The DBTL Cycle in Synthetic Biology: A Complete Guide to Engineering Biology for Drug Development
This article provides a comprehensive exploration of the Design-Build-Test-Learn (DBTL) cycle, the core engineering framework of synthetic biology.
The DBTL Cycle in Synthetic Biology: A Complete Guide to Engineering Biology for Drug Development
Abstract
This article provides a comprehensive exploration of the Design-Build-Test-Learn (DBTL) cycle, the core engineering framework of synthetic biology. Tailored for researchers and drug development professionals, it details the foundational principles of each stage, practical methodologies and applications in biomanufacturing and therapy development, strategies for overcoming bottlenecks through automation and AI, and a critical analysis of how this approach is validating new paradigms in biomedical research. The content synthesizes current advancements to offer a actionable guide for implementing and optimizing DBTL workflows in research and development.
What is the DBTL Cycle? Foundational Principles of Engineering Biology
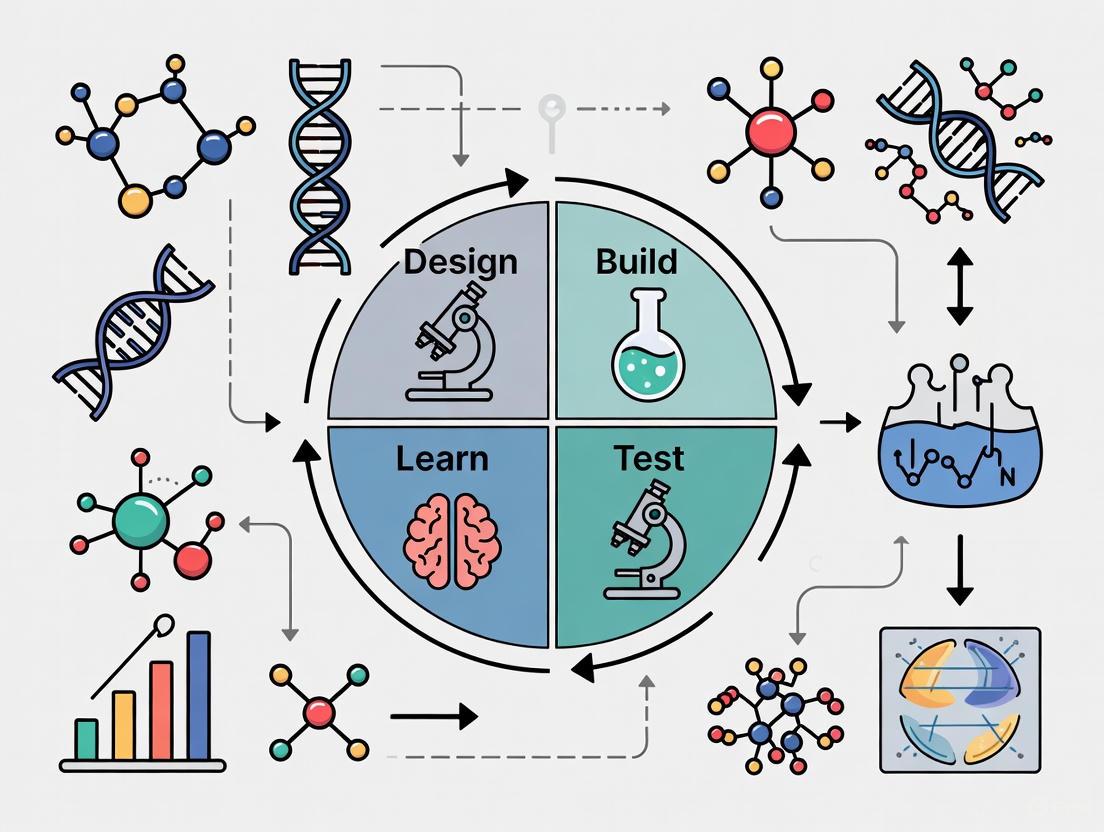
The Design-Build-Test-Learn (DBTL) cycle is a foundational framework in synthetic biology that provides a systematic, iterative methodology for engineering biological systems [1]. This engineering-based approach enables researchers to develop and optimize organisms to perform specific functions, such as producing biofuels, pharmaceuticals, or other valuable compounds [1]. The core principle of DBTL involves cycling through four distinct phases—Design, Build, Test, and Learn—where each iteration incorporates knowledge from previous cycles to refine and improve the biological system until the desired function is achieved [2].
This framework has become increasingly crucial as synthetic biology moves from demonstrating isolated successes to establishing predictable engineering principles. The iterative nature of DBTL allows researchers to navigate the complexity of biological systems, where initial designs rarely perform as expected due to the intricate and often unpredictable interactions within living cells [1]. By applying this structured cycle, synthetic biologists can methodically narrow down possibilities, optimize systems, and gain mechanistic insights into biological function [2] [3].
The Four Phases of the DBTL Cycle
Design Phase
The Design phase constitutes the initial planning stage where researchers define objectives and create a blueprint for the biological system based on a specific hypothesis or learnings from previous cycles [2]. This phase involves:
- Objective Definition: Establishing clear goals for the desired biological function, such as production of a target compound or implementation of a specific genetic circuit [4].
- Part Selection: Choosing appropriate genetic components including promoters, ribosome binding sites (RBS), coding sequences, and terminators [2].
- Circuit Assembly Planning: Determining how selected parts will be assembled into functional genetic circuits or metabolic pathways using standardized methods [2].
- Experimental Protocol Design: Defining precise protocols and success metrics that will be used to evaluate system performance [2].
The Design phase increasingly leverages computational tools and prior knowledge to create more effective initial designs. With advances in machine learning, this phase can now incorporate predictive models that have been trained on large biological datasets, enabling more informed design decisions from the outset [4].
Build Phase
In the Build phase, the theoretical design is translated into physical biological reality through molecular biology techniques [2]. This hands-on component involves:
- DNA Construction: Assembling DNA fragments into complete constructs designed for easy gene construction [1].
- Vector Cloning: Cloning assembled constructs into expression vectors appropriate for the chosen host organism [1].
- Transformation: Introducing the engineered constructs into host organisms such as bacteria, yeast, or mammalian cells [2].
- Verification: Confirming successful construction using methods like colony qPCR, sequencing, or next-generation sequencing (NGS) [1].
Automation of the assembly process significantly reduces the time, labor, and cost of generating multiple constructs, enabling higher throughput with an overall shortened development cycle [1]. The emphasis on modular design of DNA parts allows for the assembly of a greater variety of potential constructs by interchanging individual components [1].
Test Phase
The Test phase focuses on robust data collection through quantitative measurements to characterize the behavior of the engineered system [2]. This critical evaluation stage involves:
- Functional Assays: Performing various assays to measure system performance, such as measuring fluorescence to quantify gene expression, conducting microscopy to observe cellular changes, or performing biochemical assays to measure metabolic output [2].
- High-Throughput Screening: Implementing automated screening methods to efficiently test multiple variants or conditions [5].
- Data Collection: Gathering comprehensive performance data using appropriate analytical techniques and instrumentation [2].
The testing process is crucial for generating the necessary data to evaluate whether the design meets the original specifications and to inform subsequent cycles. Automation of testing significantly improves throughput, reliability, and reproducibility [5].
Learn Phase
The Learn phase represents the analytical component where data gathered during testing is analyzed and interpreted to extract meaningful insights [2]. This stage involves:
- Data Analysis: Processing and interpreting experimental results to determine if the design functioned as expected [2].
- Hypothesis Refinement: Identifying reasons for success or failure and formulating improved hypotheses [2].
- Knowledge Integration: Synthesizing new understanding to inform the next Design phase [2].
- Model Development: Creating or refining computational models to better predict system behavior in future cycles [6].
This phase has traditionally been the most weakly supported in the DBTL cycle, but advances in machine learning and data analytics are increasingly strengthening this critical component [6]. The insights gained here, whether from success or failure, are invaluable for directing subsequent engineering efforts [2].
DBTL in Action: Experimental Applications
Case Study: Engineering Dopamine Production inE. coli
A recent study demonstrated the application of a knowledge-driven DBTL cycle to develop and optimize a dopamine production strain in Escherichia coli [3]. The researchers achieved a dopamine production concentration of 69.03 ± 1.2 mg/L, representing a 2.6 to 6.6-fold improvement over state-of-the-art in vivo dopamine production [3]. Their approach combined upstream in vitro investigation with high-throughput RBS engineering to efficiently optimize the metabolic pathway.
Table 1: DBTL Cycles for Dopamine Production Optimization
| DBTL Cycle | Engineering Target | Key Approach | Outcome |
|---|---|---|---|
| Cycle 1 | Host strain development | Genomic engineering of E. coli for increased L-tyrosine production | Created precursor-optimized chassis |
| Cycle 2 | Enzyme expression balancing | In vitro cell lysate studies to test relative expression levels | Identified optimal HpaBC:Ddc expression ratio |
| Cycle 3 | Pathway fine-tuning | High-throughput RBS engineering to control translation initiation | Achieved 69.03 ± 1.2 mg/L dopamine production |
The methodology employed in this case study highlights how the DBTL framework can be adapted to incorporate mechanistic understanding while efficiently optimizing biological systems [3]. The knowledge-driven approach reduced the number of iterations needed by generating targeted insights before extensive in vivo engineering.
Case Study: Discovering a Novel Anti-Adipogenic Protein
Another research project effectively utilized the DBTL cycle to identify and validate a novel anti-adipogenic protein from Lactobacillus rhamnosus [2]. The researchers systematically narrowed down the active component from the whole bacterium to a single, purified protein through three sequential DBTL cycles:
DBTL Cycle 1: Effect of Raw Bacteria
- Design: Test hypothesis that direct contact with Lactobacillus could inhibit adipogenesis by co-culturing six different strains with 3T3-L1 preadipocytes.
- Build: Culture six bacterial strains and establish 7-day adipogenesis protocol with treatment at various multiplicities of infection (MOI).
- Test: Measure lipid accumulation using Oil Red O staining.
- Learn: Most strains, particularly L. rhamnosus, inhibited lipid accumulation by 20-30%, confirming anti-adipogenic effect [2].
DBTL Cycle 2: Effect of Bacterial Supernatant
- Design: Investigate whether secreted extracellular substances were responsible by treating cells with filtered supernatant.
- Build: Collect supernatant from all six strains and apply at concentrations of 25%, 50%, and 75%.
- Test: Quantify lipid accumulation via Oil Red O staining.
- Learn: Only L. rhamnosus supernatant showed significant, concentration-dependent inhibition (up to 45%), narrowing focus to this strain's extracellular components [2].
DBTL Cycle 3: Effect of Bacterial Exosomes
- Design: Isolate active component by testing exosomes as potential carriers of the active molecule.
- Build: Isolate exosomes from supernatant using centrifugation and Amicon tube with 100k MWCO filter.
- Test: Measure lipid accumulation and analyze expression of adipogenesis-related genes (Ppary, C/ebpa) and AMPK.
- Learn: L. rhamnosus exosomes showed 80% reduction in lipid accumulation and worked through AMPK pathway, confirming exosomes as active component [2].
This case study exemplifies the power of iterative DBTL cycles to systematically narrow down complex biological questions from a broad starting point to a specific mechanistic understanding.
Essential Research Reagents and Tools
Successful implementation of DBTL cycles requires a comprehensive suite of research reagents and tools. The table below details essential materials and their functions in synthetic biology workflows.
Table 2: Key Research Reagent Solutions for DBTL Implementation
| Reagent/Tool | Function | Application Examples |
|---|---|---|
| Expression Vectors | DNA vehicles for gene expression in host organisms | pET system for protein expression; pJNTN for library construction [3] |
| DNA Assembly Systems | Modular DNA construction methods | Golden Gate, Gibson Assembly, Ligation Chain Reaction (LCR) [7] |
| Cell-Free Expression Systems | In vitro transcription and translation | Rapid protein synthesis without cellular constraints; pathway prototyping [4] |
| RBS Libraries | Fine-tune gene expression levels | Ribosome Binding Site variants for metabolic pathway optimization [3] |
| Fluorescent Reporters | Quantify gene expression and protein production | GFP, RFP, and other fluorescent proteins for promoter characterization [5] |
| Analytical Standards | Calibrate measurement equipment | Quantification of target molecules via HPLC or mass spectrometry [7] |
Advanced Methodologies and Protocols
Automated Workflow Integration
The integration of automation into DBTL cycles has revolutionized synthetic biology by enabling higher throughput and improved reproducibility. Biofoundries—structured R&D systems where biological design, construction, testing, and modeling are performed following the DBTL cycle—have emerged as key infrastructure for advanced synthetic biology [8]. These facilities implement an abstraction hierarchy for operations:
- Level 0: Project - The overall research objective to be carried out
- Level 1: Service/Capability - Specific functions required from the biofoundry
- Level 2: Workflow - DBTL-based sequence of tasks
- Level 3: Unit Operations - Actual hardware or software performing tasks [8]
This hierarchical structure enables more modular, flexible, and automated experimental workflows while improving communication between researchers and systems [8].
Machine Learning-Enhanced DBTL
Machine learning (ML) has dramatically transformed the Learn phase of the DBTL cycle and is increasingly influencing the Design phase. Tools like the Automated Recommendation Tool (ART) leverage machine learning and probabilistic modeling to guide synthetic biology in a systematic fashion, without requiring full mechanistic understanding of the biological system [6]. ART uses sampling-based optimization to recommend strains to be built in the next engineering cycle alongside probabilistic predictions of their production levels.
More recently, a paradigm shift from DBTL to LDBT (Learn-Design-Build-Test) has been proposed, where machine learning precedes design [4]. This approach leverages the fact that data that would be "learned" by Build-Test phases may already be inherent in machine learning algorithms, potentially reducing the number of experimental cycles needed.
Diagram 1: DBTL vs. LDBT Cycle Comparison - The paradigm shift from traditional DBTL to machine learning-enhanced LDBT
AI-Powered Autonomous Enzyme Engineering
Recent advances have integrated large language models (LLMs) with biofoundry automation to create fully autonomous enzyme engineering platforms [9]. This generalized platform requires only an input protein sequence and a quantifiable way to measure fitness, enabling:
- Automated Library Design: Using protein LLMs (ESM-2) and epistasis models (EVmutation) to generate diverse, high-quality variant libraries [9]
- Integrated Construction: Implementing HiFi-assembly based mutagenesis methods that eliminate need for intermediate sequence verification [9]
- Continuous Workflow Execution: Dividing protein engineering into seven automated modules that operate without human intervention [9]
This approach has demonstrated substantial improvements in enzyme function, engineering Arabidopsis thaliana halide methyltransferase (AtHMT) for a 90-fold improvement in substrate preference and 16-fold improvement in ethyltransferase activity in just four weeks [9].
Standardized Workflow Visualization
Diagram 2: Detailed DBTL Workflow - The four phases of the DBTL cycle with their key activities
The DBTL framework continues to evolve with technological advancements. Key future directions include:
- Increased Automation: Development of fully autonomous biofoundries that can execute complete DBTL cycles with minimal human intervention [8] [9]
- Enhanced Machine Learning Integration: Deeper incorporation of AI throughout the cycle, from initial design to experimental planning and data interpretation [4] [6]
- Standardization and Interoperability: Establishment of common frameworks and data standards to enable collaboration across biofoundries and research institutions [8] [7]
- Megascale Data Generation: Utilization of cell-free systems and ultra-high-throughput methods to generate the large datasets needed to train more accurate predictive models [4]
The DBTL cycle has proven to be an essential framework for advancing synthetic biology from artisanal practices toward predictable engineering. By providing a structured approach to biological design and optimization, DBTL enables researchers to tackle increasingly complex challenges in bioengineering. As tools and technologies continue to mature, the DBTL framework will undoubtedly remain the core engine driving innovation in synthetic biology and its applications across medicine, manufacturing, and environmental sustainability.
The iterative Design-Build-Test-Learn (DBTL) cycle represents a systematic engineering framework that has become fundamental to advancing synthetic biology. Unlike classical engineering disciplines that utilize well-characterized, man-made components, synthetic biology often relies on partially characterized biological parts implemented within the complex and dynamic environment of living cells [10]. This inherent complexity necessitates an iterative approach to engineering biological systems. The DBTL cycle provides this structured framework, enabling the systematic design of biological systems at the genetic level and the elucidation of genetic design rules [10]. By continuously refining designs based on experimental data, researchers can navigate the vast biological design space to optimize microbial strains for the production of fine chemicals, therapeutics, and sustainable materials [11] [12]. This deep dive explores the core principles, technical methodologies, and transformative applications of each stage within the DBTL cycle, providing researchers and drug development professionals with a comprehensive technical guide.
The Design Stage: In Silico Blueprinting of Biological Systems
The Design stage is the foundational phase where computational tools and biological knowledge converge to create blueprints for genetic constructs. This stage encompasses both biological design—specifying desired cellular functions—and operational design—planning experimental procedures and protocols [10]. The objective is to produce one or more DNA sequences composed of multiple genetic parts that will generate the desired functions in a targeted biological context [10].
