acquires high-resolution MS1 data (for molecular weight) and MS/MS fragmentation data (for structural information) in data-dependent acquisition mode [32].
- Data Processing and Dereplication:
- MS/MS data files are converted to open formats (e.g., .mzXML).
- Data is uploaded to the Global Natural Products Social Molecular Networking (GNPS) platform or similar software [32].
- Molecular Networking: GNPS clusters MS/MS spectra based on similarity, visually grouping related compounds and revealing novel analogs [32].
- Library Search: The MS/MS spectra of unknown analytes are matched against reference spectra in embedded libraries (e.g., GNPS, MassBank) to putatively identify known compounds [32].
- Target Isolation: Efforts are then focused on isolating compounds that are either unknown or of high interest based on their presence in an active fraction and their structural novelty [32].
Bioassay-Guided Fractionation
This classical approach iteratively separates a complex mixture to pinpoint active constituents [52].
- Extraction and Initial Bioassay: The starting material (e.g., dried herb) is extracted. The crude extract is tested in a relevant bioassay (e.g., antioxidant DPPH assay, antimicrobial, cytotoxic) to establish a baseline activity [32] [52].
- Primary Fractionation: The active crude extract is subjected to a coarse separation step, typically using liquid-liquid partitioning or vacuum liquid chromatography over a normal-phase or C18 solid phase. This yields several primary fractions [32] [52].
- Bioassay and Iteration: All primary fractions are tested in the same bioassay. The most active fraction is selected for further separation using techniques like preparative HPLC or flash chromatography, yielding sub-fractions. This process of separation, bioassay, and selection is repeated until a pure, active compound is obtained [52].
- Structure Elucidation: The structure of the purified active compound is determined using spectroscopic methods, including NMR and high-resolution MS [52].
Visualization of Research Workflows
Chinmedomics Efficacy & Pathway Analysis
LC-MS/MS Dereplication Workflow
The Scientist's Toolkit: Essential Research Reagents and Materials
The following table details key reagents, materials, and software solutions essential for conducting the experiments described in this case study.
Table 2: Essential Research Reagents and Solutions for TM Analysis
| Category | Item | Function/Application | Example Use Case |
|---|---|---|---|
| Chromatography & Separation | C18 Solid-Phase Extraction (SPE) Cartridges [32] | Pre-fractionation of complex crude extracts to reduce complexity for LC-MS analysis. | Initial clean-up and fractionation of plant extracts in bioassay-guided fractionation [32]. |
| UPLC/HPLC Columns (e.g., C18) [54] | High-resolution separation of complex mixtures prior to mass spectrometry detection. | Core component of any LC-MS system for analyzing metabolites or herbal components [53] [54]. | |
| Mass Spectrometry | High-Resolution Mass Spectrometer (e.g., Q-TOF, Orbitrap, LTQ XL) [32] [54] | Provides accurate mass measurement (MS1) and structural fragmentation data (MS/MS) for compound identification. | Essential for untargeted metabolomics and serum pharmacochemistry in Chinmedomics [53] [32]. |
| Bioassay Kits & Reagents | DPPH (2,2-Diphenyl-1-picrylhydrazyl) [32] | A stable free radical used to screen for antioxidant activity in extracts and fractions. | Initial bioactivity screening in natural product research [32]. |
| Cell Viability/Cytotoxicity Assay Kits (e.g., MTT, WST-8) [52] | Measure cell proliferation or death to assess cytotoxic potential of samples. | Bioassay for anticancer drug discovery from natural products [52]. | |
| Bioinformatics & Databases | Global Natural Products Social Molecular Networking (GNPS) [32] | Web-based platform for MS/MS spectral library matching and creating molecular networks. | Dereplication and analog identification in LC-MS/MS data [32]. |
| Human Metabolome Database (HMDB) / Metlin [53] | Curated databases of metabolite spectra and information for biomarker identification. | Identifying endogenous biomarkers in metabolomics studies [53]. | |
| Cytoscape [53] [51] | Open-source software for visualizing complex molecular interaction networks. | Visualizing "compound-target-pathway" networks in network pharmacology and Chinmedomics [53] [51]. | |
| Sample Preparation | Protein Lysis Buffers (e.g., RIPA, Urea) with Protease Inhibitors [54] | Lyse cells/tissues and solubilize proteins while preventing degradation for proteomics. | Protein extraction from animal tissues in TCM mechanism studies [54]. |
| Bicinchoninic Acid (BCA) Assay Kit [54] | Colorimetric method for quantifying total protein concentration in a sample. | Determining protein content before proteomic analysis [54]. |
The integration of LC-MS technologies and robust bioassay research provides an powerful foundation for validating natural product biosynthesis and action in traditional medicine. While conventional bioassay-guided fractionation offers direct evidence for activity, and network pharmacology provides high-throughput predictive power, the Chinmedomics framework represents a particularly advanced paradigm. It successfully bridges TCM's holistic principles with modern analytical science by directly correlating the in vivo absorbed chemical profile with the reversal of disease-specific metabolic pathways. For researchers aiming to fully characterize the complex pathways and active components in traditional medicines, a synergistic strategy that leverages the strengths of all three approaches—using network pharmacology for hypothesis generation, Chinmedomics for in vivo validation and efficacy correlation, and targeted bioassays for functional confirmation—will be most effective in advancing these natural resources into evidence-based therapies.
Overcoming Analytical and Biological Challenges in Biosynthesis
The validation of natural product biosynthesis through LC-MS and bioassay research represents a cornerstone of modern drug discovery. However, this process is fraught with analytical challenges, primarily stemming from the profound complexity of biological matrices. These natural extracts contain hundreds to thousands of constituents with diverse physicochemical properties and wide concentration ranges, which can interfere with the accurate detection, quantification, and biological assessment of target compounds. Matrix effects—where co-eluting compounds suppress or enhance ionization—significantly compromise assay sensitivity, reproducibility, and the reliability of metabolic pathway validation [55] [56]. This guide objectively compares current analytical strategies and technological solutions designed to manage sample complexity, providing researchers with validated experimental protocols and data-driven comparisons to advance natural product research.
Fundamental Isolation Challenges in Natural Product Analysis
The initial stages of natural product research involve extracting compounds from complex biological materials, which presents several specific challenges that can obstruct subsequent analysis.
- Chemical Diversity and Structural Similarity: Natural extracts contain a vast array of compounds, including polyphenols, flavonoids, alkaloids, and terpenoids, often appearing as closely related structural analogues with nearly identical physicochemical properties. This complexity makes baseline separation difficult even with high-resolution chromatography [55] [57].
- Concentration Disparities: Bioactive compounds of interest frequently occur in minute quantities (trace levels) alongside highly abundant matrix components, necessitating extensive sample processing to achieve detectable levels for structure elucidation and bioactivity testing [57].
- Dynamic Range and Detection Limits: The extensive concentration range in crude extracts often exceeds the dynamic range of analytical detectors, requiring dilution or preconcentration steps that can introduce analytical bias or result in the loss of critical metabolites [57].
Matrix Effects: Impact on LC-MS Bioanalysis
Matrix effects represent a critical challenge in LC-MS analysis, particularly when investigating natural products in complex biological samples. These effects occur when co-eluting compounds from the sample matrix alter the ionization efficiency of target analytes in the mass spectrometer source [56].
Mechanisms and Consequences
Matrix components can cause ion suppression or, less commonly, ion enhancement, leading to compromised data quality. The consequences include:
- Reduced assay sensitivity and higher limits of detection
- Poor reproducibility and quantitative inaccuracy
- Masked or distorted chromatographic peaks
- False negatives/positives in metabolic profiling [56]
Biological matrices introduce numerous interfering components, including phospholipids, salts, proteins, and metabolic by-products. The extent of interference varies significantly between sample types (e.g., plant vs. microbial extracts) and preparation methods [56].
Table 1: Common Matrix Components and Their Effects in LC-MS Analysis
| Matrix Component | Source | Impact on LC-MS Analysis |
|---|---|---|
| Phospholipids | Cellular membranes | Major cause of ion suppression in ESI |
| Alkaloidal Compounds | Plant tissues | Can co-elute and interfere with target analytes |
| Proteins | Incomplete precipitation | Column fouling and signal instability |
| Carbohydrates | Plant and microbial extracts | Can affect chromatographic separation |
| Endogenous Metabolites | All biological systems | Complex interference patterns |
Comparative Analysis of Sample Preparation Techniques
Effective sample preparation is paramount for reducing matrix effects and simplifying complex mixtures. The choice of technique significantly influences downstream analytical outcomes, and researchers must select methods based on their specific sample composition and analytical goals.
Table 2: Comparison of Sample Preparation Methods for Complex Natural Product Matrices
| Method | Mechanism | Best For | Limitations | Matrix Effect Reduction |
|---|---|---|---|---|
| Protein Precipitation (PPT) | Protein denaturation with organic solvents | High-throughput workflows, simple samples | Limited selectivity, high matrix background | Low to Moderate |
| Solid-Phase Extraction (SPE) | Selective partitioning using functionalized sorbents | Pre-concentration, class-specific isolation | Method development time, cost | Moderate to High |
| Liquid-Liquid Extraction (LLE) | Differential solubility in immiscible solvents | Non-polar metabolites, large sample volumes | Emulsion formation, solvent volumes | Moderate |
| Online SPE | Automated clean-up coupled directly to LC-MS | Repetitive analysis, labile compounds | Initial setup cost, column compatibility | High |
Experimental Protocol for SPE Method Development:
- Conditioning: Sequentially flush the SPE cartridge (e.g., C18 for medium-polarity compounds) with 2-3 column volumes of methanol followed by equilibrium with water or initial mobile phase.
- Loading: Apply the sample (previously centrifuged and diluted if necessary) slowly to maximize interaction with the sorbent.
- Washing: Remove weakly retained interferents with 2-3 column volumes of water or 5-20% methanol/water.
- Elution: Collect analytes with 2-3 column volumes of strong solvent (e.g., acetonitrile, methanol, or with acid/base modifiers).
- Reconstitution: Evaporate the eluent under nitrogen or vacuum and reconstitute in initial mobile phase for LC-MS analysis [56] [58].
Advanced Chromatographic Solutions for Matrix Management
Chromatographic separation represents the first line of defense against matrix effects in LC-MS workflows. Modern stationary phases and multidimensional approaches offer significant improvements in resolving power for complex natural product mixtures.
Ultra-High-Performance Liquid Chromatography (UHPLC)
The implementation of UHPLC with sub-2μm particle columns provides superior resolution and faster analysis compared to conventional HPLC. The reduced particle size increases peak capacity, allowing better separation of complex metabolite mixtures and reducing the number of co-eluting compounds that cause matrix effects [55] [57].
Orthogonal Separation Techniques
- Reversed-Phase (RPLC): The most common approach, using C18 or pentafluorophenyl columns with water-organic mobile phases, effectively separates metabolites by hydrophobicity. It is ideal for medium to non-polar compounds but struggles with highly polar analytes [55].
- Hydrophilic Interaction Liquid Chromatography (HILIC): This technique utilizes polar stationary phases with organic-rich mobile phases, effectively retaining and separating polar compounds that elute too quickly in RPLC. It is particularly valuable for phenolic acids, catechins, and anthocyanins [55].
- Two-Dimensional LC (HILIC × RPLC): Combining these orthogonal separation mechanisms in a comprehensive 2D-LC setup significantly increases peak capacity and resolution, providing enhanced separation of complex natural product extracts [55].
LC-MS Multi-Dimensional Separation Workflow
Quantitative Method Validation in Complex Matrices
Robust quantitative analysis requires thorough method validation to ensure reliability despite matrix effects. The study by Yilmaz (2020) exemplifies a comprehensive approach, validating an LC-MS/MS method for 53 phytochemicals in 33 medicinal plants [58].
Key Validation Parameters and Experimental Data
Experimental Protocol for Method Validation:
- Linearity: Prepare calibration curves using matrix-matched standards (in blank matrix extract) across expected concentration range. Acceptable correlation coefficients (R²) should exceed 0.995 [58].
- Accuracy and Precision: Assess via spike-recovery experiments at low, medium, and high concentrations. Calculate intra-day and inter-day precision (%RSD), with values <15% generally considered acceptable [58].
- Limits of Detection and Quantification (LOD/LOQ): Determine via serial dilution of spiked matrices until signal-to-noise ratios of 3:1 (LOD) and 10:1 (LOQ) are achieved [58].
- Matrix Effects: Evaluate by comparing the analyte response in post-extraction spiked matrix to neat solution standards. Signal suppression/enhancement >25% typically requires mitigation strategies [58].
Table 3: Representative Validation Data for Selected Phytochemicals [58]
| Compound | Linearity (R²) | LOD (ng/mL) | LOQ (ng/mL) | Matrix Effect (%) | Recovery (%) |
|---|---|---|---|---|---|
| Chlorogenic Acid | 0.999 | 0.15 | 0.50 | -12.3 | 95.2 |
| Rutin | 0.998 | 0.25 | 0.83 | -8.7 | 97.8 |
| Quercetin | 0.997 | 0.32 | 1.07 | -15.2 | 92.4 |
| Kaempferol | 0.998 | 0.28 | 0.93 | -10.5 | 94.7 |
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful management of sample complexity requires specific reagents and materials designed to address matrix-related challenges.