Advanced software tools are now integral to this process. For any given target compound, tools like RetroPath enable automated pathway selection, while Selenzyme facilitates enzyme selection [12]. Subsequently, reusable DNA parts are designed with simultaneous optimization of bespoke ribosome-binding sites (RBS) and enzyme coding regions using software such as PartsGenie [12]. These genes and regulatory parts are combined in silico into large combinatorial libraries of pathway designs. A critical step in managing this complexity is the application of Design of Experiments (DoE) methodologies, such as orthogonal arrays combined with Latin squares, to statistically reduce these vast libraries into smaller, representative sets that can be tractably constructed and screened in the laboratory [12]. This compression is substantial; for instance, one documented application achieved a 162:1 compression ratio, reducing 2,592 possible configurations to just 16 representative constructs [12].
Table 1: Key Software Tools for the Design Stage
| Tool Name | Primary Function | Application Context |
|---|---|---|
| RetroPath | Automated pathway selection [12] | Identifying biosynthetic pathways for target compounds |
| Selenzyme | Automated enzyme selection [12] | Selecting optimal enzymes for specified reactions |
| PartsGenie | Design of reusable DNA parts with optimized RBS and coding regions [12] | Creating standardized, optimized genetic components |
| PlasmidGenie | Generation of assembly recipes and robotics worklists [12] | Transitioning from digital design to physical construction |
The Build Stage: From Digital Design to Physical Construct
The Build stage translates digital DNA sequences into physical biological reality through molecular biology techniques, often enhanced by robotic automation [10]. This process involves two main activities: the DNA build, which iteratively assembles the DNA sequence specified in the Design process, and the host build, which involves delivering the genetic construct into the host organism and verifying its presence [10].
The DNA assembly process employs techniques like the ligase cycling reaction (LCR) to combine multiple DNA fragments [12]. Commercial DNA synthesis often provides the starting material, followed by part preparation via PCR [12]. The assembly itself is frequently guided by automated worklists and performed on robotics platforms, ensuring precision and reproducibility. Following assembly, the constructs are transformed into a microbial host, such as Escherichia coli, a workhorse of synthetic biology. The final, crucial step of the Build stage is rigorous verification. This involves quality checks through high-throughput automated plasmid purification, restriction digest analysis by capillary electrophoresis, and definitive sequence verification [12]. This meticulous validation ensures that the physical construct perfectly matches the in silico design before proceeding to costly and time-consuming testing phases.
The Test Stage: Functional Validation of Engineered Systems
In the Test stage, researchers assess whether the biological functions encoded by the designed DNA sequence have been successfully achieved by the host organism [10]. For unicellular production hosts, this typically involves growing the engineered organism under controlled conditions and assaying for the desired function, such as the production of a target chemical [10].
Advanced pipelines automate this process using 96-deepwell plate-based growth protocols [12]. The detection and quantification of the target product and key intermediates are critical. This is achieved through automated sample extraction followed by sophisticated analytical techniques, most notably fast ultra-performance liquid chromatography coupled to tandem mass spectrometry (UPLC-MS/MS) with high mass resolution [12]. The data extraction and processing from these analytical methods are often automated using custom-developed scripts, for example, in the R programming language [12]. This stage generates the quantitative performance data—such as product titer, yield, and rate—that fuels the subsequent Learn stage. For bioprocessing, a significant challenge remains in using these small-volume measurements to predict performance in large-scale fermentation, an area of active research [10].
The Learn Stage: Data-Driven Design Optimization
The Learn stage is the analytical core of the iterative cycle, where measured data is transformed into actionable insights for the next design iteration. This process utilizes statistical methods and machine learning to identify the relationships between observed production levels and the various factors incorporated into the genetic design [12].
For example, in a pathway optimization project for the flavonoid (2S)-pinocembrin, statistical analysis of initial test data identified that vector copy number had the strongest significant positive effect on production titers, followed by the promoter strength upstream of the chalcone isomerase (CHI) gene [12]. Weaker, yet still significant, effects were observed for the promoter strengths of other genes in the pathway. These insights directly informed the constraints for the second design cycle, which focused on a more productive region of the design space [12]. The Learn process can also integrate multi-omics data with metabolic models, such as Flux Balance Analysis (FBA), to identify genetic interventions that further improve titer, rate, and yield of engineered pathways [10]. The cycle is repeated, with each iteration incorporating new knowledge, until the user-defined target function is achieved.
Table 2: Example Quantitative Analysis from a DBTL Cycle for Pinocembrin Production
| Design Factor Analyzed | Impact on Pinocembrin Titer | Statistical Significance (P-value) |
|---|---|---|
| Vector Copy Number | Strongest positive effect [12] | 2.00 x 10⁻⁸ |
| CHI Promoter Strength | Strong positive effect [12] | 1.07 x 10⁻⁷ |
| CHS Promoter Strength | Weaker positive effect [12] | 1.01 x 10⁻⁴ |
| 4CL Promoter Strength | Weaker positive effect [12] | 1.01 x 10⁻⁴ |
| PAL Promoter Strength | Weaker positive effect [12] | 3.06 x 10⁻⁴ |
| Relative Gene Order | Not significant [12] | Not Significant |
Integrated Workflow and a Case Study in Flavonoid Production
The power of the DBTL cycle is fully realized when its stages are integrated into a seamless, automated pipeline. The DOT diagram below illustrates the logical flow and iterative nature of a complete DBTL cycle, highlighting key inputs, processes, and outputs at each stage.
A concrete application of this pipeline targeted the microbial production of the flavonoid (2S)-pinocembrin in E. coli [12]. The pathway involved four enzymes converting L-phenylalanine to pinocembrin. The initial Design stage created a combinatorial library of 2,592 possible configurations, which was reduced via DoE to 16 representative constructs. The Build stage assembled these 16 constructs, all of which were successfully sequence-verified. The Test stage revealed pinocembrin titers ranging from 0.002 to 0.14 mg L⁻¹, and the subsequent Learn stage identified key limiting factors, with vector copy number and CHI promoter strength having the most significant effects [12]. A second DBTL cycle, informed by these findings, focused on a refined design space. This iterative process successfully established a production pathway improved by 500-fold, achieving competitive titers of up to 88 mg L⁻¹ [12]. This case study powerfully demonstrates the rapid optimization capability of an integrated DBTL pipeline.
Table 3: Research Reagent Solutions for a Synthetic Biology DBTL Pipeline
| Reagent / Material | Function in the DBTL Cycle |
|---|---|
| Ligase Cycling Reaction (LCR) | An enzymatic method for assembling multiple DNA fragments into a single construct during the Build stage [12]. |
| DNA Oligonucleotides | Commercially synthesized single-stranded DNA fragments used as building blocks for gene and part assembly [12]. |
| Restriction Endonucleases | Enzymes used for analytical digestion to verify the size and structure of assembled DNA plasmids during quality control [12]. |
| Selected/Multiple Reaction Monitoring (SRM/MRM) | A highly specific mass spectrometry technique for targeted quantification of metabolites, proteins, or pathway intermediates during the Test stage [13]. |
| Mass Distribution Vectors (MDVs) | Data derived from isotope labeling experiments; used with Elementary Metabolite Units (EMU) models for Metabolic Flux Analysis (MFA) in the Learn stage [13]. |
| Ribosome Binding Site (RBS) Libraries | Collections of genetic parts with varying sequences to control the translation initiation rate, a key variable optimized in the Design stage [12]. |
The DBTL cycle has firmly established itself as the central paradigm for the rigorous engineering of biological systems. Its iterative, data-driven nature is essential for managing the complexity inherent in living organisms. The ongoing integration of automation, artificial intelligence (AI), and machine learning (ML) is set to dramatically accelerate this cycle, making it faster, cheaper, and more precise [11]. As these technologies mature and community standards for data and parts sharing solidify, the DBTL framework will be instrumental in tackling global challenges, from developing sustainable manufacturing processes and advanced therapeutics to addressing climate change through carbon sequestration [11]. By deconstructing and mastering each stage of the DBTL cycle, researchers and drug development professionals are poised to unlock the full transformative potential of synthetic biology.
Synthetic biology represents a fundamental shift in the life sciences, moving from a descriptive discipline to an engineering practice focused on the design and construction of novel biological systems. This field is defined by the application of rational principles and formal design processes to biological components [1]. The core premise is that biological systems can be understood as objects endowed with a relational logic between their components not fundamentally different from those designed by computational, chemical, or electronic engineers [14]. This engineering perspective allows researchers to address how and why biological systems work by focusing on the physicochemical implementation of functions in space and time, setting aside exclusive focus on evolutionary origins [14]. The adoption of this mindset is crucial for advancing biotechnology and creating next-generation bacterial cell factories and therapeutic solutions [13].
Core Principles of Rational Biological Design
Defining the Engineering Mindset
The rational engineering approach in synthetic biology is characterized by several key principles. First and foremost is the intent to harness our understanding of biology to build a library of well-understood and characterized modular biological parts, such as genes and proteins, whose functions are predictable and reliable [15]. This approach embraces both the engineering mindset and the unique properties of biological systems, accepting "Nature on its own terms and taking advantage of the parts and tools that Nature has given us, with all of their wonderful idiosyncrasies" [15].
A crucial conceptual framework in this engineering approach is the distinction between techno-logy (rational design) and techno-nomy (the appearance of rational engineering in evolved biological systems) [14]. This parallel mirrors Monod's evolutionary paradox of teleology (finality/purpose) versus teleonomy (appearance of finality/purpose) and provides a valuable interpretive lens for understanding the logic of biological objects without implying the intervention of an actual engineer [14].
The Evolutionary Design Spectrum
Engineering design processes can be understood as existing on an evolutionary spectrum, where the number of variants tested and the number of design cycles needed differentiate approaches [16]. All design methodologies combine variation and selection iteratively, differing primarily in how they leverage exploration (searching design space) and exploitation (using prior knowledge) [16].
Table 1: Engineering Design Approaches in Synthetic Biology
| Design Approach | Key Characteristics | Exploratory Power | Knowledge Leverage | Typical Applications |
|---|---|---|---|---|
| Rational Design | High knowledge exploitation, low variant numbers | Low | High prior knowledge | Systems with well-characterized parts and predictable behavior |
| Directed Evolution | High-throughput variant testing, iterative selection | High | Low to moderate | Enzyme engineering, optimizing complex phenotypes |
| Hybrid Approaches | Combines modeling with experimental testing | Moderate to high | Moderate to high | Pathway optimization, circuit design |
Rational design aims for predictable engineering of biological systems using well-characterized modular parts [15], while directed evolution harnesses the power of evolutionary processes to direct the design of synthetic organisms through high-throughput gene editing and random mutation [15]. These approaches are not mutually exclusive but rather "highly complementary" [15], with the choice depending on the specific problem, available knowledge, and constraints.
The Design-Build-Test-Learn (DBTL) Cycle
The DBTL cycle is the fundamental framework for systematic and iterative development in synthetic biology [1]. This engineering mantra provides a structured approach for engineering biological systems to perform specific functions, such as producing biofuels, pharmaceuticals, or other valuable compounds [1]. The cycle consists of four phases:
- Design: Researchers define objectives for desired biological function and design parts or systems using domain knowledge, expertise, and computational modeling [4].
- Build: DNA constructs are synthesized, assembled into plasmids or other vectors, and introduced into characterization systems including in vivo chassis or in vitro cell-free systems [4].
- Test: Experimental measurement of engineered biological constructs' performance to determine efficacy of design and build phases [4].
- Learn: Analysis of test data compared to design objectives to inform subsequent design rounds through additional DBTL iterations [4].
This framework streamlines biological engineering by providing a systematic, iterative methodology that can be repeatedly applied until desired functionality is achieved [4] [1].
The LDBT Paradigm Shift
Recent advances are transforming the classic DBTL cycle, with some researchers proposing a paradigm shift to "LDBT" - where Learning precedes Design [4]. This reordering is enabled by machine learning algorithms that can leverage large biological datasets to make zero-shot predictions (without additional training) that improve protein functionality [4]. In this model, the data that would be "learned" by Build-Test phases may already be inherent in machine learning algorithms, potentially allowing researchers to "do away with cycling altogether" in some cases and move synthetic biology closer to a Design-Build-Work model that relies on first principles [4].
Quantitative Methodologies and Experimental Protocols
Data-Driven Engineering Decisions
Rational engineering of biological systems requires rigorous quantitative analysis to compare system performance and guide design decisions. Appropriate statistical summaries and visualization methods are essential for interpreting experimental results.
Table 2: Quantitative Comparison of Gorilla Chest-Beating Rates [17]
| Group | Mean Rate (beats/10h) | Standard Deviation | Sample Size (n) |
|---|---|---|---|
| Younger Gorillas (<20 years) | 2.22 | 1.270 | 14 |
| Older Gorillas (≥20 years) | 0.91 | 1.131 | 11 |
| Difference | 1.31 | - | - |
For quantitative data comparison between groups, researchers should employ appropriate graphical representations including back-to-back stemplots (for small datasets with two groups), 2-D dot charts (for small to moderate data across multiple groups), and boxplots (for larger datasets across multiple groups) [17]. Boxplots are particularly valuable as they visualize the five-number summary (minimum, first quartile Q1, median Q2, third quartile Q3, and maximum) and identify potential outliers using the IQR rule [17].
Experimental Workflow Automation
The implementation of automated DBTL cycles is crucial for next-generation bacterial cell factories [13]. Automated biofoundries leverage liquid handling robots and microfluidics to scale the number of reactions and accelerate the DBTL cycle [4]. For example, DropAI leveraged droplet microfluidics and multi-channel fluorescent imaging to screen upwards of 100,000 picoliter-scale reactions [4]. These automated platforms are increasingly incorporating machine learning to create closed-loop design systems where AI agents cycle through experiments [4].
Automated DBTL Workflow for Strain Engineering
This workflow diagram illustrates the iterative nature of the DBTL cycle in an automated biofoundry context, highlighting the continuous refinement process enabled by machine learning and high-throughput experimentation [4] [13].
Research Reagent Solutions Toolkit
Table 3: Essential Research Reagents for Synthetic Biology Workflows
| Reagent/System | Function | Application Examples |
|---|---|---|
| Cell-Free Expression Systems | Protein biosynthesis machinery from cell lysates or purified components for in vitro transcription/translation [4] | Rapid protein production (>1 g/L in <4 h), toxic protein expression, high-throughput variant screening [4] |
| DNA Assembly Reagents | Enzymatic tools for constructing DNA vectors (e.g., USER, LCR, MAGE) [13] | Modular assembly of genetic circuits, pathway construction, genome editing [13] |
| Analytical Standards | Isotopically-labeled internal standards for mass spectrometry [13] | Metabolic flux analysis (MFA), proteomic quantification, targeted metabolomics (SRM/MRM) [13] |
| Machine Learning Models | Pre-trained algorithms for protein design and optimization (e.g., ESM, ProGen, ProteinMPNN) [4] | Zero-shot prediction of protein structures, stability optimization, enzyme engineering [4] |
Machine Learning and Cell-Free Systems Integration
Machine Learning-Enhanced Protein Engineering
Machine learning has become a driving force in synthetic biology, with protein language models demonstrating remarkable capability in zero-shot prediction of protein structures and functions [4]. Sequence-based models like ESM and ProGen are trained on evolutionary relationships between protein sequences and can predict beneficial mutations and infer protein functions [4]. Structure-based tools like ProteinMPNN take entire protein structures as input and predict sequences that fold into that backbone, leading to nearly a 10-fold increase in design success rates when combined with structure assessment tools like AlphaFold [4].
Specialized ML tools have been developed for optimizing specific protein properties. Prethermut predicts effects of single- or multi-site mutations on thermodynamic stability, while Stability Oracle predicts the ΔΔG of protein variants using a graph-transformer architecture [4]. DeepSol employs deep learning to predict protein solubility from primary sequences [4]. These tools enable researchers to eliminate destabilizing mutations or identify stabilizing ones in silico before experimental testing.
Cell-Free Platform Advantages
Cell-free expression systems provide a powerful platform for high-throughput testing of ML predictions [4]. These systems offer multiple advantages including rapid protein production (>1 g/L in <4 hours), ability to produce toxic proteins, scalability from picoliter to kiloliter scales, and compatibility with non-canonical amino acids and post-translational modifications [4]. The open nature of cell-free systems facilitates direct sampling and manipulation of the reaction environment, making them ideal for high-throughput sequence-to-function mapping of protein variants [4].
ML-Cell-Free Integration for Protein Engineering
This integration of machine learning and cell-free testing has demonstrated significant successes in protein engineering. Researchers have coupled in vitro protein synthesis with cDNA display to achieve ultra-high-throughput protein stability mapping of 776,000 protein variants [4]. This vast dataset has been extensively utilized to benchmark various zero-shot predictors for model predictability [4]. Similar approaches have been applied to engineer amide synthetases using linear supervised models trained on over 10,000 reactions from iterative rounds of site saturation mutagenesis [4].
The rational engineering mindset represents a transformative approach to biological design that leverages engineering principles, iterative design cycles, and increasingly powerful computational tools. As machine learning and automation continue to advance, the DBTL cycle is evolving toward more predictive engineering that requires fewer iterations [4]. The integration of machine learning with high-throughput experimental platforms like cell-free systems is creating new paradigms for biological engineering that leverage megascale data generation and modeling [4]. This progression moves synthetic biology closer to established engineering disciplines where reliable outcomes can be achieved through first principles design, ultimately accelerating the development of novel therapeutic solutions and sustainable biotechnologies [16] [13].
Synthetic biology represents a fundamental redefinition of humanity's interaction with biological systems, integrating core principles from biology, engineering, and computer science to design and construct novel biological entities or systematically redesign existing systems [18]. This discipline approaches biology with an engineering mindset, aiming to program biological processes with novel functions starting from fundamental genetic components [18]. The systematic development and optimization of these biological systems are guided by the Design-Build-Test-Learn (DBTL) cycle, an iterative framework that combines experimental techniques with computational modeling [18] [1]. This cycle comprises four distinct stages: the Design phase, where researchers conjecture DNA patterns or cellular alterations to achieve specific objectives; the Build phase, involving physical development of DNA fragments and their incorporation into host cells; the Test phase, where constructs are rigorously evaluated against desired outcomes; and the Learn phase, where results inform subsequent design iterations [18] [1].
The DBTL framework has proven particularly valuable in navigating the inherent complexity of biological systems, which often creates bottlenecks in efficient and predictable engineering [18]. Traditional approaches relying on first-principles biophysical models frequently struggle with non-linear, high-dimensional interactions between genetic parts and host cell machinery, often forcing the engineering process into "ad hoc tinkering" rather than predictive design [18]. The DBTL approach provides a structured methodology to address these challenges, enabling researchers to converge on biological systems with desired functions through systematic iteration [1]. This review examines key historical successes of the DBTL approach, from pioneering genetic circuits to modern AI-enhanced strain engineering, while providing detailed methodological insights and resource guidelines for researchers pursuing DBTL-based synthetic biology campaigns.
The DBTL Cycle: Core Components and Workflow
The DBTL cycle establishes a systematic framework for biological engineering that mirrors design cycles in traditional engineering disciplines. Each phase has distinct objectives, methodologies, and output deliverables that feed into subsequent phases.
Table 1: Core Components of the Traditional DBTL Cycle
| Phase | Key Objectives | Representative Methodologies | Output Deliverables |
|---|---|---|---|
| Design | Define biological objectives; Select genetic parts; Model system behavior | Computational modeling; Parts selection from libraries; Biophysical simulations [18] [4] | DNA sequence designs; System specifications; Predictive models |
| Build | Physical DNA construction; Host cell integration; Library generation | Gene synthesis; CRISPR-Cas9 genome editing; Molecular cloning; DNA assembly [18] [1] | Engineered biological constructs; Plasmid libraries; Transformed strains |
| Test | Characterize system performance; Measure against targets; Identify unintended effects | High-throughput sequencing; Functional assays; 'Omics analyses; Phenotypic screening [18] [19] | Performance metrics; Functional data; Multi-omics datasets |
| Learn | Analyze results; Identify bottlenecks; Inform redesign | Statistical analysis; Machine learning; Pattern recognition; Data integration [18] [19] | Refined hypotheses; Design rules; Optimized parameters for next cycle |
The power of the DBTL framework emerges from its iterative application, where knowledge gained from each cycle informs subsequent iterations, progressively refining the biological system toward desired specifications [18] [1]. This cyclic process continues until the engineered system robustly achieves target functions, whether for basic biological investigation or industrial application. Recent advances have introduced significant modifications to this traditional workflow, including the emerging LDBT paradigm (Learn-Design-Build-Test) that leverages machine learning and large pre-existing datasets to generate initial designs [4].
Diagram 1: The iterative Design-Build-Test-Learn (DBTL) cycle in synthetic biology. The process continues until the engineered biological system achieves the target performance specifications.
Historical Success 1: Pioneering Genetic Circuits - The Toggle Switch and Repressilator
The earliest demonstrations of synthetic biology's engineering potential emerged through the creation of synthetic genetic circuits, with the toggle switch and repressilator representing landmark achievements. These systems established fundamental engineering principles for biological circuit design and demonstrated the effective application of DBTL cycles in constructing programmable cellular behaviors.
The genetic toggle switch constituted one of the first synthetic bistable gene networks, designed to create digital-like memory in living cells [20]. The core design comprised two repressors and two promoters arranged in a mutually inhibitory network - each repressor gene was transcribed from a promoter repressed by the other repressor protein. This configuration enabled the system to stabilize in one of two stable states, with the switch toggling between states in response to specific environmental stimuli. The DBTL process was essential to achieving this functionality: initial designs based on mathematical modeling were built using standard molecular biology techniques, tested through fluorescence and enzymatic assays, and refined through multiple iterations to optimize repressor binding strengths and promoter efficiencies to achieve robust bistability [20].
Concurrently, the repressilator demonstrated the engineering of oscillatory behavior in living cells through a synthetic gene network [20]. This pioneering work implemented a three-repressor negative-feedback loop, where each repressor protein inhibited transcription of the next gene in the cycle. The DBTL cycle guided the optimization of protein degradation rates and transcriptional kinetics necessary to sustain oscillations. Testing required sophisticated single-cell time-lapse microscopy and quantitative fluorescence measurements, with learning phases focusing on matching experimental observations to mathematical models of oscillator dynamics [20]. These foundational circuits established the conceptual and methodological framework for subsequent synthetic biology applications, proving that engineered biological systems could exhibit complex, predictable behaviors.
Table 2: Key Research Reagents for Genetic Circuit Engineering
| Reagent/Category | Specific Examples | Function in DBTL Workflow |
|---|---|---|
| Repressor Proteins | TetR, LacI, CI434 | Core components for transcriptional regulation; Provide inhibition logic for circuit function [20] |
| Promoter Parts | PLtetO-1, Ptrc | Engineered regulatory regions; Control timing and magnitude of gene expression [20] |
| Reporter Genes | GFP, RFP, LacZ | Enable quantitative measurement of circuit dynamics; Facilitate high-throughput screening [20] |
| Molecular Cloning Tools | Restriction enzymes, Ligases, Plasmid vectors | Enable physical construction of genetic designs; Allow modular assembly of genetic parts [1] |
| Inducer Molecules | IPTG, aTc | Provide external control of circuit behavior; Allow experimental perturbation of system dynamics [20] |
Historical Success 2: Microbial Cell Factories for Metabolic Engineering
The application of DBTL cycles to microbial cell factories represents a transformative advancement in metabolic engineering, enabling the production of valuable compounds ranging from pharmaceuticals to biofuels. Corynebacterium glutamicum has emerged as a particularly versatile microbial platform, with systems metabolic engineering leveraging the DBTL framework to optimize production pathways for amino acids and derivative C5 platform chemicals [21].
A representative DBTL campaign for developing L-lysine-derived C5 chemical producers involves several iterative cycles [21]. The initial Design phase employs genome-scale metabolic models (GEMs) to identify gene knockout and overexpression targets that redirect metabolic flux toward desired products while maintaining cellular viability [19] [21]. The Build phase implements these designs using advanced DNA assembly techniques and multiplex automated genome engineering (MAGE) to rapidly construct strain libraries [13]. The Test phase employs analytical methods like mass spectrometry and HPLC to quantify product titers, yields, and productivity, complemented by multi-omics analyses to understand systemic cellular responses [19] [21]. The Learn phase integrates these experimental results with computational models, identifying unforeseen bottlenecks and regulatory interactions that inform the next DBTL cycle [21].
A significant challenge in this domain is the involution of the DBTL cycle, where iterative trial-and-error leads to increased complexity without proportional gains in productivity [19]. This often occurs because removing one metabolic bottleneck reveals new rate-limiting steps, or because production stresses create deleterious metabolic imbalances. Addressing this challenge requires expanding the DBTL framework to incorporate multiscale factors, including bioreactor conditions, media composition, and substrate toxicity, which collectively influence strain performance [19]. Successfully navigating these complexities has enabled the development of C. glutamicum strains producing high-value C5 chemicals at industrial scales, demonstrating the power of systematic DBTL implementation in metabolic engineering [21].
Diagram 2: The DBTL cycle for metabolic engineering of microbial cell factories, showing the potential for cycle involution and AI/ML integration to overcome this challenge.
The Modern Toolkit: AI, Automation, and Cell-Free Systems in Next-Generation DBTL
Recent technological advances are fundamentally reshaping DBTL implementation, with artificial intelligence (AI), laboratory automation, and cell-free systems collectively addressing traditional bottlenecks in biological design cycles. Machine learning (ML) approaches have emerged as particularly powerful tools for navigating biological complexity, offering robust computational frameworks to model non-linear, high-dimensional relationships that challenge traditional biophysical models [18] [19].
The integration of AI/ML is transforming each phase of the DBTL cycle. In the Design phase, protein language models (e.g., ESM, ProGen) enable zero-shot prediction of protein structure and function, while tools like ProteinMPNN and MutCompute facilitate sequence optimization based on structural constraints [4]. For the Learn phase, ML algorithms can identify complex patterns in high-dimensional experimental data, extracting design rules that would remain opaque through conventional analysis [18] [19]. This capability is particularly valuable for avoiding DBTL involution, as ML models can incorporate features from multiple biological scales - from enzymatic parameters to bioreactor conditions - to predict strain performance and identify optimal engineering strategies [19].
Concurrently, cell-free expression systems are dramatically accelerating the Build and Test phases. These platforms leverage transcription-translation machinery from cell lysates or purified components to express proteins without time-intensive cloning steps, enabling rapid testing of thousands of design variants [4]. When combined with microfluidics and automated liquid handling, cell-free systems can screen over 100,000 protein variants in picoliter-scale reactions, generating massive datasets for ML model training [4]. This integration has enabled remarkable engineering achievements, including the development of improved PET hydrolases for plastic degradation and the design of novel antimicrobial peptides [4].
These advances have prompted a fundamental paradigm shift from DBTL to LDBT (Learn-Design-Build-Test), where machine learning on large biological datasets precedes and informs the initial design phase [4]. In this model, pre-trained algorithms generate functional designs that are subsequently validated through rapid cell-free testing, potentially reducing multiple iterative cycles to a single pass. This approach moves synthetic biology closer to the Design-Build-Work model of established engineering disciplines, potentially transforming the efficiency and predictability of biological design [4].
Table 3: Automated and AI-Enhanced Workflows in Modern DBTL Implementation
| Technology Category | Specific Tools/Methods | Impact on DBTL Efficiency |
|---|---|---|
| Protein Language Models | ESM, ProGen, ProteinMPNN | Enable zero-shot prediction of protein structure and function; Accelerate design of novel enzymes [4] |
| Stability Prediction Algorithms | Prethermut, Stability Oracle, DeepSol | Predict effects of mutations on protein stability and solubility; Reduce experimental screening burden [4] |
| Cell-Free Expression Systems | In vitro transcription/translation, cDNA display | Enable rapid testing without cloning; Allow high-throughput screening of 100,000+ variants [4] |
| Automated Strain Engineering | MAGE, automated genome editing | Accelerate construction of genetic variants; Increase reproducibility of build phase [13] |
| Multi-Omics Analytics | RNA-seq, proteomics, metabolomics | Provide comprehensive system characterization; Generate datasets for ML model training [19] |
Essential Research Reagents and Experimental Protocols
Successful implementation of DBTL cycles requires carefully selected research reagents and standardized experimental protocols. This section details key components of the synthetic biology toolkit, with particular emphasis on resources suitable for both academic and industrial research settings.
Table 4: Essential Research Reagent Solutions for DBTL Workflows
| Reagent Category | Specific Examples | Function in DBTL Workflow | Implementation Considerations |
|---|---|---|---|
| DNA Assembly Systems | Golden Gate Assembly, Gibson Assembly, BioBricks | Enable modular construction of genetic designs; Allow rapid part swapping between iterations [1] | Standardization of parts facilitates reproducibility; Automation compatibility varies by method |
| Genome Editing Tools | CRISPR-Cas9, MAGE, USER cloning | Implement precise genetic modifications; Enable multiplexed editing for library generation [13] | Off-target effects require careful validation; Efficiency varies by host organism |
| Analytical Instruments | HPLC, MS, NGS, plate readers | Quantify product titers, sequence constructs, measure performance parameters [19] [21] | Throughput and sensitivity determine testing capacity; Integration with automation platforms varies |
| Cell-Free Systems | E. coli lysates, wheat germ extracts, PURExpress | Provide rapid testing platform for DNA designs; Enable high-throughput screening [4] | Cost per reaction constraints screening scale; Predictive value for in vivo performance requires validation |
| Automation Equipment | Liquid handlers, colony pickers, microfluidics | Increase throughput of build and test phases; Reduce manual labor and improve reproducibility [13] [4] | Significant initial investment; Requires specialized programming and maintenance expertise |
For researchers establishing DBTL workflows, several core experimental protocols have emerged as particularly valuable:
High-Throughput Molecular Cloning Workflow: Modern DBTL implementations employ automated cloning pipelines to increase productivity and reduce bottlenecks [1]. This typically involves in silico design of DNA constructs using standardized parts, followed by automated assembly using restriction enzyme-based or isothermal methods. After assembly, constructs are transformed into host cells, with verification increasingly performed via colony qPCR rather than sequencing to maximize throughput [1]. Automated colony picking systems further enhance throughput by enabling rapid processing of hundreds to thousands of constructs.
Cell-Free Protein Expression Testing: For rapid testing of enzyme variants or genetic circuits, cell-free expression systems provide unparalleled speed [4]. The protocol involves preparing DNA templates via PCR or direct synthesis, setting up transcription-translation reactions with commercial cell-free systems, and quantifying outputs via colorimetric, fluorescent, or mass spectrometry-based assays. This approach can test hundreds to thousands of variants in parallel, generating data within hours rather than days [4].
Multi-Omics Analysis for Learning Phase: Comprehensive system characterization employs integrated transcriptomic, proteomic, and metabolomic analyses [19] [21]. RNA sequencing profiles transcriptional changes, while LC-MS/MS enables protein quantification and metabolite profiling. The resulting datasets are integrated with genome-scale metabolic models to identify bottlenecks and predict beneficial modifications for subsequent DBTL cycles [19].
The historical trajectory of synthetic biology, from pioneering genetic circuits to sophisticated microbial cell factories, demonstrates the transformative power of the DBTL approach as a systematic framework for biological engineering. The iterative application of Design-Build-Test-Learn cycles has enabled researchers to navigate biological complexity and progressively refine synthetic biological systems toward predetermined functions. Current advances in artificial intelligence, laboratory automation, and cell-free testing are further accelerating this paradigm, potentially enabling a fundamental shift from iterative optimization to predictive design. As these technologies mature, the DBTL framework continues to provide the conceptual scaffolding for synthetic biology's progression from empirical tinkering toward true engineering discipline, with profound implications for biomanufacturing, therapeutic development, and basic biological research.
In the realm of synthetic biology and metabolic engineering, the path to optimizing biological systems is notoriously non-linear and complex. The classical Design-Build-Test-Learn (DBTL) cycle has long been the foundational framework for this engineering effort. However, the inherent unpredictability of biological systems—where minor genetic perturbations can lead to disproportionate and unexpected outcomes—demands an iterative, cyclical approach. This technical guide explores the critical role of iteration in navigating biological complexity, drawing upon recent advances in machine learning and high-throughput experimental technologies. Framed within the context of synthetic biology's DBTL cycle, this paper provides researchers, scientists, and drug development professionals with a detailed examination of the methodologies and tools that make iterative cycles a powerful strategy for achieving robust biological design.
Biological systems are characterized by a high degree of complexity and non-linearity. Unlike predictable physical systems, they involve intricate, interconnected networks where components interact in ways that are difficult to model from first principles. A change at the genetic level—such as modifying a promoter strength or enzyme sequence—can have cascading effects on metabolic fluxes, protein-protein interactions, and overall cellular physiology, often in a non-intuitive manner [22]. For instance, combinatorial optimization of a simple linear metabolic pathway can reveal that increasing the concentration of an individual enzyme might deplete its substrate and paradoxically decrease the final product flux, while simultaneously increasing the concentrations of two different enzymes could synergistically boost output [22]. This non-linearity makes one-pass design strategies ineffective.
The synthetic biology community addresses this challenge through the Design-Build-Test-Learn (DBTL) cycle, a systematic, iterative framework for engineering biological systems [4] [22]. The power of this framework lies not in a single execution, but in its repeated application. Each cycle generates data and insights that refine the model and inform the design in the subsequent cycle, progressively closing the gap between predicted and observed system behavior. Recent proposals even suggest a paradigm shift to "LDBT," where machine Learning precedes Design, leveraging pre-trained models on vast biological datasets to make more informed initial designs, thereby accelerating the entire process [4]. This guide will dissect the quantitative evidence for iteration, provide detailed experimental protocols, and visualize the key workflows that underpin this essential approach.
Quantitative Evidence: The Data-Driven Case for Iteration
The theoretical value of iteration is well-established; however, its quantitative impact is best demonstrated through simulated and real-world experimental data. Research using mechanistic kinetic models to simulate DBTL cycles shows that iterative machine learning guidance significantly outperforms single-step optimization.
Table 1: Performance of Machine Learning Models in Successive DBTL Cycles for Metabolic Flux Optimization [22]
| DBTL Cycle | Number of Strain Designs Tested | Best Product Flux (Relative to Wild-Type) | Machine Learning Model Used | Key Learning Outcome |
|---|---|---|---|---|
| Cycle 1 | 50 | ~1.5x | Gradient Boosting / Random Forest | Identified initial correlations between enzyme expression levels and product flux. |
| Cycle 2 | 20 | ~2.8x | Gradient Boosting (retrained) | Refined understanding of non-linear enzyme interactions; exploited synergistic effects. |
| Cycle 3 | 20 | ~3.5x | Gradient Boosting (further retrained) | Discovered optimal global configuration of pathway elements, avoiding local maxima. |
Simulation studies reveal that the choice of machine learning model is crucial, especially in the low-data regime typical of early cycles. Gradient boosting and random forest models have been shown to be robust to training set biases and experimental noise, making them particularly suitable for the initial, data-scarce phases of an iterative campaign [22]. Furthermore, the strategy for allocating resources across cycles is critical. Evidence suggests that when the total number of strains to be built is limited, initiating the process with a larger initial DBTL cycle is more favorable for rapid optimization than distributing the same number of strains equally across all cycles [22]. This initial larger investment generates a richer dataset, providing a stronger foundation for machine learning models to make accurate predictions in subsequent, smaller cycles.