Table 4: Essential Research Reagents for Managing Matrix Effects
| Reagent/Material | Function | Application Example |
|---|---|---|
| Isotopically Labeled Internal Standards | Correct for analyte loss and matrix effects during sample preparation and analysis | Compensation for variable recovery in complex plant extracts [56] [58] |
| Bio-Based Solvents | Environmentally friendly alternatives for extraction following green chemistry principles | Reduced toxicity while maintaining extraction efficiency [55] |
| HILIC Stationary Phases | Retention and separation of highly polar compounds | Analysis of phenolic acids and flavonoids that poorly retain in RPLC [55] |
| Matrix-Matched Calibrators | Standard solutions prepared in blank matrix to mimic sample composition | Accurate quantification compensating for inherent matrix effects [58] |
| Silica Gel Sorbents | Classical normal-phase separation of medium to non-polar compounds | Pre-fractionation of crude extracts prior to detailed analysis [57] |
Integrated Workflows for Enhanced Natural Product Discovery
Modern approaches combine multiple strategies to address sample complexity throughout the discovery pipeline. The MATRIX platform utilizes miniaturized 24-well microbioreactors with diverse media compositions to activate silent biosynthetic gene clusters, followed by UPLC-QTOF-MS/MS analysis and GNPS molecular networking for efficient metabolite annotation [59]. Similarly, dereplication strategies employing LC-MS/MS with database searching prevent redundant compound isolation, saving significant resources in natural product discovery [32] [60].
Integrated Natural Product Analysis Workflow
Addressing sample complexity and matrix effects remains a formidable challenge in validating natural product biosynthesis. The comparative data presented demonstrates that while no single technique completely eliminates matrix interference, integrated approaches combining selective sample preparation, advanced chromatographic separations, and appropriate internal standardization deliver the most reliable results. As natural product research increasingly focuses on validating biosynthetic pathways, the systematic implementation of these rigorously validated methods will be essential for generating reproducible, biologically relevant data. Future advancements will likely focus on more intelligent online cleanup technologies, improved orthogonal separation systems, and bioinformatic tools that can computationally compensate for residual matrix effects, further accelerating natural product-based drug discovery.
Combinatorial biosynthesis and pathway engineering represent powerful synthetic biology strategies to optimize the production of valuable natural products or create novel compounds. By recombining, editing, and optimizing the genetic blueprint of biosynthetic pathways in microbial hosts, researchers can overcome the limitations of natural production systems. These approaches are fundamentally changing natural product research and development, providing a engineered, reliable, and sustainable alternative to traditional extraction from native sources. Within the context of validating natural product biosynthesis, techniques like LC-MS/MS analysis and bioassay-guided fractionation serve as critical tools for confirming successful pathway engineering and identifying the resulting bioactive molecules [32] [61].
Core Strategies for Pathway Optimization
The optimization of biosynthetic pathways leverages several distinct but complementary methodologies, each with its own applications and outcomes.
Table 1: Comparison of Key Pathway Optimization Strategies
| Strategy | Core Principle | Key Application | Representative Outcome |
|---|---|---|---|
| Combinatorial Biosynthesis [62] [63] | Recombining biosynthetic genes from different organisms to generate libraries of hybrid natural products. | Rapidly expanding structural diversity to create "unnatural" natural products. | Generation of 61 different analogs of 6-deoxyerythronolide B [62]. |
| Combinatorial Engineering [64] | Systematically testing numerous enzyme variant combinations within a pathway to find optimal configurations. | Optimizing the production levels of a specific target compound in a heterologous host. | 6-fold increase in betaxanthin production in yeast [64]. |
| Evolution-Guided Optimization [65] | Coupling product formation to cell survival and using mutagenesis to evolve high-producing strains. | Achieving high titers of a target compound without requiring prior mechanistic knowledge. | 36-fold and 22-fold increase in naringenin and glucaric acid production, respectively [65]. |
| De Novo Pathway Design [66] | Designing novel metabolic pathways using a retrosynthetic approach, combining enzymes from diverse species. | Producing both natural and non-natural compounds for which no natural pathway is known or available. | Microbial production of artemisinic acid, a precursor to the anti-malarial drug artemisinin [66]. |
Combinatorial Biosynthesis and Engineering
Combinatorial biosynthesis involves manipulating biosynthetic pathways to produce new or altered chemical structures by harnessing nature's enzymatic machinery [62]. A powerful application is domain swapping in large enzymatic complexes like polyketide synthases (PKS). For instance, swapping the starter unit acyl carrier protein transacylase (SAT) domain between different fungal PKSs has led to the production of novel polyketides with altered starter units and chain lengths [63]. In a more comprehensive approach, combinatorial engineering was used to optimize the betalain biosynthesis pathway in yeast. By testing a dozen variants of two key enzymes, researchers identified optimal combinations that resulted in a six-fold higher production of betaxanthins and achieved a betanin titer of 30.8 mg/L [64].
Evolution-Guided Optimization
This strategy uses a "toggled selection" scheme, where a biosensor is engineered to make cell survival dependent on the production of the target molecule. When combined with targeted genome-wide mutagenesis, this setup allows for the evolution of high-producing strains. This method addresses the screening bottleneck by enabling the evaluation of nearly a billion pathway variants simultaneously, enriching for the rare cells with superior production phenotypes [65].
Rational De Novo Pathway Design
Moving beyond the manipulation of existing pathways, de novo design uses a retro-biosynthetic approach to specify entirely new metabolic routes in microbial hosts. This is analogous to the retrosynthesis practiced by organic chemists and leverages a growing toolkit of well-characterized biological "Parts" – genes encoding enzymes with specific functions [66]. A landmark achievement in this area is the engineering of yeast to produce artemisinic acid, providing a scalable and sustainable source of this crucial anti-malarial drug precursor [66].
Analytical Validation: LC-MS and Bioassay Integration
The success of any pathway engineering effort must be validated through rigorous analytical techniques that confirm compound identity and biological activity.
LC-MS/MS for Metabolite Identification and Dereplication
Liquid chromatography coupled with tandem mass spectrometry (LC-MS/MS) is a cornerstone for validating engineered biosynthesis. Its primary roles include:
- Identifying and Confirming Metabolites: LC-MS/MS provides high sensitivity and specificity for detecting the products of engineered pathways. The use of tandem MS (MS/MS) allows for increased confidence in metabolite identification by comparing fragmentation patterns of unknown analytes to those of authentic standards or library spectra [32].
- Dereplication: A key step in natural product research, dereplication involves the early identification of known compounds to focus resources on novel discoveries. Platforms like Global Natural Products Social Molecular Networking (GNPS) use LC-MS/MS data to automatically compare unknown metabolites against vast MS/MS spectral libraries, preventing the rediscovery of known compounds [32].
Table 2: Key Experimental Protocols in Pathway Validation
| Technique | Protocol Summary | Key Application in Pathway Optimization |
|---|---|---|
| LC-MS/MS Analysis [32] | 1. Separate metabolites via Liquid Chromatography.2. Ionize and analyze masses in the first MS stage.3. Select precursor ions for fragmentation.4. Analyze fragment ions in the second MS stage.5. Compare MS/MS spectra to databases (e.g., GNPS). | Confirming the identity of a compound produced by an engineered pathway and ensuring it is novel. |
| Bioassay-Guided Fractionation [61] | 1. Screen crude extract for bioactivity.2. Fractionate the active extract (e.g., using HPLC).3. Test fractions for the same bioactivity.4. Iteratively fractionate the active fraction until a pure active compound is isolated.5. Identify the pure active compound (e.g., via NMR, LC-MS). | Isating and identifying the specific bioactive molecule from a library of compounds generated by combinatorial biosynthesis. |
| Fluorescence Polarization (FP) Screening [61] | 1. Incubate a protein target with a fluorescently-labeled peptide ligand and a test extract/compound.2. Measure fluorescence polarization.3. Active compounds that displace the labeled ligand cause a decrease in polarization. | Ultra-high-throughput screening (uHTS) of natural product libraries or engineered strain libraries for specific biological activities (e.g., inhibition of protein-protein interactions). |
Bioassay Integration for Functional Validation
Bioassays are essential for linking the chemical structures produced by engineered pathways to a biological function. They are used both for initial detection and for guiding the isolation of active compounds [20].
- High-Throughput Bioactivity Screening: Modern screening employs ultra-high-throughput methods to test hundreds of thousands of samples. For example, a fluorescence polarization (FP) screen of nearly 150,000 natural product extracts was conducted against six anti-apoptotic Bcl-2 family proteins to discover new anti-cancer agents [61].
- Bioassay-Guided Fractionation: This is a critical workflow where a biologically active crude extract is progressively fractionated, and each fraction is tested for activity. This iterative process ensures that the isolation efforts are focused solely on the compound(s) responsible for the desired bioactivity [61].
Essential Research Reagents and Tools
The following toolkit is fundamental for research in combinatorial biosynthesis and pathway engineering.
Table 3: The Scientist's Toolkit: Key Research Reagents and Solutions
| Reagent / Tool | Function / Application | Example Use Case |
|---|---|---|
| Heterologous Hosts (e.g., S. cerevisiae, E. coli) [64] [66] | Engineered microbial chassis for expressing heterologous biosynthetic pathways. | Production of betalains in yeast [64] and amorphadiene in E. coli [66]. |
| Biosensors [65] | Genetic circuits that couple intracellular metabolite concentration to a reporter output (e.g., fluorescence, cell survival). | Evolution-guided optimization of naringenin production [65]. |
| Global Natural Products Social Molecular Networking (GNPS) [32] | An online platform for MS/MS spectral library matching and molecular networking. | Dereplication and identification of metabolites in engineered plant extracts [32]. |
| Enzyme Parts (e.g., SAT, PT, TE domains) [63] | Well-characterized catalytic domains that can be swapped between megasynthetases. | Engineering fungal PKS to produce novel polyketides [63]. |
Visualizing Workflows and Pathways
The following diagrams illustrate the logical relationships and workflows central to optimizing and validating biosynthetic pathways.
Bioassay-Guided Fractionation and Validation Workflow
Combinatorial Biosynthesis Strategy
Combinatorial biosynthesis and pathway engineering have moved from conceptual frameworks to practical and powerful tools for natural product research and development. By integrating strategies like combinatorial enzyme engineering, evolution-guided optimization, and rational de novo design, scientists can dramatically enhance the production of valuable compounds and generate entirely new molecular entities. The continued success of this field relies on the tight integration of these engineering strategies with robust analytical validation through LC-MS/MS and bioassay, creating a virtuous cycle of design, construction, testing, and discovery. This integrated approach promises to unlock a new era of natural product-based solutions for medicine, agriculture, and industry.
The discovery and development of natural product-based therapeutics face a critical bottleneck: securing a reliable and adequate supply of bioactive compounds. Many promising molecules are produced in minuscule quantities by their native hosts—whether plants, fungi, or bacteria—or are derived from organisms that are difficult to cultivate or ethically problematic to harvest. This supply chain limitation severely hampers further research, pre-clinical testing, and clinical development. Within the context of validating natural product biosynthesis through LC-MS and bioassay research, two powerful biotechnological approaches have emerged as solutions: precursor supplementation and heterologous expression. This guide provides an objective comparison of these strategies, supported by experimental data and detailed methodologies, to help researchers select the optimal approach for their specific natural product targets.
Heterologous Expression: Establishing New Production Platforms
Heterologous expression involves transferring the entire biosynthetic machinery for a natural product—typically in the form of a biosynthetic gene cluster (BGC)—from the native producer into a well-characterized host organism suitable for laboratory manipulation and scalable fermentation [67]. This strategy effectively decouples compound production from the original source organism, creating a more reliable and controllable production platform.
Table 1: Heterologous Expression Platforms for Natural Product Production
| Host Organism | Key Modifications/Features | DNA Transfer Method | BGC Types Successfully Expressed | Reported Titers |
|---|---|---|---|---|
| Streptomyces coelicolor A3(2)-2023 | Deletion of 4 endogenous BGCs; multiple RMCE sites [68] | Conjugation from E. coli | Type II PKS (griseorhodin), Xiamenmycin BGC [68] | Increasing xiamenmycin yield with copy number (2-4 copies) [68] |
| Burkholderia thailandensis E264 | PK-NRP thailandepsin mutant; efflux mutants [69] | Conjugation, electroporation | Polyketides (PKs), PK-NRPs from Betaproteobacteria, Myxococcia [69] | 985 mg/L FK228 derivative [69] |
| Burkholderia gladioli ATCC 10248 | PK gladiolin mutant [69] | Conjugation, electroporation | NRPs, PK-NRPs from Betaproteobacteria, Gammaproteobacteria [69] | Not specified |
| Burkholderia sp. FERM BP-3421 | PK-NRP spliceostatin mutants [69] | Conjugation, electroporation mimicry by methylation | RiPPs, PK-NRP-PUFAs from Betaproteobacteria [69] | 240 mg/L capistruin [69] |
| Streptomyces albus Del14 | Minimized genome background [70] | Intergeneric conjugation from E. coli | NRPS for pyrazinones (Ichizinones A-C) [70] | Confirmed production (titer not specified) [70] |
| Phaeodactylum tricornutum (diatom) | Naturally high lipid content, precursor availability [71] | Bacterial conjugation with episomal vectors | Cannabinoid pathway (tetraketide synthase) [71] | Olivetolic acid not detected; metabolic flux alterations observed [71] |
Experimental Protocol: Heterologous Expression of Biosynthetic Gene Clusters
The following methodology for heterologous expression in Streptomyces hosts has been adapted from established protocols in the field [68] [70]:
BGC Identification and Capture: Identify the target BGC through genome mining tools (e.g., antiSMASH). Capture the complete cluster from genomic DNA using transformation-associated recombination (TAR) cloning or similar methods.