Table 2: Comparative Performance of Machine Learning Models in Simulated DBTL Cycles [22]
| Machine Learning Model | Performance in Low-Data Regime | Robustness to Noise | Robustness to Training Set Bias | Key Application |
|---|---|---|---|---|
| Gradient Boosting | High | High | High | Recommending new strain designs based on predictive distribution. |
| Random Forest | High | High | High | Predicting strain performance from combinatorial libraries. |
| Deep Learning | Lower | Medium | Medium | Requires larger datasets; more powerful in later cycles. |
| Support Vector Machines | Medium | Medium | Lower | Less effective for complex, non-linear pathway interactions. |
Experimental Protocols: Implementing Iterative DBTL Cycles
A successful iterative DBTL workflow requires the integration of precise methodologies across the design, build, test, and learn phases. Below is a detailed protocol for a combinatorial pathway optimization campaign, a common challenge in metabolic engineering.
Protocol: Machine Learning-Guided Combinatorial Pathway Optimization
Objective: To maximize the flux through a synthetic metabolic pathway by iteratively optimizing the expression levels of multiple enzymes.
Materials and Reagents:
- DNA Library: A predefined set of genetic parts (e.g., promoters, ribosomal binding sites) of varying strengths for each pathway gene [22].
- Host Chassis: An appropriate microbial host (e.g., Escherichia coli).
- Cell-Free System (Alternative): A cell-free gene expression platform derived from crude cell lysates or purified components for rapid testing [4].
- Analytical Equipment: HPLC, GC-MS, or spectrophotometric assays for quantifying target product and growth metrics.
- Computational Tools: Software for machine learning (e.g., scikit-learn for gradient boosting) and mechanistic modeling (e.g., SKiMpy for kinetic models) [22].
Methodology:
Learn & Design (LD):
- Learn (L): In the first cycle, this phase may involve initializing a machine learning model with a pre-trained protein language model (e.g., ESM, ProGen) or a foundational kinetic model of the core metabolism [4] [22]. In subsequent cycles, the model is retrained on all accumulated experimental data.
- Design (D): Using an algorithm for recommending new designs, the machine learning model's predictive distribution is sampled. For example, an automated recommendation tool can propose a set of strain designs (e.g., 50 for Cycle 1) by selecting specific promoter-gene combinations from the DNA library that are predicted to maximize product flux, balancing exploration and exploitation [22].
Build (B):
- In Vivo: Synthesize and assemble the DNA constructs designed in the previous phase. Introduce them into the host chassis using high-throughput genome engineering techniques such as MAGE (multiplex automated genome engineering) or golden gate assembly [13].
- In Vitro (Accelerated): For ultra-high-throughput, use a cell-free expression system. Synthesized DNA templates can be directly added to the cell-free reaction for protein expression without time-consuming cloning steps, enabling the testing of thousands of variants in hours [4].
Test (T):
- Cultivate the built strains in a scaled-down format (e.g., 96-well microplates) or use the cell-free reactions directly.
- Measure the key performance indicators (KPIs), primarily the product titer/yield/rate (TYR). Also, collect data on biomass growth and substrate consumption to inform physiological impact [22].
- For cell-free systems, couple the reactions with colorimetric or fluorescent assays for high-throughput sequence-to-function mapping [4].
Learn (L):
- Integrate the new experimental TYR data with the corresponding strain designs (enzyme expression levels) into the growing dataset.
- Retrain the machine learning model (e.g., gradient boosting) on this expanded dataset. Analyze the model to identify feature importance, revealing which enzymes are most influential and uncovering potential non-linear interactions.
- Compare the model's predictions against the new experimental results to assess its accuracy and identify any systematic biases.
Iterate: The cycle (steps 1-4) is repeated, with each round of designs informed by the learnings from the previous one. The number of strains built per cycle can be optimized, often starting with a larger set to seed the model and using smaller, more targeted sets in later cycles [22].
Protocol: Quantifying Microbial Interactions with an Iterative Model
Objective: To infer interaction coefficients between species in a microbial community using relative abundance data.
Materials and Reagents:
- Time-Series Metagenomic Data: Relative abundance data of microbial species across multiple time points.
- Computational Environment: Software for numerical computing (e.g., Python with SciPy).
Methodology:
Problem Framework: The generalized Lotka-Volterra (gLV) model is a standard for modeling species interactions but requires absolute abundance data, which is rarely available. The iterative Lotka-Volterra (iLV) model is designed for widely available relative abundance data [23].
Model Implementation:
- The iLV model incorporates compositional constraints into the gLV framework.
- It uses an iterative optimization strategy that combines linear approximations with nonlinear refinements to enhance the accuracy of parameter estimation for interaction coefficients and growth rates [23].
Iterative Refinement:
- The algorithm iteratively refines its parameter estimates by minimizing the difference between the predicted and observed relative abundance trajectories.
- With each iteration, the model more accurately captures the underlying ecological dynamics, such as competition, predation, and mutualism, from the compositional data.
Validation: The model's performance is validated using simulated datasets with known parameters and applied to real-world datasets (e.g., predator-prey systems, cheese microbial communities) to demonstrate its robustness in predicting species trajectories [23].
Visualizing Iterative Workflows
The following diagrams, generated with Graphviz DOT language, illustrate the core logical relationships and workflows of the iterative cycles discussed in this guide.
Diagram 1: The classic DBTL cycle shows the sequential, iterative process. The "Accelerated Feedback" arrow highlights how modern platforms can short-cycle learning directly back into design.
Diagram 2: The LDBT paradigm positions machine learning at the outset, using pre-existing knowledge to inform the initial design. Testing then generates data that strengthens foundational models for future projects.
Diagram 3: The closed-loop ML workflow shows how data drives model updates, which in turn generate new testable hypotheses, creating a self-improving system.
The Scientist's Toolkit: Essential Research Reagents and Solutions
The effective execution of iterative DBTL cycles relies on a suite of key technologies and reagents that enable high-throughput building and testing.
Table 3: Key Research Reagent Solutions for Iterative Biology
| Tool / Reagent | Function | Application in Iterative Cycles |
|---|---|---|
| Combinatorial DNA Library | A predefined set of genetic parts (promoters, RBS, coding sequences) to systematically vary component properties. | Provides the fundamental design space for exploring genetic variations in each DBTL cycle [22]. |
| Cell-Free Expression System | Protein biosynthesis machinery from cell lysates for in vitro transcription and translation. | Accelerates the Build and Test phases by removing the need for cloning and cell cultivation; enables testing of toxic compounds [4]. |
| Automated Recommendation Tool | An algorithm that uses machine learning models to propose new strain designs for the next cycle. | Automates the Learn-to-Design transition, optimizing the choice of designs to test based on exploration/exploitation trade-offs [22]. |
| Droplet Microfluidics | A technology for creating and manipulating picoliter-scale droplets. | Allows ultra-high-throughput screening by testing >100,000 cell-free or cellular reactions in a single experiment, generating massive datasets [4]. |
| Kinetic Model (e.g., SKiMpy) | A mechanistic model using ODEs to describe metabolic reaction fluxes. | Provides a "digital twin" of the pathway for in silico testing of DBTL strategies and benchmarking machine learning methods [22]. |
| Iterative Lotka-Volterra (iLV) Model | A computational framework to infer microbial interactions from relative abundance data. | Enables iterative learning and model refinement in microbial ecology from commonly available compositional data [23]. |
Iteration is not merely a useful strategy but a fundamental necessity for engineering biological systems. The inherent non-linearity and complexity of life processes mean that success is achieved through a process of progressive refinement, not one-off design. The DBTL cycle, especially when augmented with modern machine learning and accelerated by cell-free testing and biofoundries, provides a structured framework for this iterative learning. As the field evolves towards an LDBT paradigm—where learning from vast datasets precedes design—the cycles will become faster and more efficient. However, the core principle of iteration will remain key, guiding researchers as they navigate the intricate landscape of biological design to develop the next generation of cell factories, therapeutic molecules, and diagnostic tools.
Executing the DBTL Cycle: Methodologies and Applications in Drug Development
The Design-Build-Test-Learn (DBTL) cycle is the fundamental engineering framework that underpins synthetic biology, enabling the systematic and iterative development of biological systems [1]. This cycle begins with the Design phase, where researchers define objectives for a desired biological function and create a conceptual plan for the genetic system intended to achieve it [4]. In traditional DBTL, this phase relies heavily on domain knowledge, expertise, and computational modeling, after which the designed constructs are built, tested, and the resulting data is analyzed to inform the next design round [4]. The Design phase is therefore foundational, setting the trajectory for the entire engineering effort, with its precision directly influencing the number of iterative cycles required to achieve a functional system.
However, a significant paradigm shift is emerging. With recent advances in machine learning (ML), there is a growing proposition to reorder the cycle to LDBT, where "Learning" precedes "Design" [4]. In this model, learning from vast biological datasets via machine learning algorithms directly informs the initial design, potentially enabling functional solutions in a single cycle and moving synthetic biology closer to a "Design-Build-Work" model akin to more established engineering disciplines [4]. This article will explore the tools and methodologies that constitute the modern Design stage, from its traditional computational roots to its current transformation through artificial intelligence.
Computational Foundations & Machine Learning in Biological Design
The design of biological systems has been revolutionized by computational tools. Initially, this relied on parametric models based on biophysical principles, but the field is increasingly dominated by machine learning models that can detect complex patterns in high-dimensional biological data [4]. These tools operate at different levels of biological organization, from individual proteins to entire pathways.
Machine Learning Tools for Protein and Pathway Design
Machine learning provides a powerful opportunity for directly engineering proteins and pathways with desired functions, a task that is challenging due to the complex relationship between a protein's sequence, structure, and function [4]. The following table summarizes key classes of computational tools used in the design process.
Table 1: Machine Learning Tools for Biological Design
| Tool Category | Representative Tools | Primary Function | Application Example |
|---|---|---|---|
| Protein Language Models (Sequence-based) | ESM [4], ProGen [4] | Predict beneficial mutations and infer protein function by learning from evolutionary relationships in protein sequences. | Zero-shot prediction of diverse antibody sequences [4]. |
| Structure-based Design Tools | ProteinMPNN [4], MutCompute [4] | Design new protein sequences that fold into a given backbone (ProteinMPNN) or optimize residues based on the local chemical environment (MutCompute). | Designing stabilized hydrolases for PET depolymerization [4]; designing TEV protease variants with improved activity [4]. |
| Functional Prediction Tools | Prethermut [4], Stability Oracle [4], DeepSol [4] | Predict the effects of mutations on thermodynamic stability (ΔΔG) or protein solubility. | Identifying stabilizing mutations to improve protein expression and function [4]. |
| Pathway Optimization Tools | iPROBE (In vitro Prototyping and Rapid Optimization of Biosynthetic Enzymes) [4] | Uses neural networks on pathway combination data to predict optimal pathway sets and enzyme expression levels. | Improving 3-HB production in Clostridium by over 20-fold [4]. |
The effectiveness of these models, particularly for large-language models, hinges on their scaling properties and in-context learning capabilities, allowing them to be fine-tuned for specialized biological tasks [24]. Furthermore, the rise of multimodal foundation models—trained on diverse data types like DNA, RNA, protein sequences, and structures—promises to further consolidate and enhance design capabilities by providing a more integrated view of biological information [24].
Strategic Workflow for Computational Design
The application of these tools follows a logical sequence from concept to refined design. The workflow begins with objective definition, where the desired biological function (e.g., create a novel enzyme, optimize a metabolic pathway) is clearly specified. Next is tool selection, choosing the appropriate model based on the goal, whether it's de novo protein design, optimizing an existing sequence, or balancing an entire pathway.
The core of the process is in silico design and prediction, where the selected tool is used to generate candidate DNA blueprints. For proteins, this might involve using a structure-based tool like ProteinMPNN to create sequences that fold correctly, followed by a stability predictor like Stability Oracle to filter out destabilizing variants. For pathways, a tool like iPROBE can predict the optimal combination and expression level of enzymes. This step is increasingly powerful with zero-shot predictions, where models can generate functional designs without additional training on specific experimental data, potentially collapsing the number of required DBTL cycles [4]. Finally, the design validation step involves using other computational methods (e.g., AlphaFold for structure prediction) to provide a preliminary check before moving to physical construction.
Diagram 1: Computational design workflow.
Experimental Validation of Designs
Computational designs must be experimentally validated to assess their real-world functionality. This requires transitioning from digital blueprints to physical DNA, a process greatly accelerated by modern high-throughput methods.
High-Throughput Build and Test Platforms
The Build phase involves the physical assembly of the designed DNA constructs. This is often achieved in a high-throughput manner using automated biofoundries, which are facilities that automate design-build-test cycles for synthetic biology [24]. These foundries leverage automation and robotic liquid handling to assemble combinatorial libraries of genetic constructs rapidly, overcoming the limitations of manual, labor-intensive cloning methods [25] [1].
The Test phase then functionally characterizes the built constructs. Cell-free expression systems have emerged as a particularly powerful platform for this, especially for testing protein designs [4]. These systems use protein biosynthesis machinery from cell lysates or purified components to express proteins directly from synthesized DNA templates, bypassing time-intensive cloning and transformation steps in living cells [4]. Key advantages include:
- Speed and Scalability: Protein production exceeding 1 g/L in under 4 hours, scalable from picoliters to kiloliters [4].
- Throughput: Can be combined with droplet microfluidics and imaging to screen over 100,000 variants [4].
- Flexibility: Enable the production of toxic proteins and the incorporation of non-canonical amino acids [4].
Detailed Protocol: Cell-Free Expression and Testing of Protein Variants
This protocol is adapted from efforts that paired cell-free expression with machine learning to screen thousands of protein variants, such as in ultra-high-throughput protein stability mapping [4] and antimicrobial peptide validation [4].
- Objective: To rapidly express and test the function of multiple computationally designed protein variants (e.g., for stability, enzymatic activity, or binding).
- Principle: DNA templates encoding the variants are directly added to cell-free reactions, which provide the necessary components for transcription and translation. The expressed proteins are then assayed for the desired function.
Materials and Reagents:
- DNA Templates: Purified linear DNA fragments or plasmids containing the designed gene variants under a suitable promoter (e.g., T7).
- Cell-Free System: A commercial or laboratory-prepared E. coli or wheat germ extract-based cell-free protein synthesis system.
- Reaction Components: Includes amino acid mixture, energy source (e.g., phosphoenolpyruvate), RNA polymerase, nucleotides, and salts.
- Assay Reagents: Specific to the protein's function (e.g., a fluorogenic substrate for an enzyme, a labeled ligand for a binder).
- Microplates: 96-well or 384-well plates for high-throughput setup.
- Liquid Handling Robot: For automated, precise dispensing of reagents.
- Plate Reader: For detecting colorimetric, fluorescent, or luminescent signals.
Procedure:
- DNA Template Preparation: Dilute each DNA template to a standardized concentration in nuclease-free water.
- Reaction Assembly on Ice: In each well of a microplate, mix the following components:
- Cell-free extract (e.g., 10 µL of E. coli S30 extract)
- DNA template (e.g., 2 µL of 100 nM solution)
- Master mix containing amino acids, energy system, and salts
- Assay reagents (if compatible with the expression system)
- Incubation: Seal the plate to prevent evaporation and incubate at 30°C for 4-6 hours to allow for protein synthesis.
- Functional Assay:
- If the assay reagent was not included during expression, add it now.
- Measure the output signal using a plate reader (e.g., fluorescence, absorbance).
- Include appropriate controls (e.g., no-DNA control, wild-type protein control).
- Data Analysis: Normalize the signals to the controls and rank the variants based on their functional performance.
Data Interpretation: The resulting dataset provides a quantitative or semi-quantitative measure of performance for each designed variant. This data is crucial for the subsequent "Learn" phase, where it is used to refine the computational models and improve the next round of designs, ultimately accelerating the engineering campaign [4].
The Scientist's Toolkit: Essential Research Reagents and Platforms
Success in the Design phase and its subsequent validation relies on a suite of specialized reagents, tools, and platforms.
Table 2: Key Research Reagent Solutions for the Design and Build Phases
| Category | Item | Function in Design/Build Workflow |
|---|---|---|
| DNA Assembly & Editing | Gibson Assembly [26] | An in vitro method for seamlessly assembling multiple overlapping DNA fragments into a larger construct, crucial for building genetic circuits. |
| CRISPR-Cas9 [26] | A genome editing system used in top-down synthesis to introduce designed changes directly into a host organism's genome. | |
| Expression Systems | Cell-Free Expression Systems [4] | A versatile platform for rapid, high-throughput protein synthesis and testing without using living cells. |
| Automated Biofoundries [24] | Facilities that automate the Build and Test stages, enabling high-throughput assembly and screening of vast genetic libraries. | |
| Computational Resources | Protein Language Models (e.g., ESM, ProGen) [4] | Pre-trained AI models used for zero-shot prediction and design of protein sequences with desired functions. |
| Structure Prediction Tools (e.g., AlphaFold) [4] | Deep learning systems that predict the 3D structure of a protein from its amino acid sequence, vital for validating designs. |
The Design stage in synthetic biology is evolving from a knowledge-intensive, iterative process toward a predictive, data-driven engineering discipline. The integration of sophisticated machine learning models, such as protein language models and structure-based design tools, is enhancing our ability to create accurate DNA blueprints from first principles. When this advanced design capability is coupled with high-throughput build and test platforms like cell-free systems and automated biofoundries, the entire DBTL cycle is dramatically accelerated. This progress heralds a future where the design of biological systems is more precise, reliable, and efficient, ultimately unlocking new possibilities in therapeutics, biomanufacturing, and our fundamental understanding of life.