Vector Construction and Modification: Clone the BGC into an appropriate expression vector containing:
- Conjugative transfer origin (oriT)
- Selection markers (e.g., apramycin resistance)
- Site-specific integration elements (e.g., ΦC31 attP-int)
- Optional: recombinase-mediated cassette exchange (RMCE) sites (loxP, vox, rox)
BGC Transfer to Heterologous Host:
- Introduce the constructed vector into a donor E. coli strain (e.g., ET12567/pUZ8002 or similar).
- Perform intergeneric conjugation between the donor E. coli and the recipient Streptomyces host:
- Grow both donor and recipient cultures to optimal density
- Mix cells, pellet, and resuspend in minimal volume
- Plate on appropriate medium and incubate (e.g., 30°C for 16-20 hours)
- Overlay with appropriate antibiotics and incubation continue
Exconjugant Selection and Validation:
- Select for single-crossover integrants using antibiotic selection
- Verify integration through PCR and Southern blotting
- For multi-copy integration: screen for increased antibiotic resistance correlating with copy number
Metabolite Production and Analysis:
- Inoculate verified exconjugants in production media (e.g., DNPM medium)
- Incubate with shaking (e.g., 7 days at 28°C)
- Extract metabolites with organic solvents (e.g., butanol)
- Analyze production using LC-MS/MS and compare to authentic standards
Heterologous Expression Workflow: From BGC to Product Analysis
Precursor Supplementation: Enhancing Native Biosynthetic Capability
Precursor supplementation focuses on enhancing the production of natural products within native or heterologous hosts by providing key biosynthetic building blocks that may be limiting in the natural metabolic context. This approach leverages the host's existing enzymatic machinery while overcoming metabolic bottlenecks through exogenous addition of pathway intermediates.
Table 2: Precursor Supplementation Strategies in Natural Product Biosynthesis
| Target Compound/Class | Host System | Supplemented Precursors | Experimental Outcomes | Limitations/Challenges |
|---|---|---|---|---|
| Cannabinoids (Olivetolic acid) | Phaeodactylum tricornutum [71] | Endogenous malonyl-CoA, hexanoyl-CoA (precursor pathway engineering) | Enzyme expression confirmed but OA accumulation not detected; significant metabolome alterations [71] | Potential diversion of precursors to endogenous metabolism; complex pathway regulation |
| Fungal Secondary Metabolites | Various fungal cultures [72] | Amino acids, short-chain fatty acids, specialized biosynthetic intermediates | Enhanced antibiotic production in some fungal strains; activation of cryptic BGCs [72] | Variable response across different fungal taxa; precursor uptake limitations |
| Pyrazinones (Ichizinones) | Streptomyces sp. LV45-129 (native) and heterologous hosts [70] | Amino acid precursors (valine, leucine, beta-amino acids) | Production of Ichizinones A-C in native host; successful heterologous expression without supplementation [70] | Specific precursor requirements not fully elucidated |
Experimental Protocol: Precursor Supplementation Studies
Methodology for precursor supplementation experiments, as demonstrated in cannabinoid pathway engineering in diatoms [71]:
Host Engineering:
- Identify key pathway enzymes and corresponding genes from source organism (e.g., tetraketide synthase (TKS) and olivetolic acid cyclase (OAC) from Cannabis sativa)
- Clone genes into appropriate expression vectors with strong, constitutive promoters
- For photosynthetic hosts: use species-specific promoters (e.g., 40SRPS8 promoter for P. tricornutum)
- Transfer constructs to host via appropriate method (e.g., bacterial conjugation for diatoms)
Precursor Feeding Strategy:
- Determine critical pathway precursors through metabolic mapping
- Design feeding experiments with potential precursors (e.g., hexanoic acid, malonate)
- Establish optimal feeding timing (typically during active growth phase)
- Optimize precursor concentrations through dose-response experiments
Metabolomic Analysis:
- Harvest cells at multiple time points post-precursor supplementation
- Extract metabolites using appropriate solvents (e.g., methanol, butanol)
- Perform UPLC-qTOF-MS analysis with relevant standards
- Use untargeted metabolomics to assess global metabolic changes
- Statistical analysis (PCA, OPLS-DA) to identify significant metabolic alterations
Pathway Validation:
- Monitor target compound production using LC-MS with multiple reaction monitoring (MRM)
- Confirm compound identity using authentic standards when available
- Perform isotopic labeling studies to trace precursor incorporation
- Use enzyme assays to verify catalytic activity of heterologous enzymes
Precursor Supplementation and Metabolic Fate
Comparative Performance Analysis
When evaluating precursor supplementation versus heterologous expression for solving supply chain issues, each approach demonstrates distinct advantages and limitations that make them suitable for different research scenarios.
Table 3: Direct Comparison of Strategies for Natural Product Supply
| Parameter | Precursor Supplementation | Heterologous Expression |
|---|---|---|
| Technical Complexity | Moderate (requires metabolic understanding but less genetic manipulation) | High (demands specialized skills in molecular biology and genetics) |
| Development Timeline | Shorter (weeks to months for optimization) | Longer (months to years for host engineering and optimization) |
| Production Yield Potential | Variable; often limited by native regulatory mechanisms | Potentially higher; amenable to copy number and promoter optimization |
| Scalability | Limited by native host growth characteristics | Generally superior with fermentable chassis organisms |
| Applicability to Unculturable Sources | Not applicable | Enables production from unculturable organisms [73] |
| Pathway Elucidation Capability | Limited to testing specific hypotheses | Powerful for complete pathway validation and characterization |
| Representative Success Cases | Enhanced antibiotic production in fungi [72] | Griseorhodin H, xiamenmycin, ichizinones [68] [70] |
| Key Limitations | Precursor uptake, metabolic diversion, native regulation | Codon usage, post-translational modifications, precursor availability |
The Scientist's Toolkit: Essential Research Reagents and Solutions
Successful implementation of either strategy requires specific reagents and genetic tools. The following table summarizes key solutions used in the cited studies.
Table 4: Essential Research Reagents for Biosynthesis Studies
| Reagent/Tool | Function/Application | Examples from Literature |
|---|---|---|
| ΦC31-based Integration System | Site-specific integration of BGCs into actinomycete chromosomes [68] | Used in Streptomyces coelicolor and Burkholderia hosts [68] [69] |
| RMCE Cassettes (Cre-lox, Vika-vox, Dre-rox) | Recombinase-mediated cassette exchange for precise genome engineering [68] | Enables multi-copy integration in Micro-HEP platform [68] |
| pBBR1 and pRO1600 Replicons | Broad-host-range plasmids for gene expression in proteobacteria [69] | Used in Burkholderia heterologous expression systems [69] |
| Red/ET Recombineering | Efficient genetic manipulation of BGCs in E. coli intermediate hosts [68] [70] | Used for markerless gene deletions and cluster modifications [70] |
| AntiSMASH | Bioinformatics tool for BGC identification and analysis [68] [73] | Standard for genome mining and BGC prediction [68] |
| Conjugative Transfer Systems | Intergeneric DNA transfer from E. coli to recalcitrant hosts [68] [70] [71] | ET12567/pUZ8002 and similar systems for actinomycetes and diatoms [68] [71] |
| Inducible Promoter Systems | Controlled gene expression (rhamnose-, arabinose-inducible) [69] | Fine-tuned expression in heterologous hosts [69] |
Within the framework of natural product validation through LC-MS and bioassay research, both precursor supplementation and heterologous expression offer powerful—and potentially complementary—solutions to critical supply chain challenges. Heterologous expression demonstrates superior capabilities for producing complex natural products from unculturable sources and achieving scalable yields through systematic host engineering. The development of optimized chassis strains like S. coelicolor A3(2)-2023 and various Burkholderia species provides increasingly sophisticated platforms for BGC expression [68] [69]. Meanwhile, precursor supplementation offers a more rapid approach to enhancing production in native hosts, though it faces limitations from endogenous metabolic regulation and precursor uptake barriers [71]. The choice between these strategies should be guided by the specific research goals, available resources, and timeline constraints. For comprehensive natural product biosynthesis validation, a sequential approach often proves most effective: using precursor supplementation to rapidly test biosynthetic hypotheses, followed by heterologous expression to establish robust, scalable production platforms for further development and application.
Liquid Chromatography-Mass Spectrometry (LC-MS) has become an indispensable technology in the validation of natural product biosynthesis, enabling researchers to decipher complex chemical structures and their biological activities. However, the journey from raw spectral data to confident metabolite identification presents significant analytical challenges that can hinder research progress. This guide examines the core data analysis hurdles in LC-MS-based natural product research and objectively compares the computational strategies and software solutions available to overcome them, with supporting experimental data from recent studies.
Core Computational Challenges in LC-MS Data Analysis
The analysis of LC-MS data in natural product research confronts several persistent technical hurdles that impact data reliability and interpretation.
Dynamic Range and Sensitivity Limitations
The tremendous dynamic range of compound concentrations in biological samples presents a fundamental detection challenge. In natural product extracts, abundant compounds can obscure crucial low-abundance metabolites, potentially missing biologically significant molecules. Advanced fractionation techniques and high-resolution MS instruments have improved this dynamic range, but low-throughput and robustness issues remain problematic [74].
Metabolite Identification Confidence
Unlike the predictable building blocks of proteins, metabolites represent random combinations of elements with extensive isomerism. This complexity makes confident identification difficult—a single molecular ion can yield over 100 putative identifications through mass-based database searches alone [75]. This identification ambiguity necessitates sophisticated computational filtering and validation strategies.
Data Alignment and Integration Challenges
LC-MS datasets acquired across different laboratories, instruments, or even batches exhibit significant variability in retention times and mass measurements. This variability complicates data alignment, a crucial step where LC-MS features from common ions are assembled into a unified analysis matrix. Large-scale studies particularly suffer from chromatographic drift between batches, creating interoperability challenges [76].
Matrix Effects and Ionization Suppression
Unwanted matrix effects—ion suppression or enhancement—remain a persistent technical hurdle. These effects alter ionization efficiency and quantitative accuracy, particularly in complex natural product extracts. Manufacturers are actively working to improve ionization reproducibility and reduce these matrix effects through interface and ion optics innovations [77].
Comparative Analysis of Computational Strategies
Data Acquisition Methods Comparison
Table 1: Comparison of LC-MS/MS Data Acquisition Methods
| Method | Mechanism | Advantages | Limitations | Best Applications |
|---|---|---|---|---|
| Data-Dependent Acquisition (DDA) | Automatically selects precursors above abundance threshold for fragmentation | High-quality MS/MS spectra; Clear precursor-product ion relationships | May miss low-abundance ions; Limited to top N most abundant precursors | Untargeted screening of moderate-abundance metabolites |
| Data-Independent Acquisition (DIA) | Fragments all ions within specified m/z windows without precursor selection | Broader analyte coverage; Reduced intensity bias | Complex spectral deconvolution; Requires advanced software | Comprehensive lipidomics; Complex mixture analysis |
| Selected Reaction Monitoring (SRM) | Monitors specific precursor-product ion transitions | Excellent sensitivity and specificity; Gold standard for quantitation | Targeted approach only; Requires prior knowledge | Validation studies; Targeted quantitation of known compounds |
Software Solutions for Data Processing
Table 2: Software Solutions for LC-MS Data Challenges
| Software Platform | Primary Function | Key Features | Natural Product Applications | Limitations |
|---|---|---|---|---|
| metabCombiner | Multi-dataset alignment | Stepwise alignment of disparate LC-MS datasets; Handles RT variability | Inter-laboratory reproducibility studies; Multi-batch experiments | Requires programming knowledge (R package) |
| GNPS Molecular Networking | MS/MS spectral similarity analysis | Groups MS/MS spectra into structural scaffolds; Cloud-based platform | Natural product dereplication; Library reduction | Internet dependency for cloud processing |
| Skyline | Targeted data processing | Quantitative LC-MS data analysis; Support for SRM and DIA | Natural product quantitation; Method development | Steeper learning curve for complex workflows |
| Proteome Discoverer | Proteomics data analysis | Protein identification and quantification; PTM analysis | Natural product-protein interaction studies | Primarily optimized for proteomics |
Experimental data from a 2025 study demonstrates that computational scaffold-based library reduction using LC-MS/MS and molecular networking achieved an 84.9% reduction in library size while increasing bioassay hit rates from 11.3% to 22% against Plasmodium falciparum [6].
Experimental Protocols for Method Validation
Computational Workflow for Metabolite Identification
The following diagram illustrates the systematic computational framework for metabolite identification, which reduces manual verification burden by prioritizing putative identifications:
Method Validation Framework for Bioinformatic Approaches
Drawing from established bioassay validation principles, computational methods for natural product identification should undergo rigorous validation to ensure reliability:
Preliminary Development: Define method scope, endpoints, and analytical requirements including acceptable error margins [78].
Feasibility Experiments: Verify performance parameters using control compounds and draft standard operating procedures.
Internal Validation: Assess method performance characteristics including precision, accuracy, and robustness in a single laboratory setting.
External Validation: Evaluate method transferability across multiple laboratories or experimental conditions to establish fitness-for-purpose [78].
Experimental protocols should incorporate total ion current (TIC) normalization and surrogate internal standards to eliminate technical variations, with spiked-in compounds serving as quality controls for both sample preparation and data processing steps [11].