The Build stage is a critical component of the synthetic biology Design-Build-Test-Learn (DBTL) cycle, serving as the physical realization of designed genetic constructs. This stage transforms computational models and in silico designs into tangible biological entities that can be tested and characterized. The process encompasses three fundamental technical operations: DNA synthesis, which creates oligonucleotides from digital sequence data; DNA assembly, which joins these fragments into larger constructs such as genes or pathways; and host transformation, which introduces these constructs into a biological chassis for functional testing. Recent advancements in automation have significantly accelerated this stage, with automated pipetting workstations and integrated experimental equipment now handling substantial portions of these repetitive tasks, thereby reducing manual labor and enhancing overall efficiency [27]. The robustness and fidelity of the Build stage directly determine the quality and reliability of the subsequent Test phase, forming the foundation for iterative biological engineering.
DNA Synthesis: From Sequence to Oligonucleotide
Fundamentals of Oligonucleotide Synthesis
DNA synthesis begins with the chemical production of single-stranded oligonucleotides, typically using solid-phase phosphoramidite chemistry. This robust and automated method involves a four-step chain elongation cycle that adds one nucleotide per cycle to a growing oligonucleotide chain attached to a solid support matrix [28]. The cycle consists of: (1) Deprotection, where a dimethoxytrityl (DMT)-protected nucleoside phosphoramidite attached to a solid support is deprotected with acid to activate it for chain elongation; (2) Coupling, where the next DMT-protected phosphoramidite is added and couples to the activated chain; (3) Capping, where any unreacted 5'-hydroxyl groups are acetylated to render failure sequences inert; and (4) Oxidation, where the phosphite triester linkage between monomers is converted to a more stable phosphate linkage via iodine oxidation [28]. This cyclic process continues until the full-length oligonucleotide sequence is complete, after which the synthesized oligos are cleaved from the solid support and deprotected. The cost of oligonucleotide synthesis generally ranges from $0.05 to $0.17 per base, a cost floor that has remained relatively stable and directly influences the overall expense of gene synthesis [28].
Synthesis Platforms and Automation
Oligonucleotides can be synthesized using either column-based synthesizers or microarray-based synthesizers. Column-based synthesis remains the most widely used method for producing high-quality oligonucleotides for gene synthesis applications. However, emerging technologies are seeking to reduce reagent consumption, improve robustness, and increase throughput to lower the overall cost of synthetic DNA [28]. Automated high-throughput synthesis platforms have become enabling technologies for synthetic biology, allowing for the rapid production of the large oligonucleotide libraries needed for extensive genetic engineering projects. The development of accurate and high-throughput DNA synthesis platforms presents both significant challenges and opportunities for the field [27].
Table 1: Key Research Reagents for DNA Synthesis
| Reagent/Material | Function in Synthesis Process |
|---|---|
| Nucleoside Phosphoramidites | Building blocks (dA, dC, dG, dT) for oligonucleotide chain elongation |
| Solid Support Matrix | Controlled-pore glass (CPG) or polystyrene beads for immobile synthesis |
| Trichloroacetic Acid (TCA) | Deprotection reagent for removing DMT group |
| Tetrazole | Activator for coupling phosphoramidites to the growing chain |
| Acetic Anhydride | Capping reagent for blocking unreacted chains |
| N-Methylimidazole | Catalyst in capping reaction |
| Iodine Solution | Oxidation reagent for stabilizing phosphate backbone |
| Synthesis Columns | Vessels containing solid support for automated synthesizers |
DNA Assembly: From Oligonucleotides to Constructs
DNA Assembly Methodologies
Once oligonucleotides are synthesized, they are assembled into larger DNA constructs through various enzymatic methods. These assembly technologies can be broadly categorized into several groups based on their underlying mechanisms, each with distinct advantages and limitations for specific applications [29].
Restriction Enzyme-Based Methods build upon traditional cloning techniques but with enhanced efficiency and modularity. The Golden Gate method employs type IIs restriction enzymes, which cleave DNA outside their recognition sites to generate unique 4-base overhangs. This allows for multiple fragments to be assembled in a one-pot reaction through cycling between restriction digestion and ligation, with the final product lacking the original restriction sites [29]. Similarly, the BioBrick standard enables sequential assembly of standard biological parts using iterative cycles of restriction digestion and ligation, though it generates scar sequences between parts. Improved versions like the BglBrick system use more efficient and methylation-insensitive enzymes (BglII and BamHI) and produce a 6-nucleotide scar sequence suitable for protein fusions [29].
Sequence Homology-Based Methods utilize longer homologous overlapping regions between parts, avoiding restriction site dependencies. Gibson Assembly uses a one-pot isothermal reaction with three enzymes: T5 exonuclease chews back 5' ends to create single-stranded overhangs; a DNA polymerase fills in gaps; and DNA ligase seals nicks [29]. Sequence and Ligation-Independent Cloning (SLIC) employs T4 DNA polymerase in the absence of dNTPs to generate single-stranded overhangs, with recombination intermediates transformed into cells where endogenous repair machinery completes the assembly [29]. A related method, Seamless Ligation Cloning Extract (SLiCE), uses inexpensive E. coli cell extracts to drive homology-mediated assembly, significantly reducing costs [29].
Table 2: Comparison of Key DNA Assembly Methods
| Method | Mechanism | Key Features | Typical Efficiency | Modularity |
|---|---|---|---|---|
| Restriction Digestion & Ligation | Type II restriction enzymes and DNA ligase | Requires unique restriction sites; generates scars | Variable | Low |
| Golden Gate Assembly | Type IIs restriction enzymes and DNA ligase | One-pot reaction; scarless; standardized overhangs | High for ≤10 fragments | High |
| Gibson Assembly | Exonuclease, polymerase, and ligase | One-pot, isothermal (50°C); seamless | High for ≤15 fragments | High |
| SLIC/SLiCE | Homologous recombination in vitro | Sequence-independent; cost-effective | High for ≤5 fragments | Medium |
| OE-PCR | Polymerase chain reaction with overlapping ends | PCR-based; no enzymes required; seamless | Medium for 2-4 fragments | Low |
Automated Assembly and Biofoundries
Automation has revolutionized DNA assembly by enabling high-throughput construction of genetic variants. Automated pipetting workstations can execute complex assembly protocols with minimal human intervention, dramatically increasing throughput and reproducibility while reducing labor costs and human error [27]. This automation is particularly valuable in biofoundries, integrated facilities that combine laboratory automation with advanced computational workflows to streamline the entire DBTL cycle [27]. The modular design of DNA parts is essential for these automated workflows, as it enables the assembly of a greater variety of potential constructs by interchanging individual components [1]. Automated assembly processes reduce the time, labor, and cost of generating multiple constructs, allowing for an increased throughput with an overall shortened development cycle—a critical advantage for comprehensive pathway optimization and genetic circuit prototyping [1].
Diagram 1: Automated vs manual DNA assembly workflow
Host Transformation and Strain Engineering
Transformation Techniques
The final technical operation in the Build stage involves introducing the assembled DNA constructs into a host organism, typically a microbial chassis such as Escherichia coli. Traditional transformation methods include chemical transformation (using calcium chloride to make cells competent) and electroporation (using an electrical pulse to create temporary pores in cell membranes). For high-throughput workflows, automated transformation and colony picking are essential. Traditional screening methods of transformed bacterial colonies using sterile pipette tips, toothpicks, or inoculation loops are highly prone to human error, labor-intensive, and time-consuming, creating bottlenecks in molecular cloning workflows [1]. Automated systems address these limitations by enabling robust and repeatable processing of hundreds to thousands of transformations simultaneously.
Verification and Quality Control
Following transformation, constructed strains must be verified before proceeding to the Test phase. Verification methods include colony qPCR for rapid screening of positive clones and Next-Generation Sequencing (NGS) for comprehensive sequence validation [1]. In some high-throughput workflows, complete sequence verification may be optional for initial screening rounds, with only functional hits undergoing full sequence analysis. After verification, the sequence-verified constructs are transformed into a production chassis and assayed for function, completing the Build stage and initiating the Test phase of the DBTL cycle [28].
Advanced Integration: LDBT Paradigm and Cell-Free Systems
Recent advances are reshaping the traditional DBTL cycle, particularly through the integration of machine learning and alternative testing platforms. There is a growing proposal for an LDBT paradigm, where "Learning" precedes "Design" [4]. In this model, machine learning provides a new opportunity for directly engineering proteins and pathways with desired functions by leveraging large biological datasets to detect patterns in high-dimensional spaces, enabling more efficient and scalable design [4]. Pre-trained protein language models—such as ESM and ProGen—can perform zero-shot prediction of diverse protein sequences and functions, effectively moving learning to the beginning of the cycle [4].
The adoption of cell-free platforms can further accelerate the Build and Test phases. Cell-free gene expression leverages protein biosynthesis machinery from cell lysates or purified components to activate in vitro transcription and translation [4]. These systems are rapid (>1 g/L protein in <4 hours), enable production of products that might be toxic to live cells, are readily scalable, and can be coupled with assays for high-throughput sequence-to-function mapping of protein variants [4]. When combined with liquid handling robots and microfluidics, cell-free systems can dramatically increase throughput, as demonstrated by platforms like DropAI that can screen upwards of 100,000 picoliter-scale reactions [4]. This approach is particularly valuable for generating the large datasets needed to train machine learning models, creating a virtuous cycle of improvement for synthetic biology.
Diagram 2: LDBT cycle with machine learning and cell-free testing
The Build stage represents a critical juncture in the synthetic biology DBTL cycle where digital designs transition to physical biological entities. DNA synthesis, assembly, and host transformation technologies have advanced significantly through automation, standardized protocols, and integrated workflows. The ongoing development of accurate, high-throughput, and cost-effective DNA synthesis and assembly methods continues to present both challenges and opportunities for the field [27]. As machine learning approaches become increasingly integrated with experimental biology and cell-free systems enable faster prototyping, the efficiency and predictability of the Build stage will continue to improve. These advancements are gradually closing the gap between DNA sequence design and functional implementation, moving synthetic biology closer to a true engineering discipline with transformative potential for therapeutic development, bio-manufacturing, and fundamental biological research.
In the synthetic biology Design-Build-Test-Learn (DBTL) cycle, the "Test" phase is where designed biological constructs are experimentally evaluated to measure their performance and functional outcomes [1]. This stage is critical for generating high-quality, quantitative data that feeds directly into the "Learn" phase, informing the next round of design iterations. High-throughput functional assays and multi-omics characterization represent two powerful, complementary approaches that dominate modern test phase strategies. These methodologies enable researchers to move beyond simplistic, single-measurement outputs to gain comprehensive, systems-level understanding of how engineered genetic modifications affect biological function across multiple molecular layers. The integration of these approaches provides the empirical data necessary to refine biological designs and accelerate the development of optimized strains for therapeutic applications, bio-production, and diagnostic tools.
High-Throughput Functional Assays
Core Principles and Applications
High-throughput screening (HTS) is a cornerstone methodology for the rapid, large-scale testing of biological systems against thousands of experimental conditions or genetic variants [30]. HTS relies on miniaturized formats (e.g., 96-, 384-, or 1536-well plates), automation and robotics for liquid handling and plate reading, and robust detection chemistries to quickly generate functional data at scale [30]. In synthetic biology and drug discovery, HTS functional assays provide a powerful path to identify active compounds, validate drug targets, and accelerate hit-to-lead development [30]. A key application is the functional characterization of gene variants, where high-throughput assays measure effects on macromolecular function to aid in classifying variants of uncertain clinical significance [31]. These assays generate continuous functional scores that help distinguish between functionally normal and abnormal variants, providing critical evidence for pathogenicity assertions [31].
Key Assay Types and Detection Methodologies
HTS encompasses diverse assay formats tailored to different biological questions. Biochemical assays directly measure enzyme activity, receptor binding, or nucleic acid processing in a defined system, providing highly quantitative, interference-resistant readouts [30]. Examples include kinase activity assays to find small-molecule enzymatic modulators within compound libraries [30]. In contrast, cell-based assays capture pathway or phenotypic effects in living cells, using reporter gene assays, viability measurements, or second messenger signaling [30]. Phenotypic screening compares multiple compounds to identify those producing a desired phenotype, such as proliferation assays to determine how a drug affects cell growth [30].
Detection methods are chosen based on sensitivity requirements and assay format:
- Fluorescence polarization (FP) and fluorescence intensity (FI) offer sensitive, homogeneous detection suitable for kinetic measurements [30].
- TR-FRET (Time-Resolved Förster Resonance Energy Transfer) provides reduced background interference.
- Luminescence and absorbance assays offer alternative detection chemistries with different dynamic ranges.
Table 1: Key Performance Metrics for HTS Assay Validation
| Metric | Target Value | Interpretation |
|---|---|---|
| Z'-factor | 0.5 - 1.0 | Excellent assay robustness and reproducibility [30] |
| Signal-to-Noise Ratio (S/N) | Higher is better | Measure of assay window between positive and negative controls |
| Coefficient of Variation (CV) | <10% | Measure of well-to-well and plate-to-plate reproducibility |
| Dynamic Range | Higher is better | Ability to distinguish active vs. inactive compounds [30] |
Experimental Protocol: High-Throughput Variant Functional Characterization
The following protocol outlines a generalized workflow for high-throughput functional characterization of genetic variants, adaptable for enzymes, signaling proteins, or regulatory elements:
Assay Design: Define biological objectives and select appropriate assay format (biochemical vs. cell-based). For clinical variant classification, determine score thresholds that maximize separation between known benign and pathogenic variants [31].
Plate Preparation: Dispense assay components into 384-well or 1536-well plates using automated liquid handlers. Include appropriate controls (positive, negative, blank) distributed across plates.
Reaction Initiation: Add test variants (compound library or genetic variant collection) using pin tools or acoustic dispensers. For enzyme assays, initiate reactions by adding substrate.
Incubation and Kinetic Reading: Incubate plates under controlled temperature conditions. Monitor reaction progress kinetically if measuring residence time or enzyme velocity [30].
Signal Detection: Read endpoint or kinetic signals using appropriate detectors (plate readers equipped with FP, TR-FRET, luminescence, or absorbance capabilities).
Data Processing: Normalize raw data to controls, calculate Z'-factor and other quality metrics to validate assay performance [30]. For variant classification, model score distributions using approaches like multi-sample skew normal mixture models to calculate variant-specific evidence strengths [31].
Multi-Omics Characterization
Integrating Multiple Biological Layers
Multi-omics research involves the simultaneous or integrated analysis of multiple biological data layers—including genomics, transcriptomics, proteomics, epigenomics, and metabolomics—to obtain a comprehensive view of biological systems [32]. Where single-omics approaches provide limited, siloed insights, multi-omics reveals how different molecular layers interact and contribute to overall function or dysfunction. This approach is particularly valuable for understanding complex diseases and engineering biological systems, as disease states often originate within different molecular layers [32]. By measuring multiple analyte types within a pathway, biological dysregulation can be better pinpointed to single reactions, enabling elucidation of actionable targets [32]. The integration of multiomics is driving the next generation of cell and gene therapy approaches, including CRISPR-based therapeutics [32].
Advanced Multi-Omics Applications
Single-cell multiomics represents a cutting-edge approach that enables correlated measurements of genomic, transcriptomic, and epigenomic changes from the same individual cells [32]. This technology allows investigators to determine which molecular changes co-occur within specific cell types, providing unprecedented resolution of cellular heterogeneity in complex tissues. As the field advances, researchers are examining larger fractions of each cell's molecular content alongside larger cell numbers, complemented by technologies like long-read sequencing to examine complex genomic regions and full-length transcripts [32].
Spatial multiomics extends these capabilities by retaining spatial information within tissues, revealing how cellular organization influences function. Liquid biopsy approaches analyze biomarkers like cell-free DNA (cfDNA), RNA, proteins, and metabolites from blood samples, offering non-invasive diagnostic capabilities that are expanding beyond oncology into other medical domains [32].
Table 2: Multi-Omics Technologies and Their Applications in Synthetic Biology
| Omics Layer | Key Technologies | Applications in DBTL Cycle |
|---|---|---|
| Genomics | Whole genome sequencing (WGS), long-read sequencing | Identifying structural variations, verifying construct integration [32] |
| Transcriptomics | RNA-seq, single-cell RNA-seq | Measuring expression levels of engineered pathways [32] |
| Proteomics | Mass spectrometry, intracellular signaling assays | Verifying protein expression, post-translational modifications [32] |
| Epigenomics | ChIP-seq, methylation sequencing | Assessing epigenetic effects of genetic engineering |
| Metabolomics | LC/MS, GC/MS | Profiling metabolic flux through engineered pathways |
Experimental Protocol: Integrated Multi-Omics Workflow
Sample Preparation: Process identical biological samples across multiple omics platforms. For single-cell multiomics, use tissue dissociation protocols that preserve cell viability while enabling partitioning into single-cell suspensions.
Multi-Omic Data Generation:
- Genomics: Extract DNA and perform whole genome sequencing using short-read (Illumina) or long-read (PacBio, Oxford Nanopore) platforms.
- Transcriptomics: Extract RNA and prepare RNA-seq libraries, preferably with strand-specific protocols to accurately determine transcription direction.
- Proteomics: Perform protein extraction followed by tryptic digestion and liquid chromatography-mass spectrometry (LC-MS/MS) analysis.
Data Integration: Use network integration approaches where multiple omics datasets are mapped onto shared biochemical networks to improve mechanistic understanding [32]. In this process, analytes (genes, transcripts, proteins, metabolites) are connected based on known interactions (e.g., transcription factors mapped to regulated transcripts).
Statistical Analysis and Modeling: Apply machine learning and artificial intelligence tools specifically designed for multiomics data to extract meaningful insights [32]. These tools can detect intricate patterns and interdependencies across molecular layers.
Implementation and Integration Strategies
The Scientist's Toolkit: Essential Research Reagent Solutions
Successful implementation of high-throughput functional assays and multi-omics characterization requires specialized reagents and platforms optimized for scale, reproducibility, and sensitivity.
Table 3: Essential Research Reagents and Platforms for High-Throughput Testing
| Reagent/Platform | Function | Application Examples |
|---|---|---|
| Transcreener ADP² Assay | Universal biochemical assay for kinase, ATPase, GTPase, and helicase activity detection [30] | Measuring enzyme activity and inhibitor residence times across diverse target classes |
| Cell-Free Expression Systems | Protein biosynthesis machinery from cell lysates or purified components for in vitro transcription/translation [4] | Rapid protein synthesis without cloning steps; expression of toxic proteins; pathway prototyping |
| MAVE (Multiplex Assays of Variant Effect) Platforms | Systematic measurement of functional effects for thousands of genetic variants in parallel [31] | Clinical variant classification, variant effect maps, deep mutational scanning |
| Single-Cell Multi-Omic Kits | Simultaneous measurement of genomic, transcriptomic, and epigenomic features from same cells [32] | Cellular heterogeneity studies, tumor microenvironment characterization, developmental biology |
| Liquid Biopsy Assay Panels | Analysis of cfDNA, RNA, proteins, and metabolites from blood samples [32] | Non-invasive disease monitoring, early cancer detection, treatment response assessment |
Computational and Analytical Approaches
The massive data output from high-throughput functional assays and multi-omics studies requires sophisticated computational infrastructure and analytical pipelines. For functional assay calibration in clinical variant classification, statistical approaches like multi-sample skew normal mixture models can jointly model score distributions of different variant classes (synonymous, gnomAD, known pathogenic/benign) using constrained expectation-maximization algorithms that preserve the monotonicity of pathogenicity posteriors [31]. For multi-omics data integration, artificial intelligence and machine learning tools are essential for detecting patterns and interdependencies across molecular layers [32]. These include:
- Network integration methods that map multiple omics datasets onto shared biochemical networks based on known interactions [32].
- Data harmonization tools that reconcile disparate datasets with varying formats, scales, and biological contexts [32].
- Multi-analyte algorithmic analysis that simultaneously analyzes genomics, transcriptomics, proteomics, and metabolomics datasets [32].
Workflow Integration with DBTL Cycle
Emerging Trends and Future Directions
The field of high-throughput testing is evolving rapidly with several transformative trends. The integration of machine learning is shifting the traditional DBTL cycle toward an LDBT (Learn-Design-Build-Test) paradigm, where learning from large datasets precedes design [4]. Pre-trained protein language models (e.g., ESM, ProGen) enable zero-shot prediction of protein function and stability, potentially reducing experimental iterations [4]. Cell-free systems combined with microfluidics allow ultra-high-throughput testing of >100,000 protein variants, generating massive datasets for training machine learning models [4]. In multi-omics, the development of purpose-built analysis tools specifically designed for integrated multi-omics data rather than single data types is addressing critical bottlenecks [32]. The clinical application of these technologies is expanding through liquid biopsies and integrated molecular profiling for personalized treatment strategies [32]. Finally, 3D culture systems and organoids are providing more physiologically relevant contexts for high-throughput screening, bridging the gap between traditional cell culture and in vivo models [30].
The Design-Build-Test-Learn (DBTL) cycle is a fundamental framework in synthetic biology for systematically engineering biological systems. The "Learn" stage represents the critical phase where data collected from the "Test" stage is analyzed to extract meaningful insights about the performance of the engineered biological constructs [1]. This analytical process directly informs the subsequent design iterations, enabling researchers to refine their genetic designs, optimize system performance, and progressively approach desired functions such as optimized production of biofuels, pharmaceuticals, or other valuable compounds [1]. In modern synthetic biology, the Learn stage is increasingly being transformed by advanced data analysis techniques and machine learning, which can leverage large datasets to predict beneficial modifications, potentially accelerating the entire engineering process [4]. This phase closes the loop of the DBTL cycle, transforming raw experimental data into actionable knowledge that drives scientific discovery and biotechnological innovation.
Data Analysis Methods for Biological Data
The analysis of biological data in the Learn stage employs a variety of statistical and computational methods, chosen based on the nature of the data and the specific research questions. These methods can be broadly categorized to guide appropriate selection and application.
Table 1: Essential Data Analysis Methods for the "Learn" Stage
| Method | Primary Purpose | Common Applications in Synthetic Biology |
|---|---|---|
| Regression Analysis [33] | Models relationships between variables; predicts outcomes. | Predicting protein expression levels based on promoter strength or codon usage. |
| Factor Analysis [33] | Reduces data dimensionality; identifies latent variables. | Identifying underlying factors (e.g., metabolic burdens) from multivariate readouts. |
| Cohort Analysis [33] | Groups and tracks entities with shared characteristics over time. | Analyzing sub-populations of microbial producers with different genetic stability. |
| Time Series Analysis [33] | Models data points collected sequentially over time. | Monitoring dynamic metabolite production or gene expression profiles in bioreactors. |
| Cluster Analysis [33] | Groups objects so that those in the same group are more similar. | Classifying enzyme variants based on functional performance metrics. |
| Qualitative Analysis [33] | Examines non-numeric data to understand qualities and meanings. | Thematic analysis of literature and existing experimental knowledge for hypothesis generation. |
| Quantitative Analysis [33] | Examines numeric data to identify patterns and quantify relationships. | Statistical analysis of fluorescence levels, yield, growth rate, and other numerical assays. |
Quantitative Data Presentation
Effective presentation of analyzed quantitative data is crucial for interpretation and decision-making. The choice of graphical representation depends on the data's structure and the insights to be communicated.
- Frequency Tables and Histograms: For data spanning a wide range, grouping into class intervals is recommended [34]. The general guidance is to create between 5 and 20 equal-sized intervals. A histogram is then the appropriate graphical representation, providing a visual summary of the distribution of a continuous dataset [35] [34]. The area of each bar in a histogram is proportional to the frequency of the data within that interval [35].
- Frequency Polygons and Line Diagrams: A frequency polygon, derived by joining the midpoints of the tops of the bars in a histogram, is particularly useful for comparing the distributions of two or more different datasets on the same graph [35] [34]. Line diagrams are ideal for displaying time trends, such as the production of a target molecule over the course of a fermentation [35].
- Scatter Diagrams: This graphical method is used to visualize the correlation or relationship between two quantitative variables, for instance, plotting the relationship between gene copy number and protein expression level [35].
Machine Learning and the Paradigm Shift to LDBT
Machine learning (ML) is fundamentally reshaping the Learn stage and the entire DBTL paradigm. By leveraging large biological datasets, ML models can detect complex patterns in high-dimensional spaces, enabling more efficient and predictive design. This has given rise to a proposed paradigm shift from DBTL to LDBT (Learn-Design-Build-Test), where learning precedes design [4]. In this model, pre-trained models are used for zero-shot predictions, generating initial designs that are highly likely to be functional, potentially reducing the number of iterative cycles required.
Machine Learning Approaches in Protein Engineering
Table 2: Machine Learning Models for Biological Design in the "Learn" Stage
| Model Type | Description | Example Tools | Application Example |
|---|---|---|---|
| Sequence-Based Models [4] | Trained on evolutionary relationships in protein sequences. | ESM [35], ProGen [36] | Predicting beneficial mutations for antibody sequences and inferring protein function. |
| Structure-Based Models [4] | Trained on databases of protein structures to associate sequence with 3D structure. | MutCompute, ProteinMPNN [37] | Designing stable hydrolase variants for PET depolymerization [34] or improved TEV protease. |
| Hybrid & Physics-Informed Models [4] | Combines statistical power of ML with explanatory strength of biophysical principles. | Physics-informed ML [33] | Exploring evolutionary landscapes while incorporating energy-based constraints for enzyme engineering. |
| Functional Prediction Models [4] | Focused on predicting specific protein properties from sequence or structure. | Prethermut, Stability Oracle, DeepSol | Predicting the thermodynamic stability (ΔΔG) or solubility of protein variants. |
The following diagram illustrates the flow of information and decision-making in the ML-enhanced LDBT cycle:
ML-Driven LDBT Cycle
Experimental Protocols for High-Throughput Learning
To generate the high-quality, megascale data required for effective machine learning, the Build and Test phases must be highly parallelized and rapid. Cell-free expression systems have emerged as a key technology for this purpose.
Protocol: Cell-Free Protein Expression and Testing
This protocol enables the rapid production and testing of thousands of protein variants without the need for live cells, drastically accelerating the Test phase [4].
1. Key Research Reagent Solutions:
Table 3: Essential Reagents for Cell-Free Protein Synthesis
| Reagent / Material | Function / Description |
|---|---|
| DNA Template | Linear PCR product or plasmid encoding the gene of interest; no cloning required. |
| Cell Lysate | Crude extract from organisms like E. coli, wheat germ, or HEK293 cells, containing the transcription/translation machinery [4]. |
| Energy Solution | Provides ATP, GTP, and other nucleotides and energy sources to drive protein synthesis. |
| Amino Acid Mixture | Contains all 20 canonical amino acids as building blocks for translation. |
| Reaction Buffer | Optimized buffer to maintain pH and provide necessary cofactors like Mg²⁺. |
| Reporting Reagents | Colorimetric or fluorescent substrates (e.g., for an enzymatic assay) to measure function directly in the reaction [4]. |
2. Methodology:
- DNA Template Preparation: Use synthesized linear DNA fragments or plasmids as templates. In high-throughput workflows, DNA can be assembled directly in microtiter plates using automated liquid handling systems [4].
- Cell-Free Reaction Assembly: Combine the following components on ice in a defined order to a final volume of 10-50 µL:
- 5-15 µL of cell lysate (e.g., E. coli S30 extract).
- 1x Energy Solution.
- 1x Amino Acid Mixture (1 mM final concentration per amino acid).
- 1x Reaction Buffer.
- 10-500 ng of DNA template.
- Nuclease-free water to volume.
- Incubation for Protein Synthesis: Incubate the reaction mix at a defined temperature (e.g., 30-37°C for E. coli systems) for 2-6 hours to allow for protein expression. Reactions can be performed in 96-well or 384-well plates [4].
- Functional Testing: Directly assay the expressed protein's function in the same reaction vessel. For an enzyme, add its specific substrate and measure the generation of a colored or fluorescent product over time using a plate reader. This provides a direct readout of activity [4].
- Data Collection and Analysis: Collect kinetic or endpoint data from the plate reader. Analyze the data (e.g., calculating initial rates) and compile it into a dataset linking DNA sequence to functional output for the Learn phase.
The workflow for this high-throughput protocol is visualized below:
Cell-Free Testing Workflow
From Analysis to Design: Closing the Loop
The final and most crucial step of the Learn stage is translating analytical insights and model predictions into concrete design plans for the next DBTL cycle. This involves generating specific, testable hypotheses for new genetic constructs.
Protocol: Designing the Next Construct Library
1. Analyzing Sequence-Function Landscapes: Use the data from cell-free testing to train or refine machine learning models that map DNA or protein sequence to function (e.g., enzymatic activity, solubility) [4]. Models like Stability Oracle can predict the effect of new, untested mutations [4].
2. Prioritizing Mutations and Combinations: - Beneficial Mutations: Identify individual mutations that confer improved properties. - Synergistic Effects: Use the model to predict which beneficial mutations might work well together, avoiding predicted destabilizing combinations. - Exploration vs. Exploitation: Balance the design between variants that are predicted to be highly optimal (exploitation) and those that are more uncertain but could yield valuable new information (exploration).
3. Library Design Strategy: - Saturation Mutagenesis: For a critical residue, design a library that includes all possible amino acid variations at that position. - Combinatorial Assembly: Design a library that combines a selected set of beneficial mutations from different parts of the protein or pathway in different arrangements. - CDS Optimization: Based on expression data, re-design the coding sequence (CDS) to optimize codon usage for the host chassis, improving translation efficiency and protein yield.
The decision-making process for the next design iteration is summarized below:
Next Iteration Design Strategy
The engineering of microbial systems for therapeutic applications represents a frontier in synthetic biology, driven by iterative Design-Build-Test-Learn (DBTL) cycles. This whitepaper details the practical application of these frameworks to develop live biotherapeutic products (LBPs) capable of treating human metabolic diseases. We examine the integration of advanced machine learning to accelerate the DBTL cycle into an LDBT (Learn-Design-Build-Test) paradigm and provide methodologies for constructing and testing engineered microbial chassis. Supported by quantitative data and experimental protocols, this guide serves as a technical resource for researchers and drug development professionals advancing microbial therapeutics.
The DBTL Cycle: Foundation for Engineering Biology
The Design-Build-Test-Learn (DBTL) cycle is a systematic framework central to synthetic biology, enabling the rational development and optimization of biological systems [1]. This iterative process allows researchers to methodically engineer organisms for specific functions, such as producing therapeutic compounds.
- Design: Researchers define objectives and computationally design biological parts or systems. This phase leverages domain knowledge, bioinformatics, and increasingly, machine learning models to predict protein structures and functions [4].
- Build: DNA constructs are synthesized and assembled into vectors (e.g., plasmids) and introduced into a chosen chassis organism, such as E. coli or Lactobacillus [1].
- Test: Engineered constructs are experimentally characterized to measure performance against design objectives using functional assays [1].
- Learn: Data from testing phases are analyzed to inform subsequent design rounds, creating a feedback loop for continuous improvement [4] [1].
High-throughput automation and modular DNA assembly are critical for scaling this process, allowing rapid generation and testing of numerous design variants [1]. A paradigm shift towards LDBT (Learn-Design-Build-Test) is emerging, where machine learning and pre-existing large datasets precede the design phase, enabling more predictive engineering and potentially reducing iterative cycles [4].
Engineering Microbial Therapeutics for Metabolic Diseases
Engineered live biotherapeutic products (LBPs) are being developed to treat diseases by modulating host metabolism directly within the gastrointestinal tract. These recombinant microorganisms are designed to sense, respond to, and rectify pathological metabolic states.
Table 1: Engineered Bacterial Therapeutics for Metabolic Disorders
| Target/Disease | Chassis Organism | Engineered Function | In Vivo Model | Key Outcome |
|---|---|---|---|---|
| Hyperammonemia [38] | Lactobacillus plantarum | Hyperconsumption of ammonia | Ornithine transcarbamylase-deficient mice | Reduced blood ammonia levels |
| Hyperammonemia [38] | E. coli Nissle (SYNB1020) | Overproduction of arginine | Murine model | Reduced blood ammonia |
| Obesity / Metabolic Syndrome [39] | E. coli Nissle | Production of N-acylphosphatidylethanolamine (NAPE) | High-fat diet murine model | Reduced adiposity, insulin resistance, hepatosteatosis |
| Fructose-induced Metabolic Disorders [39] | E. coli Nissle | Conversion of fructose to mannitol | Preclinical models | Protection against metabolic syndrome |
Chassis Selection and Engineering Strategies
Selecting an appropriate microbial chassis is fundamental. Ideal chassis are safe, genetically tractable, and suited to the host environment. Common chassis include:
- Escherichia coli Nissle 1917 (EcN): A well-characterized probiotic with a long history of safe use [39]. Its genetic tractability allows for sophisticated engineering, such as using anaerobic-inducible promoters to activate gene expression specifically in the gut [39].
- Lactobacillus and Bifidobacterium spp.: Gram-positive probiotics often used for their safety profile [38].
Genetic modifications are introduced via methods like CRISPR-Cas9, homologous recombination, and site-specific recombination [38]. For complex pathways, bacterial artificial chromosomes (BACs) can accommodate large DNA inserts [38]. A critical safety consideration is the removal of antibiotic resistance genes used in construction to prevent horizontal gene transfer [39].
Experimental Protocol: In Vivo Testing of an Engineered LBP
Objective: Evaluate the efficacy of an engineered ammonia-consuming Lactobacillus plantarum strain in a murine model of hyperammonemia [38].
Methodology:
- Animal Model Induction: Use ornithine transcarbamoylase-deficient Sparse-fur mice or rats with carbon tetrachloride-induced liver failure.
- Bacterial Administration: Administer the engineered L. plantarum strain orally to the experimental group daily for 3 days. A control group receives a placebo or wild-type strain.