The Scientist's Computational Toolkit
Essential Research Reagent Solutions
Table 3: Key Computational Tools for Natural Product LC-MS Research
| Tool Category | Specific Solutions | Function | Application in Natural Product Research |
|---|---|---|---|
| Spectral Libraries | NIST MS/MS, MassBank, GNPS Libraries | Reference fragmentation patterns | Metabolite identification by spectral matching |
| Data Processing Packages | metabCombiner, XCMS, MZmine | Feature detection, alignment, and normalization | Multi-batch data integration; Metabolic fingerprinting |
| Molecular Networking | GNPS Classical Molecular Networking | Scaffold-based compound grouping | Library redundancy reduction; Bioactive compound discovery |
| Quantitation Platforms | Skyline, Chromeleon | Targeted and untargeted quantitation | Natural product potency assessment; Biosynthetic yield optimization |
| Cloud-Based Technologies | Thermo Fisher Ardia Platform | Data sharing, collaboration, and remote analysis | Multi-institutional natural product discovery projects |
Case Study: Rational Natural Product Library Reduction
A 2025 study demonstrated an innovative application of LC-MS/MS and molecular networking to address structural redundancy in natural product libraries. Using a collection of 1,439 fungal extracts, researchers applied computational scaffold-based selection to create minimal libraries representing maximum chemical diversity [6].
Experimental Protocol:
- Acquired untargeted LC-MS/MS data for all extracts
- Processed data through GNPS molecular networking to group MS/MS spectra into structural scaffolds
- Used custom R code to iteratively select extracts with the greatest scaffold diversity
- Validated approach through bioactivity testing against multiple targets
Performance Metrics:
- The method achieved 100% scaffold diversity with only 216 extracts (6.6-fold reduction)
- Bioassay hit rates increased from 2.57% to 8.00% for neuraminidase inhibition
- 16 of 17 activity-correlated features were retained in the reduced library [6]
Managing complex LC-MS datasets and confidently identifying metabolites remains challenging in natural product biosynthesis research. However, as computational strategies evolve, they offer increasingly robust solutions to these hurdles. The integration of advanced data acquisition methods, sophisticated alignment algorithms, and rigorous validation frameworks provides a pathway to more efficient and reliable natural product discovery. By strategically implementing the tools and methodologies compared in this guide, researchers can significantly enhance their capability to validate natural product biosynthesis and accelerate drug development pipelines.
Orthogonal Validation Strategies: LC-MS, Bioassay, and Comparative Methodologies
In the rigorous fields of natural product research and drug development, the validation of analytical methods is paramount. Analytical figures of merit are quantitative metrics that provide objective evidence that an analytical method is fit for its intended purpose, ensuring that experimental data generated is reliable, accurate, and reproducible. For researchers employing LC-MS and bioassays to validate natural product biosynthesis, three figures of merit are particularly critical: sensitivity, specificity, and reproducibility. Sensitivity refers to the ability of a method to detect small changes in analyte concentration; specificity is the capacity to distinguish the analyte from other components in a complex mixture; and reproducibility denotes the precision of the method under varied conditions over time [11] [79]. These metrics form the foundation for trusting data that leads to discoveries in cellular targeting, mechanism of action, and the therapeutic potential of natural products like artemisinin, paclitaxel, and berberine [11].
The process of establishing these metrics is formalized through method validation, a process that demonstrates a technique is suitable for its intended purpose and that the results obtained are reliable. This is especially crucial when developing new bioassays for novel classes of insecticides or applying LC-MS proteomics to uncover how natural products influence cellular processes [79]. Without a rigorous validation process, the high variability inherent in biological tests can lead to unreliable results, hindering scientific progress and drug development. This guide provides a comparative framework for establishing these vital figures of merit, framed within the context of validating natural product biosynthesis.
Comparative Analysis of Method Validation Approaches
The approach to validation can differ significantly between established fields like clinical chemistry and more application-specific areas like bioassay development. The table below compares the core frameworks and their applicability to LC-MS and bioassay research.
Table 1: Comparison of Validation Frameworks Across Disciplines
| Feature | General Analytical Chemistry & Clinical | LC-MS-Based Proteomics | Bioassay for Vector Control/Natural Products |
|---|---|---|---|
| Core Philosophy | Highly standardized, quantitative parameters defined by regulatory bodies. | Adapts general principles to high-throughput protein identification and quantification. | Modular framework acknowledging biological variability; draws from chemical and healthcare fields [79]. |
| Key Validation Parameters | Accuracy, precision, linearity, range, specificity, limit of detection (sensitivity) [79]. | Sensitivity, specificity, reliability in large-scale protein data and post-translational modifications [11]. | Precision (imprecision), accuracy (trueness/inaccuracy), robustness, defined endpoints [79]. |
| Defining Acceptability Criteria | Strict, predefined allowable error limits. | Based on the required specificity and sensitivity for protein network analysis [11]. | Allowable error is defined during development, should be as small as possible yet practically achievable (e.g., CV < 20%) [79]. |
| Primary Challenge | Meeting stringent regulatory requirements. | Managing complex data analysis and confounding experimental variations [11]. | Accounting for inherent variability in live biological material (e.g., insects, cell lines) and non-homogeneous products [79]. |
| Typical Experimental Replication | Defined by statistical power and regulatory guidelines. | Multiple technical and biological replicates for statistical confidence in protein expression. | Validation stages (feasibility, internal, external) with experiments designed to measure analytical error [79]. |
Establishing Figures of Merit in LC-MS-Based Proteomics
Liquid Chromatography-Mass Spectrometry (LC-MS) has become a powerful platform for identifying and quantifying proteins affected by natural product (NP) exposure, providing insights into cellular targeting and mechanisms of action [11]. Validating these methods is crucial for generating reliable data.
Experimental Protocols for LC-MS Workflows
A typical LC-MS proteomics workflow for studying natural products involves several key stages, with validation metrics embedded throughout:
- Cell Line Selection and Treatment: Select biologically relevant cell lines (e.g., MCF-7 for breast cancer). Cells are exposed to the NP and a vehicle control [11].
- Protein Extraction and Digestion: Proteins are extracted and digested into peptides using enzymes like trypsin. Robustness is tested by varying digestion time or enzyme-to-protein ratios [11].
- LC-MS Analysis:
- Liquid Chromatography (LC): Peptides are separated. Specificity is demonstrated by the method's ability to resolve complex peptide mixtures.
- Mass Spectrometry (MS): Peptides are ionized and detected. Sensitivity is determined by the limit of detection for low-abundance peptides.
- Data Normalization and Analysis: Raw data is processed using specialized software (e.g., Skyline, Proteome Discoverer). Normalization methods, such as Total Ion Current (TIC) or quantile normalization, are applied to eliminate experimental variations and control for reproducibility [11]. Statistical analysis (e.g., t-tests) identifies differentially expressed proteins, but as noted in microarray studies, relying solely on statistical significance (p-values) can reduce reproducibility; incorporating fold-change criteria is often essential [80].
Diagram: LC-MS Proteomics Workflow for Natural Product Validation
Key Reagent Solutions for LC-MS Proteomics
The following reagents and materials are essential for successful LC-MS-based proteomics.
Table 2: Key Research Reagent Solutions for LC-MS Proteomics
| Item Name | Function/Brief Explanation |
|---|---|
| Trypsin (Protease) | Enzyme used for bottom-up proteomics; digests proteins into smaller peptides for LC-MS analysis [11]. |
| Stable Isotope Labels (SILAC, TMT) | Label-based quantification reagents. Incorporate stable isotopes into peptides, allowing for precise multiplexed quantification of protein expression across samples [11]. |
| Spiked-In Internal Standards | Synthetic isotope-labeled peptides of known quantity and sequence. Used for data normalization, controlling for experimental variation, and improving reproducibility [11]. |
| LC-MS Grade Solvents | High-purity solvents (e.g., water, acetonitrile) for mobile phases. Essential for minimizing background noise and maximizing sensitivity and specificity. |
| Specific Software (Skyline, Proteome Discoverer) | Specialized bioinformatics tools for processing LC-MS raw data, enabling peptide identification, quantification, and statistical analysis [11]. |
Establishing Figures of Merit in Bioassay Research
Bioassays used to evaluate vector control tools, or more broadly, the biological activity of natural products, face unique validation challenges due to their reliance on live biological material [79].
A Framework for Bioassay Method Validation
A proposed validation framework for bioassays involves four key stages to ensure reliability [79]:
- Preliminary Development: The method's scope, endpoints (e.g., mosquito mortality, oviposition inhibition), and acceptability criteria are defined. Allowable analytical error is established.
- Feasibility Experiments: Initial small-scale tests verify performance parameters and endpoints. A draft Standard Operating Procedure (SOP) is written.
- Internal Validation: The method's analytical performance (precision, accuracy) is rigorously tested in a single laboratory. A method claim is drafted.
- External Validation: The method is evaluated in multiple independent laboratories or semi-field sites to confirm its reproducibility and transferability. The final method claim is produced.
Diagram: Bioassay Method Validation Framework
Case Study: The m:n:θb Procedure for Precision
A common procedure for validating assay precision is the m:n:θb procedure, where m levels of an analyte are measured with n replicates at each level. The assay passes if all m estimates of the coefficient of variation (CV) are less than a bound, θb (e.g., a 3:5:15% procedure) [81]. However, this procedure's statistical properties are often overlooked. Under a constant CV model, if the true CV equals θb, the probability of passing can be as low as 10-20% for some recommended implementations, meaning a truly precise assay might fail. Conversely, with extreme heterogeneity, the passing probability can be over 50% even if one level has a CV at the bound [81]. This highlights the need for robust statistical understanding during validation. For relative potency assays (e.g., growth inhibition assays), a constant standard deviation (SD) model often fits better than a constant CV model, requiring a different validation approach [81].
Comparative Data on Reproducibility and Statistical Analysis
The MAQC (MicroArray Quality Control) project provided seminal insights into the reproducibility of biomarker lists, with lessons directly applicable to proteomics and bioassay data analysis. A key finding was that ranking and selecting differentially expressed genes (or proteins) solely by statistical significance (P-value) from simple t-tests led to highly irreproducible lists between similar experiments [80]. This is a mathematical consequence of the high variability of t-values when sample sizes are small.
Table 3: Impact of Gene Selection Method on List Reproducibility
| Gene Selection / Ranking Criterion | Inter-Site Reproducibility (POG for ~20 genes) | Cross-Platform Reproducibility | Comment |
|---|---|---|---|
| P-value ranking alone | Low (20-40%) [80] | Much lower [80] | High variability; more stringent P-value thresholds yield less reproducible lists. |
| Fold Change (FC) ranking alone | High (Near 90%) [80] | Markedly improved (70-85%) [80] | Enhances reproducibility by incorporating magnitude of change. |
| FC-ranking + non-stringent P-value cutoff | Highest and most stable [80] | Highest and most stable [80] | Recommended practice: Balances reproducibility (FC) with sensitivity/specificity (P). |
The recommended practice to generate more reproducible results is to use FC-ranking plus a non-stringent P-value cutoff. The P-value cutoff should not be too small, and the FC should be as large as possible. This joint criterion enhances reproducibility while balancing sensitivity and specificity [80].
The Scientist's Toolkit: Essential Materials for Validation
Beyond the specific reagents for LC-MS or bioassays, a core set of conceptual tools is essential for any scientist establishing figures of merit.
Table 4: Essential Methodological Tools for Analytical Validation
| Tool / Concept | Function/Brief Explanation |
|---|---|
| Standard Operating Procedure (SOP) | A detailed, step-by-step document describing the entire method. Critical for ensuring consistency and reproducibility during internal and external validation [79]. |
| Allowable Analytical Error | A predefined threshold combining random (imprecision) and systematic (inaccuracy) errors. The total method error must be within this limit for the method to be considered valid [79]. |
| Coefficient of Variation (CV) | A standardized measure of precision (CV = Standard Deviation / Mean). Used to set acceptability criteria for precision (e.g., within-day CV < 20%) [79]. |
| Controls of Known Value | Samples with a known concentration or response. Used during method verification and routine use to monitor the method's accuracy and precision over time [79]. |
| Fold Change (FC) Criterion | A predefined threshold for the magnitude of change (e.g., 2-fold). Using FC as a primary ranking criterion, alongside statistical tests, dramatically improves the reproducibility of hit lists in 'omics' studies [80]. |
In the fields of natural product research and drug development, the accurate identification and quantification of target molecules are paramount. Analytical techniques form the backbone of research aimed at validating natural product biosynthesis, profiling metabolites, and advancing therapeutic candidates. For decades, immunoassays have served as the workhorse for bioanalysis in clinical and research settings, offering simplicity and rapid results. However, the emergence of liquid chromatography-tandem mass spectrometry (LC-MS/MS) has fundamentally shifted the analytical paradigm, establishing a new gold standard for specificity and accuracy. This comparison guide objectively examines the performance characteristics, applications, and limitations of LC-MS/MS versus immunoassays and other traditional methods, providing researchers with the experimental data necessary to select the optimal analytical approach for their specific needs in natural product research.
Fundamental Principles and Technological Comparison
Immunoassay Techniques: Mechanism and Evolution
Immunoassays (IAs) are biochemical tests that measure the presence or concentration of biological molecules, known as analytes, through the highly specific binding between an antigen and its antibody. This interaction, often described as a "lock and key" relationship, allows for the detection and quantification of diverse analytes in complex samples [82]. The technology has evolved significantly since Rosalyn Yalow and Solomon Berson developed radioimmunoassay (RIA) in the 1950s, for which Yalow became the second woman to win a Nobel Prize [82]. Modern immunoassays are categorized by several key characteristics:
- Reaction Method: Competitive vs. non-competitive formats. Competitive immunoassays are typically used for small molecules, where the target analyte and a labeled analog compete for a limited number of antibody binding sites, producing a signal inversely proportional to the analyte concentration. Non-competitive assays (e.g., sandwich ELISA) use excess antibody binding sites and produce a signal directly proportional to the amount of analyte [82].
- Detection Configuration: Direct vs. indirect detection. Direct assays use a primary antibody conjugated with a detection label, while indirect assays employ an unlabeled primary antibody followed by a labeled secondary antibody, offering signal amplification at the cost of additional procedural steps [82].
- Detection Labels: Radioactive (largely phased out due to safety concerns), chromogenic (colorimetric changes), fluorescence, and luminescence (the most sensitive method) [82].
Common immunoassay platforms include Western Blots (qualitative/semi-quantitative, low reproducibility), Enzyme-Linked Immunosorbent Assays (ELISA; quantitative, medium reproducibility), and bead-based immunoassays (quantitative, high reproducibility, capable of multiplexing) [82].
LC-MS/MS Technology: Unparalleled Specificity
Liquid chromatography-tandem mass spectrometry (LC-MS/MS) combines the physical separation capabilities of liquid chromatography with the exceptional mass analysis power of tandem mass spectrometry. This technique first separates components in a sample by liquid chromatography, then ionizes them and introduces the ions into the mass spectrometer. The core strength of LC-MS/MS lies in its tandem mass spectrometry component, which typically consists of three quadrupoles (Q1, Q2, Q3) that enable multiple types of experiments [83]:
- Full Scan: Identifies all ions in a sample.
- Product Ion Scan: Fragments a selected precursor ion and identifies all resulting product ions.
- Precursor Ion Scan: Identifies all precursors that fragment to a common product ion.
- Neutral Loss Scan: Identifies all precursors that lose a common neutral fragment.
- Selective/Multiple Reaction Monitoring (SRM/MRM): The workhorse for quantitative analysis, SRM/MRM selects a specific precursor ion in Q1, fragments it in Q2 (collision cell), and monitors one or more specific product ions in Q3. This double selection process provides exceptional specificity by effectively filtering out interferences [83].
The instrumentation typically uses an atmospheric pressure ionisation source, most commonly electrospray ionisation (ESI) or atmospheric pressure chemical ionisation (APCI), coupled to the tandem mass spectrometer [83]. For natural product research, LC-MS/MS has become indispensable for metabolite profiling and identification, particularly through LC-MS/MS-based molecular networking, which clusters metabolites based on common MS/MS fragmentation patterns to annotate compounds in complex extracts [32].
Table 1: Fundamental Characteristics of Analytical Platforms
| Feature | Immunoassays | LC-MS/MS |
|---|---|---|
| Basic Principle | Antibody-antigen binding | Physical separation followed by mass-based detection |
| Specificity Source | Antibody specificity | Chromatographic retention time and mass-to-charge ratio |
| Typical Workflow | Relatively simple, often automated | Multistep, complex, requires specialized expertise |
| Sample Throughput | High | Moderate (lower than automated immunoassays) |
| Key Experiment Types | Competitive, Non-competitive (Sandwich) | Full Scan, Product Ion, Precursor Ion, SRM/MRM |
| Primary Applications | Clinical diagnostics, protein detection | Metabolite profiling, steroid analysis, biomarker validation |
Comparative Performance Data: Analytical Figures of Merit
Sensitivity and Specificity: Fundamental Analytical Parameters
Sensitivity and specificity represent two critical parameters where LC-MS/MS demonstrates distinct advantages over immunoassays. The superior specificity of LC-MS/MS stems from its ability to differentiate between molecular isoforms, modifications, and structurally similar compounds that often cross-react in immunoassays [84]. This is particularly valuable in natural product research where complex mixtures of structurally similar metabolites must be distinguished.
Experimental data from clinical chemistry highlights this advantage. For instance, in hormone analysis, immunoassays suffer from interference from cross-reacting substances, especially at low analyte concentrations, as demonstrated for testosterone in neonates and for 25-hydroxyvitamin D [85]. Binding proteins can also cause interference, as seen with cortisol measurements [85]. LC-MS/MS mitigates these issues through its separation power and selective detection, minimizing matrix effects and interference common in immunoassays [84].
In a direct comparison of urinary free cortisol (UFC) measurement—a crucial diagnostic test for Cushing's syndrome—four new direct immunoassays showed strong correlations with LC-MS/MS (Spearman coefficients ranging from 0.950 to 0.998), but all immunoassays demonstrated proportionally positive biases compared to the LC-MS/MS reference method [26]. This systematic bias underscores the potential for immunoassays to overestimate concentrations due to residual cross-reactivity, even in modern platforms.
Quantitative Performance: Precision, Accuracy, and Dynamic Range
When evaluating quantitative performance, LC-MS/MS generally provides superior accuracy, precision, and wider dynamic ranges compared to immunoassays. The integration of stable isotope-labeled internal standards in LC-MS/MS methods corrects for variability in sample preparation, ionization efficiency, and matrix effects, resulting in more precise and accurate measurements [85].
External quality assurance data reveals that while the overall bias for LC-MS/MS methods is better than for immunoassays, there remains significant between-laboratory variation for some analytes [85]. This variation highlights the importance of standardized protocols and rigorous validation, even for LC-MS/MS methods. For immunoassays, the dynamic range typically spans only a few orders of magnitude, whereas LC-MS/MS maintains linearity over three to five orders of magnitude, facilitating the simultaneous quantification of analytes present at vastly different concentrations in the same sample [82].
Table 2: Quantitative Performance Comparison of UFC Measurement for Cushing's Syndrome Diagnosis
| Platform | Correlation with LC-MS/MS (Spearman r) | Bias vs. LC-MS/MS | AUC for CS Diagnosis | Optimal Cut-off (nmol/24h) |
|---|---|---|---|---|
| Autobio A6200 | 0.950 | Proportionally positive | 0.953 | 178.5 |
| Mindray CL-1200i | 0.998 | Proportionally positive | 0.969 | 194.5 |
| Snibe MAGLUMI X8 | 0.967 | Proportionally positive | 0.963 | 272.0 |
| Roche 8000 e801 | 0.951 | Proportionally positive | 0.958 | 196.0 |
| LC-MS/MS (Reference) | 1.000 | - | - | Established by lab |
Experimental Protocols and Workflows
Representative Protocol: Urinary Free Cortisol Comparison
A recent systematic comparison of four new immunoassays with LC-MS/MS for urinary free cortisol measurement provides an excellent case study for understanding experimental design in method comparison studies [26]. The protocol details are as follows:
Sample Preparation: Residual 24-hour urine samples from 337 patients (94 with Cushing's syndrome and 243 non-CS patients) were used. The LC-MS/MS method involved diluting urine specimens 20-fold with pure water, followed by the addition of an internal standard solution containing cortisol-d4. After centrifugation, the supernatant was injected into a SCIEX Triple Quad 6500+ mass spectrometer [26].
LC-MS/MS Analysis: Separation was achieved on an ACQUITY UPLC BEH C8 column using a binary mobile phase of water and methanol. The instrument operated in positive electrospray ionization mode with multiple reaction monitoring (MRM) tracking the following transitions: 363.2 → 121.0 (quantifier) and 363.2 → 327.0 (qualifier) for cortisol, and 367.2 → 121.0 for cortisol-d4 (internal standard) [26].
Immunoassay Analysis: The four immunoassay platforms (Autobio A6200, Mindray CL-1200i, Snibe MAGLUMI X8, and Roche 8000 e801) were operated according to manufacturers' instructions using direct methods without organic solvent extraction. All instruments were properly calibrated and quality controls were implemented as specified by manufacturers [26].
Statistical Analysis: Method comparison utilized Passing-Bablok regression and Bland-Altman plot analyses. Diagnostic performance was evaluated through ROC analysis, with optimal cut-off values determined using Youden's index [26].
LC-MS/MS Proteomics Protocol for Natural Product Research
In natural product research, LC-MS/MS-based proteomics has become a powerful platform for identifying protein targets and understanding mechanisms of action. A typical workflow includes [11]:
Cell Line Selection: Choosing biologically relevant cell lines (e.g., MCF-7 for breast cancer, A549 for lung cancer, HCT-116 for colon cancer) that accurately represent the disease or biological system being studied.
Sample Preparation: Processing cell lines after natural product exposure, including protein extraction, reduction, alkylation, and digestion (typically with trypsin).
LC-MS/MS Analysis: Separating peptides using nano-flow or ultra-performance liquid chromatography coupled to a tandem mass spectrometer capable of high-resolution mass measurements.
Data Acquisition: Employing data-dependent acquisition (DDA) or data-independent acquisition (DIA) to fragment peptides and generate MS/MS spectra.
Data Analysis: Searching MS/MS data against protein databases using specialized software (e.g., Skyline, Proteome Discoverer) and conducting bioinformatic analysis to identify pathways and processes affected by natural product treatment.
Application in Natural Product Biosynthesis Validation
Dereplication and Metabolite Profiling
In natural product research, dereplication—the process of quickly identifying known compounds in complex mixtures—is crucial for avoiding rediscovery of known metabolites and focusing resources on novel compounds [32]. LC-MS/MS has become indispensable for this application, particularly through LC-MS/MS-based molecular networking, which visualizes relationships between metabolites based on similar fragmentation patterns [32].
The integration of LC-MS/MS with database searching platforms like the Global Natural Products Social Molecular Networking (GNPS) allows researchers to compare MS/MS spectra of unknown metabolites against extensive spectral libraries, significantly accelerating identification [32]. This approach is particularly valuable in medicinal plant research, where students and researchers have successfully utilized LC-MS/MS to identify antioxidant metabolites in plants like rosemary, aloe, echinacea, and ashwagandha [32].
Targeted Protein Analysis and Mechanism Elucidation
LC-MS/MS-based proteomics provides unique insights into natural product-directed cellular targeting by enabling high-throughput identification and quantification of proteins affected by natural product exposure [11]. This platform allows researchers to map protein-protein interactions, signaling pathways, and post-translational modifications (PTMs) that underlie the biological effects of natural products [11].
For example, proteomic studies have clarified the mechanisms of various natural products:
- Berberine: Chemoproteomics revealed that berberine directly binds to PKM2 to inhibit colorectal cancer progression [11].
- Withaferin A: SILAC-based quantitative MS identified proteins regulated by this natural product in prostate cancer models [11].
- Nigella sativa seed extract, green tea extract, and Kerra: Proteomic approaches have elucidated their antiproliferative and apoptotic effects in various cancer cell lines [11].
These studies demonstrate how LC-MS/MS platforms, including bottom-up proteomics, top-down proteomics, and targeted proteomics, provide a comprehensive view of protein dynamics in response to natural product treatment [11].
Visualizing Workflows and Signaling Pathways
Diagram 1: Comparative Workflows of LC-MS/MS and Immunoassays in Natural Product Research. The LC-MS/MS pathway emphasizes physical separation and mass-based detection, while the immunoassay pathway relies on specific antibody-antigen interactions.
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Research Reagents and Materials for Analytical Methods in Natural Product Research
| Reagent/Material | Function | Application in LC-MS/MS | Application in Immunoassays |
|---|---|---|---|
| Stable Isotope-Labeled Internal Standards | Corrects for variability in sample preparation and ionization; improves quantification accuracy | Essential for precise quantification; e.g., cortisol-d4 for cortisol analysis [26] | Not typically used |
| Specific Antibodies | Binds target analyte with high specificity | Limited use in immunocapture prior to MS analysis | Core component; critical for assay specificity and sensitivity [82] |
| LC Columns (C8, C18) | Separates compounds in complex mixtures based on hydrophobicity | Critical for resolving analytes prior to MS detection; e.g., ACQUITY UPLC BEH C8 [26] | Not used |
| Enzyme Conjugates | Generates detectable signal through enzymatic reaction | Not used | Core detection component in ELISA; e.g., horseradish peroxidase conjugates [82] |
| Mass Spectrometry-Grade Solvents | Mobile phase for chromatographic separation | Essential for minimal background and optimal ionization | Not critical; standard HPLC-grade often sufficient |
| Solid-Phase Extraction Cartridges | Pre-concentrates and purifies analytes from complex matrices | Used for sample clean-up to reduce matrix effects; e.g., C18 SPE for plant extracts [32] | Occasionally used to remove interfering substances |
| Reference Standards | Provides known quantities for calibration and identification | Essential for method development and calibration | Used for standard curve generation in quantitative assays |
| Bioinformatic Tools (GNPS, Skyline) | Analyzes complex MS/MS data and facilitates metabolite identification | Critical for dereplication and metabolite annotation [32] | Not applicable |
The comparative analysis of LC-MS/MS and immunoassays reveals a clear technological landscape where each platform offers distinct advantages for specific applications in natural product research. LC-MS/MS stands as the unequivocal gold standard for applications demanding high specificity, the ability to distinguish closely related molecular structures, and comprehensive metabolite profiling. Its superiority in quantifying small molecules with minimal cross-reactivity makes it particularly valuable for validating natural product biosynthesis and elucidating mechanisms of action through proteomic approaches.
Immunoassays, despite their limitations in specificity, maintain important roles in scenarios requiring high-throughput analysis, point-of-care testing, and detection of proteins where LC-MS/MS methods remain challenging. The emergence of digital ELISA and other advanced immunoassay formats continues to push the sensitivity boundaries of antibody-based detection.
For researchers validating natural product biosynthesis, the strategic selection between these platforms should be guided by the specific research questions, required level of specificity, throughput requirements, and available resources. LC-MS/MS provides the definitive analytical validation for structural characterization and quantification, while immunoassays offer practical solutions for rapid screening and high-volume clinical applications. As both technologies continue to evolve, their synergistic application promises to accelerate natural product discovery and development, ultimately advancing therapeutic options for various diseases.