- Sample Collection: Collect blood samples at predetermined time points post-administration.
- Biomarker Analysis: Measure blood ammonia levels using a clinical biochemistry analyzer or specific enzymatic assays.
- Histological Examination: In endpoint studies, analyze brain cortex tissue for astrocyte swelling, a marker of ammonia-induced neurotoxicity.
- Data Analysis: Compare ammonia levels and histological outcomes between treatment and control groups using statistical tests (e.g., t-test) to determine significance.
Optimizing Metabolic Pathways in Microbial Chassis
Optimizing metabolic pathways for high-yield production of therapeutic phytochemicals requires balancing enzyme expression and host metabolism [40]. Cell-free systems are valuable for rapid pathway prototyping.
- In Vitro Prototyping and Rapid Optimization of Biosynthetic Enzymes (iPROBE): This method uses cell-free systems to test pathway combinations and enzyme expression levels. Data from these experiments train a neural network to predict optimal pathway sets for implementation in living cells, leading to >20-fold product yield improvements [4].
- Machine Learning-Guided Optimization: Supervised linear models can be trained on data from iterative site-saturation mutagenesis libraries (e.g., from 10,000+ reactions) to identify enzyme variants with improved properties like stability or activity [4].
Table 2: Market Overview: Synthetic Biology Tools and Technologies
| Product / Technology | Market Size (2029 Projection) | Compound Annual Growth Rate (CAGR) | Primary Drivers |
|---|---|---|---|
| Synthetic Biology Market (Overall) [41] [42] | $31.52 - $61.6 Billion | 20.6% - 26.1% | Demand for bio-based products, increased R&D funding |
| Oligonucleotides & Synthetic DNA [41] | Dominant product segment | Not Specified | Rising demand for synthetic genes for research, diagnostics, therapeutics |
| Genome Engineering [41] | Fastest-growing technology segment | Not Specified | Ease of editing with technologies like CRISPR |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents and Materials
| Reagent / Material | Function / Application | Example Use Case |
|---|---|---|
| Oligonucleotides & Synthetic DNA [41] | Gene synthesis, assembly of genetic constructs, PCR | Building genetic circuits and pathway genes for insertion into a chassis. |
| Chassis Organisms (e.g., EcN, Lactobacillus) [38] [39] | Engineered host for therapeutic functions | Serving as the delivery vehicle for therapeutic genes in the gut environment. |
| CRISPR-Cas9 Systems [41] | Precision genome editing | Knocking in therapeutic genes or creating auxotrophies for biocontainment. |
| Cell-Free Expression Systems [4] | Rapid in vitro protein synthesis and pathway testing | High-throughput prototyping of enzyme variants or metabolic pathways without culturing live cells. |
| Specialized Plasmids & Cloning Kits [1] | Vector systems for gene delivery and expression | Maintaining and expressing therapeutic genetic circuits in the chassis organism. |
Visualizing Workflows and Pathways
The following diagrams illustrate the core workflows and logical relationships in engineering microbial therapeutics.
DBTL Cycle in Synthetic Biology
LDBT Paradigm with Cell-Free Testing
Engineered Bacteria for Metabolic Therapy
Overcoming DBTL Bottlenecks: AI, Automation, and Workflow Optimization
Identifying Common Bottlenecks in Traditional DBTL Workflows
The Design-Build-Test-Learn (DBTL) cycle is a fundamental engineering framework in synthetic biology, enabling the systematic development of engineered biological systems [1]. This iterative process aims to design and optimize biological entities, such as microorganisms, to perform specific functions like producing biofuels, pharmaceuticals, or other valuable compounds [1]. Despite its structured approach, the traditional DBTL cycle often faces significant bottlenecks that hinder its efficiency and predictability. These limitations arise primarily from the inherent complexity of biological systems, where introducing foreign DNA into a host cell can lead to unpredictable outcomes due to non-linear, high-dimensional interactions between genetic parts and host cell machinery [18]. This complexity forces researchers to test numerous permutations, making the process laborious and time-consuming [1] [18].
The DBTL cycle begins with the Design phase, where DNA patterns or cellular alterations are conceived to achieve specific objectives. The Build phase involves the physical construction of DNA fragments and their insertion into a host cell. The Test phase assesses how well the engineered system performs against desired outcomes, and the Learn phase uses these results to refine and improve the next design iteration [18]. While this framework is conceptually sound, its practical implementation often deviates from rational design into a "regime of ad hoc tinkering" because biological systems frequently violate assumptions about part modularity that are critical for predictable engineering [18]. This guide examines the common bottlenecks in each stage of the DBTL cycle, provides quantitative insights into their impacts, and outlines emerging solutions that leverage automation and artificial intelligence (AI).
Bottlenecks in the Design Phase
The Design phase involves planning genetic constructs or cellular modifications to achieve desired functions. Bottlenecks in this stage significantly impact all subsequent steps in the DBTL cycle.
Predictive Modeling Limitations
Traditional DBTL workflows rely heavily on first-principles biophysical models to predict biological system behavior. However, these models struggle with the non-linear interactions and high-dimensional design spaces characteristic of biological systems [18]. The vast combinatorial space of potential genetic configurations makes comprehensive exploration impractical. For example, a relatively simple pathway with four genes can generate 2,592 possible configurations when considering variables like promoter strength, ribosome binding sites, and gene order [12]. This complexity often forces researchers to make suboptimal design choices based on incomplete information.
Knowledge Gaps in Biological Systems
A fundamental challenge in biological design is the incomplete understanding of how genetic sequences translate into functional outcomes within living systems. Biological systems operate through intricate networks of interactions that are not fully captured by current models [43] [18]. This knowledge gap becomes particularly evident when designing complex biological systems such as nonribosomal peptide synthetases (NRPS) and polyketide synthases (PKS), where the structural complexity and tightly coordinated interactions between domains make reprogramming these systems exceptionally challenging [44].
Quantitative Impact of Design Limitations
Table 1: Quantitative Impact of Design Phase Bottlenecks
| Bottleneck Category | Specific Challenge | Experimental Impact | Data Source |
|---|---|---|---|
| Combinatorial Complexity | 4-gene pathway optimization | 2,592 possible configurations requiring evaluation | [12] |
| Design Space Reduction | Application of Design of Experiments (DoE) | Compression ratio of 162:1 (2,592 to 16 constructs) | [12] |
| Predictive Modeling | Traditional biophysical models | Struggle with non-linear, high-dimensional interactions | [18] |
Bottlenecks in the Build Phase
The Build phase encompasses the physical construction of genetic designs and their implementation in host organisms. This stage has traditionally been hampered by manual, low-throughput techniques.
Manual Laboratory Techniques
Traditional Build processes rely heavily on manual manipulation by researchers, including techniques such as pipetting, colony picking, and transformation [45]. These methods are not only time-consuming and labor-intensive but also introduce significant variability and human error [1] [45]. The reliance on manual techniques creates a fundamental throughput bottleneck, as noted by researchers: "Synthetic biology is not limited by technology anymore. It's limited by a throughput bottleneck, because at the end of the day, a researcher still has only two hands and a finite number of number of hours to spend in a lab" [45].
DNA Synthesis and Construction Limitations
The cost and time required for DNA synthesis presents another critical bottleneck. In high-throughput protein engineering workflows, DNA synthesis can account for over 80% of the total expense [46]. Traditional gene synthesis methods often involve lengthy processes including colony picking, sequencing, and verification, which dramatically slow down the Build phase. While automated DNA assembly systems exist, they often require significant capital investment and specialized expertise, placing them out of reach for many academic laboratories [46].
DNA Construction Cost Analysis
Table 2: DNA Construction Cost Analysis in Build Phase
| Cost Factor | Traditional Workflow | Optimized Workflow (DMX) | Improvement | Data Source |
|---|---|---|---|---|
| DNA Synthesis Cost | >80% of total expense | 5-8 fold reduction | ~85% cost reduction | [46] |
| Cloning Accuracy | Requires sequencing verification | ~90% accuracy with suicide gene (ccdB) system | Eliminates sequencing step | [46] |
| Gene Variant Recovery | Low-throughput | 78% recovery from oligo pool (1,500 designs) | High multiplexing capability | [46] |
Bottlenecks in the Test Phase
The Test phase involves characterizing and evaluating the performance of built biological systems. This stage often becomes a major bottleneck due to low-throughput analytical methods.
Low-Throughput Screening Methods
Traditional screening methods in synthetic biology rely on manual techniques using sterile pipette tips, toothpicks, or inoculation loops to handle transformed bacterial colonies [1]. These approaches are highly prone to human error and do not scale effectively for evaluating large libraries of biological variants [1]. Even when more sophisticated analytical methods are employed, such as liquid chromatography-mass spectrometry, they often lack the throughput necessary to keep pace with the Build phase, particularly when dealing with thousands of variants [12] [46].
Data Generation and Analysis Challenges
The Test phase generates complex datasets that require sophisticated analysis, but traditional workflows often lack automated data processing pipelines. Researchers must manually process and interpret results, which becomes impractical with large experimental datasets [12] [18]. Without standardized analytical protocols, data quality and consistency can vary, complicating the Learn phase and hindering the iterative improvement process [46]. The absence of integrated data management systems further exacerbates these challenges, as experimental parameters and results are often recorded in disconnected formats.
Bottlenecks in the Learn Phase
The Learn phase involves analyzing experimental data to extract insights that will inform the next Design cycle. This critical translation step faces several significant challenges.
Data Integration and Interpretation Barriers
A primary bottleneck in the Learn phase is the difficulty in extracting meaningful design principles from complex experimental data. Biological systems exhibit multivariate interactions where multiple factors influence outcomes in non-additive ways [12]. Traditional statistical methods often fail to capture these complex relationships, particularly when working with limited datasets. Furthermore, the lack of standardized data formats and experimental metadata makes it challenging to compare results across different cycles or research groups, limiting the accumulation of knowledge [18].
Limited Feedback to Design
In traditional DBTL workflows, the feedback from experimental results to subsequent design iterations is often slow and incomplete. The manual nature of data analysis and interpretation creates delays, while cognitive biases may lead researchers to focus on familiar design paradigms rather than exploring novel solutions [12] [18]. This limitation is particularly evident in the context of the "black box" problem of biological complexity, where the mechanisms underlying successful designs remain obscure, making it difficult to systematically apply these insights to new problems [18].
Case Study: Automated DBTL for Flavonoid Production
A published case study demonstrates both the bottlenecks in traditional DBTL workflows and how automation addresses them. Researchers applied an automated DBTL pipeline to optimize the microbial production of the flavonoid (2S)-pinocembrin in Escherichia coli [12].
Experimental Protocol and Workflow
The study implemented a highly automated DBTL pipeline with the following key methodological components:
Pathway Design: Four enzymes (PAL, CHS, CHI, 4CL) were selected to convert L-phenylalanine to (2S)-pinocembrin. A combinatorial library was designed with variations in vector copy number, promoter strength, and gene order, generating 2,592 possible configurations [12].
Design of Experiments (DoE): Statistical reduction using orthogonal arrays combined with a Latin square for gene arrangement compressed the library from 2,592 to 16 representative constructs (compression ratio of 162:1) [12].
Automated Assembly: Robotic platforms performed ligase cycling reaction for pathway assembly, with automated quality control through capillary electrophoresis and sequence verification [12].
High-Throughput Testing: Automated 96-deepwell plate growth protocols were implemented, followed by quantitative analysis using ultra-performance liquid chromatography coupled to tandem mass spectrometry [12].
Data Analysis and Learning: Custom R scripts performed statistical analysis to identify factors significantly influencing production titers, informing the second DBTL cycle [12].
Quantitative Results and Workflow Diagram
After two DBTL cycles, the optimized pathway achieved a 500-fold improvement in (2S)-pinocembrin production, with titers reaching 88 mg L⁻¹ [12]. Statistical analysis revealed that vector copy number had the strongest significant effect on production levels (P value = 2.00 × 10⁻⁸), followed by CHI promoter strength (P value = 1.07 × 10⁻⁷) [12].
Diagram 1: Automated DBTL workflow for flavonoid production, showing two cycles with statistical learning
Enabling Technologies and Solutions
Emerging technologies are addressing DBTL bottlenecks through automation, artificial intelligence, and advanced molecular biology techniques.
Laboratory Automation and Robotics
Integrated robotic systems are transforming DBTL workflows by enabling high-throughput experimentation. For example, the UCSB BioFoundry employs custom-designed robotic workflows for synthetic biology that can operate without human intervention, allowing for miniaturized cultivation of cells and automated sampling, testing, and analysis [45]. These systems provide the "experimental firepower of a mid-size biotechnology or pharmaceutical company" to academic researchers, dramatically increasing throughput while reducing human error [45].
AI and Machine Learning Integration
Artificial intelligence and machine learning are playing an increasingly crucial role in overcoming DBTL bottlenecks. AI-driven tools can rapidly screen and predict enzyme performance, design optimal biological parts, and guide experimental planning [47] [43]. The integration of AI creates a powerful synergy with synthetic biology—while synthetic biology generates large datasets for training AI models, these models in turn inform and optimize biological design [18]. This mutually reinforcing relationship accelerates the entire DBTL cycle, potentially reducing development timelines from years to months [18].
Innovative Molecular Biology Methods
Novel molecular biology techniques are addressing specific bottlenecks in the Build and Test phases:
Semi-Automated Protein Production (SAPP): This workflow achieves a 48-hour turnaround from DNA to purified protein with only about six hours of hands-on time, using sequencing-free cloning with a suicide gene system for high cloning accuracy (~90%) and miniaturized parallel processing in 96-well plates [46].
DMX DNA Construction: This method reduces DNA synthesis costs by 5-8 fold through construction of sequence-verified clones from inexpensive oligo pools, using isothermal barcoding and nanopore sequencing to recover multiple gene variants from a single pool [46].
Research Reagent Solutions
Table 3: Key Research Reagents and Platforms for DBTL Workflows
| Reagent/Platform | Function | Application in DBTL | Source/Example |
|---|---|---|---|
| Ligase Cycling Reaction (LCR) | DNA assembly method | Automated pathway construction in Build phase | [12] |
| Gibson SOLA Platform | On-demand DNA/mRNA synthesis | Accelerates DNA construction in Build phase | [48] |
| Golden Gate Assembly with ccdB | Cloning with negative selection | High-efficiency cloning without sequencing in Build phase | [46] |
| Selenzyme & RetroPath | Enzyme selection software | Computational enzyme selection in Design phase | [12] |
| PartsGenie & PlasmidGenie | DNA part design software | Automated biological part design in Design phase | [12] |
Traditional DBTL workflows in synthetic biology face significant bottlenecks at each stage of the cycle, including predictive modeling limitations in the Design phase, manual techniques in the Build phase, low-throughput screening in the Test phase, and knowledge extraction challenges in the Learn phase. These limitations collectively hinder the efficient engineering of biological systems for applications in therapeutics, sustainable chemicals, and biomaterials. However, emerging solutions centered on automation, artificial intelligence, and innovative molecular biology methods are actively addressing these constraints. The integration of these technologies enables more iterative and data-driven DBTL cycles, as demonstrated by case studies showing 500-fold improvements in product titers through automated, statistically-guided optimization [12]. As these solutions mature and become more accessible, they promise to transform synthetic biology from an empirical practice into a truly predictive engineering discipline, dramatically accelerating the development of biological solutions to global challenges.
In the contemporary landscape of biotechnology, biofoundries represent a transformative shift from artisanal research methods to industrialized, automated workflows. These facilities are integrated platforms that combine robotic automation, high-throughput measurement, and computational analytics to streamline and accelerate synthetic biology research and applications through the Design-Build-Test-Learn (DBTL) engineering cycle [49] [50] [51]. The core challenge biofoundries address is the inherent slowness, expense, and inconsistency of manual biological engineering, which traditionally limited the exploration of the vast biological design space [52]. By automating the highly repetitive but critical "Build" and "Test" stages of the DBTL cycle, biofoundries enable a massive increase in experimental throughput, allowing researchers to prototype and iterate biological systems with unprecedented speed and scale [49]. This capability is crucial for developing economically important bioengineered products and organisms, positioning biofoundries as foundational infrastructure for strengthening the global bioeconomy [49] [50]. This technical guide delves into the core architectures, methodologies, and operational frameworks that make these high-throughput facilities a reality.
The DBTL Cycle: The Conceptual Backbone of Biofoundries
The DBTL cycle is the core operational and conceptual model for all biofoundry activities, transforming biological engineering into a rigorous, iterative process [50] [53].
- Design: The cycle initiates with a software-driven design phase where researchers use computational tools to design new nucleic acid sequences, genetic circuits, or metabolic pathways. This stage leverages computer-aided design (CAD) software, retrobiosynthesis algorithms, and increasingly, artificial intelligence (AI) to predict optimal genetic designs for a desired function [50] [52].
- Build: This phase involves the physical construction of the biological components designed in the previous step. Automation is pivotal here, with liquid-handling robots executing DNA assembly, cloning, and transformation protocols in high-density microplates (96-, 384-, or 1536-well formats), drastically increasing the number of constructs that can be assembled in parallel [49] [51].
- Test: The constructed biological systems are then subjected to high-throughput functional characterization. Automated analytical equipment, such as plate readers, flow cytometers, and mass spectrometers, screens and characterizes the performance of the built constructs, generating large, quantitative datasets [49] [50].