Correlating Chemical Fingerprints with Bioactivity Profiles
For researchers in natural product drug discovery, accurately predicting the biological activity of a compound from its chemical structure is a central challenge. Molecular fingerprints, which encode chemical structures into bit-string or numerical vectors, serve as a fundamental tool for this task, enabling computational comparisons and bioactivity predictions. However, the unique structural complexity of natural products—characterized by high stereochemical diversity, extensive sp3 carbon frameworks, and intricate ring systems—presents specific challenges for their representation via conventional fingerprints [86]. The core thesis is that the strategic selection and application of these fingerprints are critical for validating the biosynthesis of natural products, effectively bridging the gap between LC-MS-based metabolite profiling and bioassay-guided isolation research. This guide provides an objective comparison of fingerprint performance, supported by experimental data, to inform the workflows of researchers, scientists, and drug development professionals.
Molecular Fingerprints: Types and Principles
Molecular fingerprints are computational representations that transform a molecule's structural information into a standardized format, facilitating rapid similarity searches and quantitative structure-activity relationship (QSAR) modeling. Their utility is paramount for processing the vast chemical space of natural products [87]. Fingerprints can be broadly categorized based on the algorithmic approach used to generate them.
Fingerprint Classification
- Dictionary-based Fingerprints (Structural Keys): These fingerprints use a predefined dictionary of functional groups or substructural motifs. Each bit in the fingerprint vector signifies the presence (1) or absence (0) of a specific fragment. Examples include MACCS and PubChem (PC) fingerprints. They are efficient for fast substructure searching [87].
- Circular Fingerprints: These algorithms dynamically generate fragments from a molecule's graph structure without a predefined list. They start from each atom and iteratively incorporate information from neighboring atoms within a specified radius, leading to a set of unique, "circular" fragments. Common examples are Extended Connectivity Fingerprints (ECFP) and Functional Class Fingerprints (FCFP), with the latter focusing on pharmacophoric features rather than atomic identities [86] [87].
- Path-based (Topological) Fingerprints: This category generates features by analyzing the paths through the molecular graph. Atom Pairs (AP) and Topological Torsion (TT) are classic examples that capture connectivity and distance relationships between atoms [86].
- Pharmacophore Fingerprints: These representations focus on the spatial arrangement of functional features critical for molecular recognition, such as hydrogen bond donors/acceptors and hydrophobic regions. They describe a molecule based on its potential for biological interactions rather than its exact atomic structure [87].
- String-based Fingerprints: These operate directly on the SMILES (Simplified Molecular-Input Line-Entry System) string of a compound. Methods like MinHashed Fingerprints (MHFP) and MinHashed Atom Pair Fingerprints (MAP4) fragment the SMILES string and use hashing techniques to create a fixed-size representation [86].
Table 1: Categories and Characteristics of Molecular Fingerprints
| Fingerprint Category | Representative Examples | Underlying Principle | Key Characteristics |
|---|---|---|---|
| Dictionary-Based | MACCS, PubChem (PC) | Predefined list of structural fragments | Fast; interpretable; may miss novel scaffolds |
| Circular | ECFP, FCFP | Dynamically generated circular neighborhoods from molecular graph | Captures novel structures; robust; widely used |
| Path-Based | Atom Pairs (AP), Topological Torsion (TT) | Enumeration of paths or torsions in molecular graph | Encodes topological distance information |
| Pharmacophore | Pharmacophore Pairs (PH2), Triplets (PH3) | 2D/3D arrangement of chemical features (e.g., H-bonding) | Linked to bioactivity; describes interaction potential |
| String-Based | MHFP, MAP4 | Fragmentation of SMILES strings using hashing techniques | Alignment-free; captures SMILES syntax nuances |
Comparative Performance of Fingerprints with Natural Products
The structural uniqueness of natural products means that fingerprint performance benchmarks established with synthetic, drug-like compound libraries do not always translate directly. Comprehensive benchmarking studies are essential to guide method selection.
Benchmarking Studies and Key Findings
A landmark 2009 study compared fingerprint methods based on their ability to reproduce similarities in biological activity space, using the BioPrint database of biological activity profiles. It concluded that fingerprints describing global molecular features, such as CHEMGPS or TRUST4 (which incorporate physicochemical properties and pharmacophore patterns), were often superior at identifying compounds with similar biological activity profiles, even in the presence of significant structural differences, compared to purely structural fingerprint methods [88].
A more recent 2024 benchmark evaluated 20 different fingerprinting algorithms on over 100,000 unique natural products from the COCONUT and CMNPD databases. The study focused on their performance in QSAR modeling across 12 bioactivity prediction tasks. A critical finding was that while Extended Connectivity Fingerprints (ECFP) are the de-facto standard for drug-like compounds, other fingerprints could match or outperform them for natural product bioactivity prediction [86]. This underscores the necessity of evaluating multiple fingerprint types for optimal performance on NP-centric tasks.
Quantitative Performance Data
The following table summarizes key quantitative results from the 2024 benchmark study, providing a direct comparison of fingerprint efficacy for natural product bioactivity prediction [86].
Table 2: Fingerprint Performance in Natural Product Bioactivity Prediction (Adapted from [86])
| Fingerprint Category | Representative Examples | Average Balanced Accuracy (Range across 12 datasets) | Key Strengths and Context |
|---|---|---|---|
| Circular | ECFP4 | ~0.75 (0.68 - 0.82) | Robust baseline performance; widely applicable. |
| Circular | FCFP4 | ~0.76 (0.69 - 0.83) | Can outperform ECFP by focusing on pharmacophoric features. |
| String-Based | MHFP6 | ~0.77 (0.70 - 0.84) | Matches or outperforms ECFP; useful for complex NP structures. |
| String-Based | MAP4 | ~0.78 (0.71 - 0.85) | Top performer; combines topological info with hashing. |
| Path-Based | Atom Pair (AP) | ~0.73 (0.66 - 0.80) | Good performance; provides a different similarity perspective. |
| Pharmacophore | PH2/PH3 | ~0.74 (0.67 - 0.81) | Higher biological relevance; can identify functionally similar NPs. |
| Dictionary-Based | MACCS | ~0.70 (0.63 - 0.77) | Computationally efficient; performance can be lower for novel NPs. |
Experimental Protocols for Fingerprint Validation
Validating the correlation between chemical fingerprints and bioactivity requires a rigorous experimental workflow that integrates analytical chemistry, biological testing, and computational analysis. The following protocol outlines a hybrid strategy, combining metabolomics with bioassay-guided principles.
Integrated LC-MS/MS and Bioassay Workflow
This protocol is designed for the analysis of a complex natural extract, such as a medicinal plant specimen, to identify bioactive metabolites.
Step 1: Sample Preparation and Extraction
- Procedure: Plant material is dried, ground, and extracted using a solvent of choice (e.g., acetone or methanol/water mixtures) via maceration or accelerated solvent extraction. The extract is vacuum-filtered and concentrated in vacuo [32].
- Rationale: Standardized extraction ensures a representative sample of the metabolome for downstream analysis.
Step 2: Bioactivity Screening
- Procedure: The crude extract is subjected to a relevant in vitro bioassay (e.g., DPPH assay for antioxidant activity, or an enzyme inhibition assay). This establishes a baseline level of bioactivity for the extract [32] [20].
- Rationale: This initial bioassay confirms the presence of the desired biological effect and provides a benchmark for tracking activity during fractionation.
Step 3: Fractionation and Activity Tracking
- Procedure: The active crude extract is fractionated using techniques like solid-phase extraction (SPE) or medium-pressure liquid chromatography (MPLC). All generated fractions are then re-tested in the same bioassay [32].
- Rationale: This bioassay-guided isolation step narrows down the chemical complexity by pinpointing the fractions responsible for the observed bioactivity.
Step 4: LC-MS/MS Analysis of Active Fractions
- Procedure: The active fraction(s) are analyzed using Liquid Chromatography-tandem Mass Spectrometry (LC-MS/MS).
- Chromatography: Typically, Reversed-Phase (C18) UHPLC with a water-acetonitrile gradient.
- Mass Spectrometry: Data-Dependent Acquisition (DDA) on a high-resolution instrument (e.g., Q-TOF or Orbitrap). This method first performs a full MS scan and then automatically selects the most intense ions for fragmentation (MS/MS) [32] [11].
- Rationale: LC-MS/MS provides two critical pieces of data: the accurate mass of metabolites (from MS1) and their characteristic fragmentation patterns (from MS2), which are essential for identification.
Step 5: Metabolite Identification and Dereplication
- Procedure: The acquired MS/MS spectra are compared against reference libraries, such as the Global Natural Products Social Molecular Networking (GNPS) platform. This dereplication step identifies known compounds and helps avoid rediscovery [32].
- Rationale: Dereplication is a crucial efficiency step in natural product discovery, allowing researchers to focus resources on novel or unconfirmed metabolites [32].
Step 6: Chemical Fingerprint Calculation and Correlation Analysis
- Procedure: The chemical structures of identified (and unknown) metabolites in the active fraction are converted into one or more types of molecular fingerprints (e.g., ECFP4, MAP4, PH3).
- Similarity Analysis: Pairwise similarities between all compounds are calculated using an appropriate metric (e.g., Jaccard-Tanimoto similarity).
- Bioactivity Correlation: The computed chemical similarities are then compared to the observed biological activity profiles. For instance, compounds clustering closely in fingerprint space should, in theory, exhibit similar bioactivity [88] [86].
- Rationale: This step directly tests the hypothesis that the chosen fingerprint can accurately map the chemical space to the biological activity space for the given set of natural products.
The Scientist's Toolkit: Essential Research Reagents and Materials
The following table details key reagents, software, and databases essential for executing the experimental and computational workflows described in this guide.
Table 3: Essential Research Reagents and Computational Tools
| Item Name | Function/Application | Relevance to Workflow |
|---|---|---|
| C18 Solid-Phase Extraction (SPE) Cartridges | Fractionation of complex natural extracts based on compound polarity. | Critical for bioassay-guided isolation to simplify the mixture and track activity [32]. |
| DPPH (2,2-Diphenyl-1-picrylhydrazyl) | A stable free radical used to screen for antioxidant activity in extracts and fractions. | A common, reliable initial bioassay for natural products [32]. |
| UHPLC-Q-TOF Mass Spectrometer | High-resolution separation and accurate mass measurement for metabolite profiling and identification. | Generates the high-quality MS and MS/MS data required for dereplication and identification [32] [89]. |
| Global Natural Products Social Molecular Networking (GNPS) | A web-based platform for MS/MS spectral library matching and molecular networking. | The primary tool for dereplication, preventing the rediscovery of known compounds [32]. |
| RDKit | An open-source cheminformatics toolkit. | Used for calculating molecular fingerprints (ECFP, etc.), standardizing structures, and performing similarity searches [86]. |
| Python with NumPy, scikit-learn | Programming environment for data analysis and machine learning. | Essential for computing similarity metrics, building QSAR models, and analyzing the correlation between fingerprints and bioactivity [86]. |
The effective correlation of chemical fingerprints with bioactivity profiles is not a one-size-fits-all endeavor, especially within the structurally diverse realm of natural products. While traditional workhorses like ECFP provide a strong baseline, emerging evidence strongly suggests that researchers should adopt a more nuanced strategy. Fingerprints that capture global, pharmacophoric, or topology-informed features—such as FCFP, MAP4, and pharmacophore fingerprints—often demonstrate superior performance in capturing biological similarity [88] [86]. A hybrid research strategy, which integrates the broad profiling power of LC-MS/MS-based metabolomics with the targeted precision of bioassay-guided isolation, provides the most robust framework for validating these computational tools. By strategically selecting fingerprints and employing integrated experimental workflows, scientists can more effectively navigate the complex chemical space of natural products, accelerating the discovery and development of novel bioactive compounds.
The therapeutic potential of plant-derived natural products (NPs) is immense, with an estimated one-quarter of all modern medicines being plant-based [4]. However, the journey from plant material to clinically viable therapeutics faces significant challenges, including complex metabolite mixtures, low abundance of bioactive compounds, and batch-to-batch variability [90] [91]. For engineered natural products—those produced or optimized through biosynthetic approaches—these challenges necessitate exceptionally rigorous quality control (QC) frameworks. The convergence of advanced analytical technologies and biological validation systems has created unprecedented opportunities for standardizing the quality assessment of these complex therapeutics, ensuring their safety, efficacy, and consistency from laboratory discovery to clinical application [92] [11].
This guide objectively compares the current QC methodologies centered on LC-MS-based characterization and bioassay-driven functional assessment, providing researchers with experimental protocols, performance data, and implementation frameworks. The critical thesis underpinning this analysis is that robust validation of natural product biosynthesis requires an integrated approach that couples detailed chemical profiling with relevant biological activity measurements, creating a comprehensive understanding of both composition and function [11] [91]. As the field moves toward more sophisticated engineering of biosynthetic pathways in heterologous systems [90] [93], the QC strategies must evolve to address both the chemical complexity of the products and their intended biological mechanisms.
Analytical Foundation: LC-MS Technologies for Metabolic Profiling
Liquid chromatography-mass spectrometry (LC-MS) has become the cornerstone technology for quality control of engineered natural products due to its exceptional sensitivity, resolution, and ability to handle complex mixtures [92] [91]. The technological evolution of LC-MS platforms has dramatically enhanced our capacity to characterize the intricate metabolic profiles of natural product preparations, moving beyond simple fingerprinting to comprehensive structural elucidation and quantitative analysis.
Core LC-MS Platforms and Configurations
Table 1: Comparison of LC-MS-Based Metabolomics Platforms for Natural Product Quality Control
| Analytical Platform | Key Strengths | Throughput Capacity | Metabolite Coverage | Implementation Complexity |
|---|---|---|---|---|
| LC-ESI-MS/MS (Targeted) | Excellent sensitivity for known compounds; precise quantification | Medium to High | Limited to pre-defined metabolites | Moderate |
| LC-HRMS (Untargeted) | Comprehensive detection; no prior knowledge required | Low to Medium | Very broad | High |
| Multi-dimensional LC | Superior separation of complex mixtures | Low | Extensive | Very High |
| LC-DIA-MS | Comprehensive MS2 data; reduced missing values | Medium | Broad | High |
| LC-MS with Ion Mobility | Additional separation dimension; isomer differentiation | Medium | Broad | High |
The selection of appropriate LC-MS configurations depends heavily on the specific QC objectives. LC-ESI-MS/MS (electrospray ionization tandem mass spectrometry) provides exceptional sensitivity for detecting known bioactive compounds, as demonstrated in fenugreek studies where it accurately quantified 237 metabolic features including trigonelline and 4-hydroxyisoleucine, which increased by 33.5% and 33.3% respectively during germination [94]. For more exploratory quality assessment, LC-HRMS (high-resolution mass spectrometry) enables untargeted metabolomics without predetermined analytical targets, capturing a broader chemical landscape [92] [91].
The emerging field of multi-dimensional liquid chromatography significantly enhances separation capabilities for complex natural product mixtures, particularly valuable for resolving structurally similar compounds like ginsenoside analogs in Panax species [92]. When combined with advanced scanning modes such as data-independent acquisition (DIA), which captures comprehensive MS2 data without precursor ion selection, these platforms provide deeply informative datasets for quality assessment [92] [11].
Experimental Protocol: LC-MS-Based Metabolomic Profiling
Sample Preparation:
- Harvesting and Stabilization: Rapidly freeze plant materials or cell cultures in liquid nitrogen to halt enzymatic activity and preserve metabolic profiles [91].
- Extraction: Use a two-step liquid-liquid extraction with methyl tert-butyl ether (MTBE)/methanol/water (3:1:1 ratio) for comprehensive coverage of both polar and non-polar metabolites [91]. Alternative: chloroform/methanol/water for specific lipid classes.
- Concentration and Reconstitution: Dry extracts under nitrogen gas and reconstitute in initial mobile phase compatible with LC-MS analysis.
- Quality Controls: Include pooled quality control (QC) samples by combining equal aliquots from all samples to monitor instrument performance [91].
LC-MS Analysis:
- Chromatographic Separation:
- Column: C18 reversed-phase column (100 × 2.1 mm, 1.8 μm)
- Mobile Phase A: Water with 0.1% formic acid
- Mobile Phase B: Acetonitrile with 0.1% formic acid
- Gradient: 5% B to 95% B over 25 minutes, hold 5 minutes, re-equilibrate
- Flow Rate: 0.3 mL/min
- Injection Volume: 5 μL
- Mass Spectrometric Detection:
- Ionization: Electrospray ionization (ESI) in both positive and negative modes
- Mass Analyzer: High-resolution time-of-flight (TOF) or Orbitrap
- Resolution: >30,000 full width at half maximum
- Mass Range: 50-1500 m/z
- Collision Energy: Ramped 10-40 eV for MS/MS fragmentation
Data Processing:
- Use software such as MS-DIAL or XCMS for peak picking, alignment, and normalization [91].
- Apply quality filters: retain features present in ≥80% of QC samples with <30% RSD.
- Perform metabolite identification through spectral matching to databases (GNPS, MassBank) and retention time alignment with authentic standards when available [91].
Figure 1: LC-MS Metabolomic Workflow for Natural Product Quality Control
Bioassay Integration: Functional Assessment of Bioactivity
While LC-MS provides detailed chemical characterization, bioassays deliver the critical functional dimension to quality control, assessing whether engineered natural products maintain their intended biological activity [11] [95]. This integration is particularly vital for complex natural product mixtures where therapeutic effects often emerge from synergistic interactions between multiple compounds rather than single constituents [4] [91].
Cell-Based Bioassay Systems
The selection of biologically relevant cell lines forms the foundation of meaningful bioassay design. For quality control of natural products with known molecular targets, engineered cell lines with specific reporter constructs offer high sensitivity and mechanistic insight. For natural products with complex or poorly understood mechanisms, more phenotypic cell-based assays provide broader activity assessment [11].
Table 2: Cell-Based Bioassay Systems for Natural Product Quality Control
| Bioassay System | Measured Endpoints | Throughput | Relevance to Therapeutic Action | Technical Complexity |
|---|---|---|---|---|
| Cancer Cell Lines (e.g., MCF-7, A549) | Cytotoxicity, Apoptosis, Cell Cycle Arrest | Medium | High for anticancer applications | Moderate |
| Reporter Gene Assays | Pathway activation (e.g., Nrf2, NF-κB) | High | Mechanism-specific | High |
| Primary Cell Cultures | Functional responses in normal cells | Low | High physiological relevance | High |
| Stem Cell-Derived Models | Differentiation, tissue-specific functions | Low | Emerging relevance for complex diseases | Very High |
| Microfluidic Organ-on-Chip | Complex tissue-level responses | Low | High physiological mimicry | Very High |
Case studies demonstrate the power of combining cell-based bioassays with proteomic analysis. For example, treatment of MCF-7 breast cancer cells with Nigella sativa seed extract followed by LC-MS-based proteomics revealed specific protein networks involved in apoptosis and cell cycle regulation, providing both activity confirmation and mechanistic insight [11]. Similarly, green tea extract treatment of A549 lung cancer cells combined with proteomic analysis identified proteins associated with cell migration inhibition [11].
Experimental Protocol: Cell-Based Bioassay with Proteomic Readout
Cell Culture and Treatment:
- Cell Line Selection: Choose biologically relevant cell lines (e.g., MCF-7 for breast cancer, A549 for lung cancer, HCT-116 for colon cancer) [11].
- Culture Conditions: Maintain cells in appropriate medium (RPMI-1640 or DMEM) with 10% fetal bovine serum at 37°C with 5% CO₂.
- Natural Product Treatment:
- Prepare natural product extracts in DMSO (final concentration ≤0.1%).
- Include vehicle control (DMSO only) and positive control (known bioactive compound).
- Treat cells at multiple concentrations (typically 1-100 μg/mL) for 24-72 hours.
Viability and Functional Assessment:
- Cell Viability: Measure using MTT or resazurin reduction assays after 24-48 hours treatment.
- Apoptosis Detection: Assess using Annexin V/propidium iodide staining with flow cytometry.
- Cell Cycle Analysis: Fix cells with 70% ethanol, stain with propidium iodide, and analyze DNA content by flow cytometry.
Sample Preparation for Proteomic Analysis:
- Cell Lysis: Lyse cells in RIPA buffer with protease and phosphatase inhibitors.
- Protein Digestion:
- Reduce with dithiothreitol (5 mM, 30 minutes, 60°C)
- Alkylate with iodoacetamide (15 mM, 30 minutes, room temperature in dark)
- Digest with trypsin (1:50 enzyme-to-protein ratio, 37°C, overnight)
- Peptide Desalting: Desalt using C18 solid-phase extraction cartridges.
LC-MS Proteomic Analysis:
- Chromatography:
- Column: C18 nano-flow column (75 μm × 25 cm, 2 μm particles)
- Gradient: 5-30% acetonitrile over 120 minutes
- Flow Rate: 300 nL/min
- Mass Spectrometry:
- Instrument: Q-Exactive HF-X or similar high-resolution mass spectrometer
- MS1 Resolution: 120,000
- MS2 Resolution: 30,000
- Top N: 20 most intense ions for fragmentation
Data Analysis:
- Protein Identification: Search MS/MS data against appropriate protein databases using Sequest or MaxQuant.
- Quantification: Use label-free quantification based on precursor intensity or spectral counting.
- Pathway Analysis: Perform enrichment analysis using Gene Ontology, KEGG, or Reactome databases.
Comparative Performance Assessment: Case Studies in Natural Product QC
Ginseng Metabolomics and Standardization
Ginseng research provides an exemplary case study in systematic quality control implementation. With the global ginseng market expected to reach $17.7 billion by 2030 [92], robust QC frameworks are essential. LC-MS profiling combined with metabolomics has proven highly effective in discriminating between different ginseng varieties and authenticating commercial products [92].
Advanced analytical approaches include:
- Multi-dimensional LC: Significantly improves separation performance for complex ginsenoside mixtures [92].
- Mass Spectrometry Imaging: Provides spatial distribution information of ginsenosides in plant tissues [92].
- Three-level fingerprinting: Establishes comprehensive structural characterization of ginseng polysaccharides [92].
These approaches enable identification of novel oligosaccharide or monosaccharide markers for differentiation among six root ginseng drugs, demonstrating the power of modern analytical techniques in natural product standardization [92].
Fenugreek Germination Monitoring
A quality-controlled LC-ESI-MS food metabolomics study on fenugreek seeds demonstrated precise tracking of metabolic changes during germination [94]. This approach accurately quantified 237 metabolic features and revealed significant biochemical transformations:
- Bioactive Compounds: Trigonelline and 4-hydroxyisoleucine concentrations increased by 33.5% and 33.3% respectively during 72-hour germination.
- Specialized Metabolites: 9 putative flavonoids increased 1.19- to 2.77-fold; 19 steroid saponins rose by 1.08- to 31.86-fold.
- Primary Metabolites: Showed extreme variability with abundance changes in amino acid derivatives, peptides, and saccharides falling in the 0.09- to 22.25-fold, 0.93- to 478.79-fold and 0.36- to 941.58-fold ranges, respectively [94].
This study exemplifies how targeted and untargeted LC-MS approaches can monitor complex biochemical changes during natural product processing, providing critical quality parameters for standardized preparation.
Table 3: Performance Comparison of QC Approaches for Engineered Natural Products
| QC Approach | Chemical Resolution | Functional Assessment | Batch Consistency Monitoring | Implementation Cost |
|---|---|---|---|---|
| Traditional Phytochemical | Limited to marker compounds | No direct assessment | Moderate | Low |
| LC-MS Metabolomics | Comprehensive chemical profiling | Indirect via compound identification | Excellent | High |
| Bioassay-Guided Fractionation | Correlates chemistry with activity | Direct functional measurement | Challenging | Very High |
| Integrated LC-MS + Bioassay | Comprehensive chemical profiling | Direct functional measurement | Excellent | Very High |
| Proteomic Response Profiling | Indirect chemical assessment | Mechanism-based functional insight | Good | High |
Biosynthesis Validation: From Pathway Engineering to Clinical Translation
The emergence of engineered biosynthesis platforms represents a paradigm shift in natural product production [90] [93]. Transient plant expression systems, particularly agro-infiltration of Nicotiana benthamiana, enable rapid reconstruction of complex plant biosynthetic pathways, producing gram-scale amounts of target compounds within days [90]. This approach successfully reconstituted the 20-step biosynthetic pathway for QS-21, a valuable vaccine adjuvant normally sourced from the bark of the Chilean soapbark tree [90].
Quality Framework for Engineered Biosynthesis
Validating the quality of natural products from engineered systems requires additional considerations beyond traditional sources:
Pathway Fidelity Assessment:
- Intermediate Tracking: Monitor expected biosynthetic intermediates to confirm pathway functionality.
- Byproduct Screening: Identify unexpected shunt products that may indicate pathway inefficiency or enzyme promiscuity.
- Isotopic Labeling: Use ¹³C-labeled precursors to confirm predicted carbon flow through engineered pathways.
Product Characterization:
- Structural Validation: Combine NMR spectroscopy with HRMS to confirm chemical structures of engineered products.
- Stereochemical Analysis: Verify correct stereochemistry using chiral chromatography or optical rotation.
- Impurity Profiling: Identify and quantify process-related impurities specific to the production system.
Figure 2: Quality Control Framework for Engineered Natural Product Biosynthesis
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 4: Essential Research Reagents for Natural Product Quality Control
| Reagent Category | Specific Examples | Function in QC Workflow | Performance Considerations |
|---|---|---|---|
| Chromatography Columns | C18 reversed-phase (1.8-2.2 μm particles); HILIC for polar compounds | Metabolic separation | Particle size affects resolution; surface chemistry determines selectivity |
| Mass Spec Standards | Stable isotope-labeled internal standards (¹³C, ¹⁵N) | Quantification accuracy | Should be added early in extraction to correct for losses |
| Cell Line Models | MCF-7 (breast cancer), A549 (lung cancer), HCT-116 (colon cancer) | Bioactivity assessment | Select based on biological relevance to expected activity |
| Proteomics Reagents | Trypsin/Lys-C mix; TMT isobaric labels; iRT peptides | Protein digestion and quantification | Digestion efficiency affects proteome coverage |
| Extraction Solvents | LC-MS grade methanol, acetonitrile; MTBE for lipidomics | Metabolite isolation | Purity critical to avoid background interference |
| Bioassay Kits | MTT/resazurin viability; Caspase-3 apoptosis; ELISA cytokine kits | Functional assessment | Validate linear range and sensitivity for each application |
The validation of engineered natural products requires increasingly sophisticated quality control ecosystems that integrate advanced analytical technologies with biologically relevant assessment systems. LC-MS-based metabolomics provides unprecedented chemical resolution, while complementary bioassays deliver essential functional validation [11] [91]. The most robust frameworks emerge from the strategic integration of these approaches, creating comprehensive understanding of both composition and biological activity.
As the field advances, several trends are shaping future quality control paradigms: the adoption of multi-omics integration (combining metabolomics, proteomics, and transcriptomics) [11] [93], implementation of microfluidic organ-on-chip platforms for more physiologically relevant bioactivity assessment [4], and utilization of artificial intelligence for pattern recognition in complex datasets [90] [93]. Additionally, the emergence of biosynthetic engineering enables more sustainable production of complex natural products while introducing new quality considerations [90] [93].
For researchers and drug development professionals, successful translation of engineered natural products from laboratory to clinic will depend on implementing these integrated quality systems early in development pipelines. This proactive approach ensures that critical quality attributes are defined based on comprehensive chemical and functional understanding, ultimately accelerating the development of safe, effective, and consistent natural product-based therapeutics.
Conclusion
The synergistic integration of advanced LC-MS technologies and targeted bioassays provides a powerful, validated framework for natural product biosynthesis research. This multi-faceted approach, spanning from foundational exploration to rigorous comparative validation, is crucial for accelerating the discovery and development of novel therapeutics. Future directions will be shaped by emerging trends in synthetic biology, the use of machine learning for data analysis and pathway prediction, and the continuous innovation in LC-MS instrumentation, such as the increased use of ion mobility for isomeric separation. This robust validation paradigm is essential for successfully translating engineered natural products from the laboratory into clinical applications, addressing the urgent need for new therapeutic agents.
References
- 1. Natural products in drug discovery: advances and opportunities [https://pmc.ncbi.nlm.nih.gov/articles/PMC7841765/]
- 2. A Historical Overview of Natural Products in Drug Discovery [https://pmc.ncbi.nlm.nih.gov/articles/PMC3901206/]
- 3. Unlocking the potential of natural products in drug discovery [https://www.drugtargetreview.com/article/141908/unlocking-the-potential-of-natural-products-in-drug-discovery/]
- 4. Natural Products for Drug Discovery in the 21st Century [https://pmc.ncbi.nlm.nih.gov/articles/PMC6032166/]
- 5. Development and Validation of a New LC-MS/MS Method ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12194892/]
- 6. Rationally minimizing natural product libraries using mass ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11915828/]
- 7. Natural products in drug discovery: advances and ... [https://www.nature.com/articles/s41573-020-00114-z]
- 8. probe synthesis, target fishing and protein identification [https://www.nature.com/articles/s41392-020-0186-y]
- 9. Plant and marine-derived natural products: sustainable ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11743463/]
- 10. Toward Hybrid Strategies in Natural Product Drug Discovery [https://chemrxiv.org/engage/chemrxiv/article-details/68a957a823be8e43d69b88e7]
- 11. LC-MS-based proteomics: insights into natural product- ... [https://www.sciencedirect.com/science/article/pii/S1359644625002478]
- 12. A Proteomics Approach to Discovery of Natural Products ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2782881/]
- 13. Accessing Natural Product Biosynthetic Processes by Mass ... [https://pubmed.ncbi.nlm.nih.gov/18706516/]
- 14. Terpenoid Biosynthesis Dominates among Secondary ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8070225/]
- 15. Improving polyketide biosynthesis by rescuing the ... [https://www.nature.com/articles/s41467-025-55973-0]
- 16. Structural insights and rational engineering strategies for ... [https://www.sciencedirect.com/science/article/abs/pii/S0141813025058544]
- 17. An updated catalogue of diverse type II polyketide ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10132072/]
- 18. A strategy for the identification of patterns in ... [https://www.nature.com/articles/s41598-017-11314-w]
- 19. Screening for Expressed Nonribosomal Peptide Synthetases and... [https://pubmed.ncbi.nlm.nih.gov/26831706/]
- 20. Some aspects of bioassay methods in natural-product ... [https://pubmed.ncbi.nlm.nih.gov/9237404/]
- 21. Frontiers | LC - MS based analysis reveal antimicrobial compounds from... [https://www.frontiersin.org/journals/natural-products/articles/10.3389/fntpr.2024.1478361/full]
- 22. Strategy of employing plug-and-play vectors and LC – MS screening to... [https://bioresourcesbioprocessing.springeropen.com/articles/10.1186/s40643-024-00833-w]
- 23. The Future of Metabolic Engineering and Synthetic Biology [https://pmc.ncbi.nlm.nih.gov/articles/PMC3615475/]
- 24. A novel panel of yeast assays for the assessment of thiamin ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9303440/]
- 25. Development and validation of an LC–MS/MS method ... [https://www.nature.com/articles/s41598-025-96764-3]
- 26. Comparative evaluation of four new immunoassays and LC ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12205834/]
- 27. Biosynthetic investigation of γ-lactones in Sextonia rubra ... [https://www.nature.com/articles/s41598-018-37577-5]
- 28. Combining UHPLC and high-resolution MS [https://www.sciencedirect.com/science/article/abs/pii/S0165993614001927]
- 29. Six Key Differentiators Between Liquid Chromatography ... [https://www.chromatographyonline.com/view/six-key-differentiators-between-liquid-chromatography-and-high-resolution-ion-mobility]
- 30. New HPLC, MS, and CDS Products from 2024–2025 [https://www.chromatographyonline.com/view/new-hplc-ms-and-cds-products-from-2024-2025-a-brief-review]
- 31. Comparison of liquid chromatography-high-resolution ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10590993/]
- 32. Incorporating LC-MS/MS Analysis and the Dereplication of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10468906/]
- 33. Bioassay- and liquid chromatography/mass spectrometry ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3798136/]
- 34. Advanced LC×LC–MS Technique for Food Analysis [https://www.chromatographyonline.com/view/advanced-lc-lc-ms-technique-for-food-analysis-insight-into-natural-products]
- 35. Automated Integration and Quality Assessment of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12257450/]
- 36. Offline Two-dimensional Liquid Chromatography-Mass ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12177891/]
- 37. An automatic LC-MS/MS data analysis workflow for herbal ... [https://www.sciencedirect.com/science/article/abs/pii/S094471132400850X]
- 38. Metabolomics - Bindley Bioscience Center - Purdue University [https://bindley.research.purdue.edu/facilities/chemistry-and-biochemistry/metabolomics/]
- 39. Integrated LC-MS/MS system for plant metabolomics - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC3962214/]
- 40. Streamlined identification of strain engineering targets for ... [https://www.nature.com/articles/s41598-023-39661-x]
- 41. Optimal LC-MS metabolomic profiling reveals emergent ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10077522/]
- 42. Bioassays - NCATS Toolkit [https://toolkit.ncats.nih.gov/module/discovery/developing-translational-research-tools/bioassays/]
- 43. Bioassay [https://en.wikipedia.org/wiki/Bioassay]
- 44. Understanding LC/MS-Based Metabolomics: A Detailed ... [https://pubmed.ncbi.nlm.nih.gov/39474988/?utm_source=Other&utm_medium=rss&utm_campaign=None&utm_content=12Su8VomptfNPoFdeXIhFJmeRIxUf-74CII-8UNdnrsgF82kXk&fc=None&ff=20241030143602&v=2.18.0.post9+e462414]
- 45. connecting chemical structure to biological profiles - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC5554950/]
- 46. Bioassay - an overview [https://www.sciencedirect.com/topics/agricultural-and-biological-sciences/bioassay]
- 47. Validation challenges in liquid chromatography-tandem ... [https://www.sciencedirect.com/science/article/abs/pii/S0308814619307319]
- 48. Optimising cell-based bioassays via integrated design of ... [https://www.sciencedirect.com/science/article/pii/S2472555222137056]
- 49. Certain Approaches to Understanding Sources of Bioassay ... [https://www.bioprocessintl.com/assays/certain-approaches-to-understanding-sources-of-bioassay-variability]
- 50. The Traditional Medicine and Modern Medicine from Natural ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6273146/]
- 51. Frontiers | Network Pharmacology Databases for Traditional Chinese Medicine: Review and Assessment [https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2019.00123/full]
- 52. A guide to the use of bioassays in exploration of natural ... [https://pubmed.ncbi.nlm.nih.gov/38185432/]
- 53. Chinmedomics: a potent tool for the evaluation of traditional Chinese medicine efficacy and identification of its active components | Chinese Medicine | Full Text [https://cmjournal.biomedcentral.com/articles/10.1186/s13020-024-00917-x]
- 54. Frontiers | Application of multi-omics in the study of traditional Chinese medicine [https://www.frontiersin.org/journals/pharmacology/articles/10.3389/fphar.2024.1431862/full]
- 55. The Role of LC-MS in Profiling Bioactive Compounds from ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12348853/]
- 56. Matrix Effects—A Challenge toward Automation of ... [https://www.sciencedirect.com/science/article/pii/S1535553510000158]
- 57. from metabolite profiling to pure chemical entities [https://link.springer.com/article/10.1007/s11101-024-09928-w]
- 58. A detailed, robust and comprehensive LC–MS/MS method ... [https://www.sciencedirect.com/science/article/abs/pii/S0926669020302636]
- 59. MATRIX: a platform for cultivation profiling and bioactive ... [https://www.frontiersin.org/journals/natural-products/articles/10.3389/fntpr.2024.1532543/full]
- 60. Advanced Methods for Natural Products Discovery [https://www.mdpi.com/1660-3397/21/5/308]
- 61. Ultra High Throughput Screening of Natural Product ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4521994/]
- 62. Combinatorial Biosynthesis – Potential and Problems - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC1865499/]
- 63. Combinatorial biosynthesis for the engineering of novel fungal natural products | Communications Chemistry [https://www.nature.com/articles/s42004-024-01172-9]
- 64. Combinatorial engineering of betalain biosynthesis pathway in yeast Saccharomyces cerevisiae - PubMed [https://pubmed.ncbi.nlm.nih.gov/37592353/]
- 65. Evolution-guided optimization of biosynthetic pathways - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4273373/]
- 66. De novo biosynthetic pathways: rational design of ... [https://www.sciencedirect.com/science/article/abs/pii/S0958166908000918]
- 67. Heterologous expression facilitates the discovery and ... [https://www.sciencedirect.com/science/article/pii/S2667370323000693]
- 68. A highly efficient heterologous expression platform to facilitate ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12076808/]
- 69. The development of Burkholderia bacteria as heterologous ... [https://pubs.rsc.org/en/content/articlehtml/2025/np/d5np00024f]
- 70. Identification and heterologous expression of an NRPS ... [https://microbialcellfactories.biomedcentral.com/articles/10.1186/s12934-025-02753-6]
- 71. Impact of heterologous expression of Cannabis sativa ... [https://biotechnologyforbiofuels.biomedcentral.com/articles/10.1186/s13068-025-02638-1]
- 72. Analyzing the Challenges and Opportunities Associated With ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12284604/]
- 73. Recent Advances in the Heterologous Expression of ... [https://www.mdpi.com/1660-3397/20/6/341]
- 74. Advances and Challenges in Liquid Chromatography-Mass ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC1781927/]
- 75. Metabolite identification and quantitation in LC-MS ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3278153/]
- 76. Disparate Multi-Dataset Feature Alignment for LC-MS ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10891690/]
- 77. Analytical challenges and current limitations of LCMS [https://www.shimadzu.com/an/service-support/technical-support/technical-information/fundamental/analytical_challenges.html]
- 78. A bioassay method validation framework for laboratory and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10540336/]
- 79. A bioassay method validation framework for laboratory and ... [https://malariajournal.biomedcentral.com/articles/10.1186/s12936-023-04717-w]
- 80. The balance of reproducibility, sensitivity, and specificity of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2537561/]
- 81. Measuring Precision in Bioassays: Rethinking Assay ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5771887/]
- 82. Types Of Immunoassay - And When To Use Them [https://www.quanterix.com/immunoassay-technologies-past-present-and-future/]
- 83. LC-MS/MS in the Clinical Laboratory – Where to From Here? [https://pmc.ncbi.nlm.nih.gov/articles/PMC3052391/]
- 84. The advantages and the limitations of LC-MS/MS and Elisa ... [https://www.biotrial.com/bioanalysis-lc-ms-ms-vs-elisa/]
- 85. LC-MS/MS in clinical chemistry: Did it live up to its promise? [https://www.sciencedirect.com/science/article/abs/pii/S0009898123001936]
- 86. Effectiveness of molecular fingerprints for exploring the ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-024-00830-3]
- 87. Concepts and applications of chemical fingerprint for hit ... [https://www.sciencedirect.com/science/article/abs/pii/S135964462200349X]
- 88. Comparison of molecular fingerprint methods on the basis ... [https://pubmed.ncbi.nlm.nih.gov/19434835/]
- 89. The Role of LC-MS in Profiling Bioactive Compounds from ... [https://www.mdpi.com/2223-7747/14/15/2284]
- 90. Harnessing plant biosynthesis for the development of next ... [https://journals.plos.org/plosbiology/article?id=10.1371/journal.pbio.3002886]
- 91. Metabolomics in the Context of Plant Natural Products ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7023240/]
- 92. Phytochemistry, quality control and biosynthesis in ginseng ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11589329/]
- 93. Systems and synthetic biology for plant natural product ... [https://www.sciencedirect.com/science/article/pii/S2211124725004863]
- 94. Quality-controlled LC-ESI-MS food metabolomics of ... [https://www.sciencedirect.com/science/article/pii/S0963996922014053]
- 95. Natural bioactive products as promising therapeutics [https://www.sciencedirect.com/science/article/pii/S0254629922002551]