- Learn: In the final phase, data from the Test phase is analyzed using bioinformatic tools and machine learning (ML) models. The goal is to extract meaningful insights, identify correlations between design and function, and inform the redesign for the next DBTL iteration, progressively optimizing the system toward the desired specification [50] [52].
The following diagram illustrates the continuous, iterative nature of this core engineering cycle.
To manage the complexity of diverse experiments and ensure interoperability, a standardized framework for describing biofoundry operations is essential. A recently proposed abstraction hierarchy organizes activities into four distinct levels, facilitating clear communication and modular design [8] [54].
- Level 0: Project: This is the highest level, representing the overarching goal of an external user, such as "develop a yeast strain to produce a novel biofuel" [8] [54].
- Level 1: Service/Capability: This level defines the specific, offerable functions the biofoundry provides to fulfill a project. Services can be tiered, from providing access to individual equipment to supporting a full DBTL cycle [8] [54].
- Level 2: Workflow: A service is broken down into sequential, modular workflows, each assigned to a single stage of the DBTL cycle (e.g., "DNA Assembly" for Build, "Microplate Assay" for Test). This modularity allows for the reconfiguration of standard workflows to create new services [8] [54].
- Level 3: Unit Operation: This is the most fundamental level, representing an individual task performed by a single piece of hardware or software. Examples include "Liquid Transfer" by a liquid handler or "Protein Structure Generation" by a specific software application [8] [54].
The relationship between these levels is visualized in the following hierarchy diagram.
Tiered Service Models
Biofoundries engage with their users through different service tiers, which define the scope of their involvement in the DBTL cycle, as summarized in the table below.
Table: Tiered Service Models in Biofoundries
| Tier | Description | Example |
|---|---|---|
| Tier 1 | Provides access to individual pieces of automated equipment. | Access to a liquid handling robot for user-led experiments [8]. |
| Tier 2 | A service focused on a single stage of the DBTL cycle. | Providing a protein sequence library designed by an AI tool like Protein MPNN [8]. |
| Tier 3 | A service combining two or more DBTL stages. | A common service involving the construction of a genetic library (Build) and its sequence verification (Test) [8]. |
| Tier 4 | A comprehensive service supporting the full DBTL cycle. | Applying the full DBTL cycle to engineer a microorganism for plastic degradation or to discover a new therapeutic enzyme [8] [50]. |
Automating the Build Phase: From Genetic Design to Physical Construct
The Build phase translates digital genetic designs into physical DNA constructs and introduces them into a host organism. Automation in this phase brings precision, reproducibility, and massive parallelism to molecular biology protocols.
Core Build Workflows and Unit Operations
A standard high-throughput Build workflow for microbial strain engineering might include DNA synthesis, construct assembly, transformation, and colony picking [49]. Each of these steps comprises multiple unit operations.
Table: Key Unit Operations in the Build Phase
| Workflow | Example Unit Operations (Hardware/Software) | Function |
|---|---|---|
| DNA Assembly | Liquid Handling (e.g., Opentrons), Thermocycling (e.g., PCR machine), DNA Design Software (e.g., j5) | Assembles smaller DNA fragments (e.g., oligomers or genetic parts) into larger functional constructs like plasmids [8] [50]. |
| Transformation | Liquid Handling, Electroporation, Heat Block Incubation | Introduces assembled DNA constructs into microbial host cells (e.g., E. coli or yeast) [49]. |
| Colony Picking | Robotic Colony Picker, Liquid Handling | Selects and transfers individual microbial colonies from an agar plate to a culture microplate for further growth and screening [49] [55]. |
Experimental Protocol: High-Throughput DNA Assembly using Golden Gate Assembly
The following is a generalized methodology for automated DNA assembly, a cornerstone of the Build phase.
- Design (Prerequisite): Using DNA assembly design software (e.g., j5), design the assembly strategy. The software defines the final sequence and output the instructions for a liquid-handling robot, specifying the volumes and locations of DNA parts and reagents in the labware [50].
- Reagent Dispensing (Unit Operation: Liquid Handling): A liquid-handling robot is programmed to transfer the required volumes of DNA fragments (e.g., promoters, coding sequences), Golden Gate enzyme mix (e.g., Type IIS restriction enzyme and ligase), and buffer from source tubes or plates into a destination microplate [8].
- Enzymatic Assembly (Unit Operation: Thermocycling): The destination microplate is transferred to a thermal cycler, which runs a programmed temperature cycle. A typical Golden Gate cycle involves periods of digestion (37°C) and ligation (16°C), repeated multiple times to efficiently assemble the DNA parts [8].
- Transformation (Unit Operation: Electroporation/Heat Shock): The assembly reaction is then introduced into competent host cells. This can be automated using integrated systems that perform electroporation or manage heat shock steps in a high-throughput format [49].
Automating the Test Phase: High-Throughput Functional Characterization
The Test phase is where the functionality of the built constructs is rigorously assessed. Automation enables the quantitative screening of thousands of variants in parallel, generating the high-quality data essential for the Learn phase.
Core Test Workflows and Unit Operations
Test phase workflows are designed to measure the performance of engineered biological systems against project-specific metrics, such as metabolite production, enzyme activity, or growth.
Table: Key Unit Operations in the Test Phase
| Workflow | Example Unit Operations (Hardware) | Function |
|---|---|---|
| Cell Culturing | Microbioreactor Fermentation (e.g., BioLector), Liquid Handling for media exchange | Grows engineered strains under controlled conditions in small volumes (e.g., in 96-well plates) to produce biomass and target molecules [49] [8]. |
| High-Throughput Screening | Microplate Reading (Absorbance, Fluorescence), Flow Cytometry | Measures optical density (growth), fluorescence (reporter gene expression), or other spectrophotometric properties in a high-density format [49] [50]. |
| Analytics & Omics | Liquid Handling for sample prep, connection to Mass Spectrometry, Next-Generation Sequencing | Prepares and analyzes samples to identify and quantify specific metabolites (metabolomics) or verify genetic sequences (genomics) [49] [51]. |
Experimental Protocol: Screening a Microbial Library for Chemical Production
A standard workflow for identifying high-producing strains from a library involves cultivation and automated assay.
- Inoculation and Cultivation (Unit Operation: Microbioreactor Fermentation): A liquid-handling robot is used to inoculate sterile growth medium in a 96-well microplate from the culture plate created in the Build phase. The plate is then placed in an automated micro-bioreactor system that provides controlled temperature, shaking, and gas exchange, while periodically measuring optical density [49].
- Sample Preparation (Unit Operation: Liquid Handling): After a defined growth period, the robot is used to transfer a small, precise aliquot of the culture from each well to a new assay plate. It may also add reagents to lyse cells or develop a colorimetric/fluorometric signal proportional to the target chemical's concentration.
- Assay and Detection (Unit Operation: Microplate Reading): The assay plate is transferred to a multi-mode microplate reader. The instrument measures the signal (e.g., absorbance, fluorescence) from each well, quantifying the production level of the target molecule in every variant [49].
- Data Export: The raw quantitative data from the plate reader is automatically formatted and exported to a central database for the subsequent Learn phase.
The Scientist's Toolkit: Essential Research Reagent Solutions
The high-throughput operation of a biofoundry relies on a standardized set of reagents and materials compatible with automated platforms.
Table: Essential Research Reagents and Materials for Biofoundries
| Item | Function in Automated Workflows |
|---|---|
| Enzymatic Assembly Mixes | Pre-mixed, standardized reagents (e.g., for Golden Gate or Gibson Assembly) ensure consistent, robust DNA construction when dispensed by robots [49]. |
| Lyophilized Reagents | Pre-dispensed, stable reagents in microplates simplify workflow setup and increase reliability by reducing liquid handling steps and variability [56]. |
| Synthetic DNA Oligomers & Parts | Defined, sequence-verified DNA fragments are the fundamental building blocks for automated construction of larger genetic designs [49] [52]. |
| High-Throughput Media Kits | Pre-formulated, soluble powders or liquid concentrates for rapid preparation of microbial growth media in multi-well plates [49]. |
| Cryogenic Storage Plates | Specially designed microplates for archiving thousands of engineered strains at -80°C, integral to library management and reproducibility [49]. |
Biofoundries have firmly established themselves as a cornerstone of modern synthetic biology by mastering the automation of the Build and Test stages. Through the implementation of robust DBTL cycles, standardized abstraction hierarchies, and interconnected automated hardware, they have transformed biological engineering from a craft into a quantitative, high-throughput discipline. The ongoing integration of artificial intelligence and machine learning is set to further revolutionize these facilities, creating "self-driving labs" that can autonomously propose and run experiments [50] [51]. While challenges in sustainability, standardization, and data management persist, the continued growth of collaborative networks like the Global Biofoundry Alliance (GBA) ensures that these powerful platforms will continue to accelerate innovation, paving the way for a more sustainable, bio-based economy [49] [50].
Integrating Machine Learning to Decipher Complex Biological Data and Predict Outcomes
The field of biological research is undergoing a profound transformation, driven by the integration of machine learning (ML) methodologies capable of decoding complex, high-dimensional datasets. Machine learning, a branch of artificial intelligence (AI), provides a robust framework for analyzing intricate biological questions by developing computational systems that learn directly from data, enhancing their performance without explicit programming [57]. This paradigm shift is particularly significant in the context of synthetic biology, where the traditional Design-Build-Test-Learn (DBTL) cycle has long served as the foundational engineering approach for developing biological systems. However, recent advances are reshaping this landscape, suggesting a new paradigm where "Learning" can strategically precede "Design" [4].
Machine learning addresses three fundamental challenges in computational biology: the scale problem of enormous biological datasets encompassing billions of genomic sequences and terabytes of multi-omics data; the complexity problem of biological systems exhibiting non-linear relationships and emergent behaviors; and the integration problem of harmonizing heterogeneous data types from genomics, transcriptomics, proteomics, metabolomics, and clinical records [58]. By tackling these challenges, ML enables researchers to move beyond traditional reductionist approaches and embrace the complexity of living systems through integrative, data-driven methodologies, accelerating discovery timelines that once spanned decades into processes measurable in months or weeks [58].
Key Machine Learning Algorithms and Their Biological Applications
Foundational Algorithms
Several machine learning algorithms have demonstrated particular utility in biological research due to their complementary strengths in handling different data types and biological questions. These algorithms form the foundation for more advanced techniques and are selected based on their widespread adoption, balance between predictive accuracy and interpretability, and scalability across diverse dataset sizes [57].
Figure 1: Machine Learning Algorithms in Biological Research: This diagram illustrates the categorization of key ML algorithms and their primary applications in biological research, showing how different learning paradigms support various analytical tasks.
Algorithm Comparison and Selection Guidelines
Table 1: Key Machine Learning Algorithms in Biological Research
| Algorithm | Core Functionality | Advantages | Common Biological Applications | Key Considerations |
|---|---|---|---|---|
| Ordinary Least Squares (OLS) Regression | Minimizes sum of squared residuals to estimate linear relationship parameters [57] | Computational efficiency, interpretability, well-understood theoretical foundation | Genomic prediction, metabolic flux analysis, gene expression modeling [57] | Sensitive to outliers, assumes linearity and independence of observations [57] |
| Random Forest | Ensemble method combining multiple decision trees via bagging [57] [59] | Handles high-dimensional data, robust to outliers, provides feature importance metrics [59] | Patient stratification, cell type classification, variant effect prediction [59] | Can be computationally intensive, less interpretable than single trees [57] |
| Gradient Boosting Machines | Ensemble method that iteratively builds decision trees to minimize errors from previous trees [57] [59] | High predictive accuracy, handles mixed data types, effective with complex interactions [59] | Disease outcome prediction, genetic risk assessment, protein function prediction [59] | Prone to overfitting without careful tuning, requires extensive parameter optimization [57] |
| Support Vector Machines (SVMs) | Finds optimal hyperplane to separate classes in high-dimensional space [57] [13] | Effective in high-dimensional spaces, memory efficient, versatile with kernel functions [57] | Disease classification from omics data, protein structure prediction, metabolic pathway analysis [13] | Performance depends on kernel selection, less effective with noisy data [57] |
| Neural Networks/Deep Learning | Multiple layered networks that learn hierarchical representations through nonlinear transformations [58] [59] | Captures complex nonlinear relationships, state-of-art for many pattern recognition tasks [58] | Protein structure prediction (AlphaFold), single-cell analysis, drug discovery [58] | "Black box" nature reduces interpretability, requires large datasets [60] |
The selection of appropriate ML algorithms depends on multiple factors including dataset size, dimensionality, required interpretability, and the specific biological question. For instance, while simple linear models offer transparency and computational efficiency for initial explorations, more complex ensemble methods and neural networks provide superior predictive power for modeling intricate biological interactions at the cost of interpretability and greater computational requirements [57] [59].
Integrating Machine Learning with Synthetic Biology DBTL Cycles
The Traditional DBTL Framework
The Design-Build-Test-Learn (DBTL) cycle represents a systematic, iterative framework commonly used in synthetic biology to engineer biological systems [1]. In this established paradigm, researchers first Design biological components based on objectives for desired function, then Build DNA constructs through synthesis and assembly into appropriate vectors or chassis. The Test phase experimentally measures the performance of these engineered constructs, and the Learn phase analyzes the resulting data to inform the next design iteration [4]. This cyclic process has become the cornerstone of biological engineering, enabling the development of organisms for specific functions such as producing biofuels, pharmaceuticals, or other valuable compounds [1].
The Machine Learning-Enhanced LDBT Paradigm
Recent advances in machine learning are fundamentally transforming this traditional workflow. The integration of ML capabilities has prompted a proposed paradigm shift from DBTL to "LDBT" (Learn-Design-Build-Test), where Learning precedes Design [4]. This reordering leverages the predictive power of pre-trained ML models that have learned from vast biological datasets, enabling more informed initial designs and potentially reducing the number of experimental iterations needed.
Figure 2: DBTL to LDBT Paradigm Shift: This diagram contrasts the traditional Design-Build-Test-Learn cycle with the emerging Learn-Design-Build-Test paradigm enhanced by machine learning and cell-free testing technologies.
Implementation with Cell-Free Systems and Automation
The integration of cell-free expression systems with machine learning further accelerates the Build and Test phases of the cycle [4]. These systems leverage protein biosynthesis machinery from cell lysates or purified components to activate in vitro transcription and translation, enabling rapid protein production without time-intensive cloning steps. When combined with liquid handling robots and microfluidics, cell-free platforms can screen hundreds of thousands of reactions, generating the massive datasets required for training effective ML models [4].
This synergistic combination of machine learning and rapid experimental prototyping is transforming synthetic biology from an empirical, iterative discipline toward a more predictive engineering science. As zero-shot prediction capabilities improve—where models can make accurate predictions without additional training—the field moves closer to a Design-Build-Work model similar to established engineering disciplines like civil engineering [4].
Technical Implementation: Architectures and Workflows
Data Integration Architectures
Modern computational biology faces the significant challenge of integrating diverse data types, from DNA sequences and protein structures to cellular images and clinical records. Machine learning frameworks address this through sophisticated architectural designs capable of processing and integrating multi-modal biological data.
Table 2: Essential Research Reagent Solutions for ML-Enhanced Biology
| Reagent/Technology | Function | Application in ML Workflows |
|---|---|---|
| Cell-Free Expression Systems | Protein biosynthesis machinery for in vitro transcription and translation [4] | Rapid testing of ML-designed protein variants without cloning; megascale data generation [4] |
| DNA Assembly Technologies | Modular construction of genetic circuits and pathways [1] | Building ML-designed genetic constructs for experimental validation [1] |
| Single-Cell RNA Sequencing | High-resolution profiling of gene expression at single-cell level [58] | Generating training data for cell type classification and developmental trajectory models [58] |
| Next-Generation Sequencing | High-throughput DNA and RNA sequencing [57] [1] | Validating synthetic constructs; generating genomic datasets for model training [57] |
| Mass Spectrometry | Proteomic and metabolomic profiling [13] | Quantitative protein and metabolite data for multi-omics integration [13] |
Figure 3: Multi-Omics Integration Architecture: This diagram illustrates a machine learning framework for integrating diverse biological data types through specialized encoders and cross-attention mechanisms.
Implementation Example: Multi-Omics Integration
A representative implementation for multi-omics integration employs modality-specific processing followed by cross-modal integration:
Code Example 1: Conceptual framework for multi-omics integration using specialized encoders and attention mechanisms [58].
This architecture enables several advanced capabilities: (1) Modality-specific processing where each biological data type receives specialized preprocessing through domain-appropriate neural architectures; (2) Cross-modal learning where attention mechanisms enable the model to learn relationships between different biological layers; and (3) Adaptive integration where sophisticated fusion networks weight different modalities based on their relevance and data quality [58].
Applications and Experimental Protocols
Protein Engineering and Design
Machine learning has revolutionized protein engineering through both sequence-based and structure-based approaches. Sequence-based protein language models—such as ESM and ProGen—are trained on evolutionary relationships between protein sequences and can perform zero-shot prediction of beneficial mutations and protein functions [4]. Structural models like ProteinMPNN take entire protein structures as input and predict new sequences that fold into specified backbones, achieving nearly 10-fold increases in design success rates when combined with structure assessment tools like AlphaFold [4].
Experimental Protocol: ML-Guided Protein Optimization
- Library Design: Use pre-trained protein language models (e.g., ESM, ProGen) to generate candidate sequences with desired properties [4].
- Construct Assembly: Employ cell-free DNA assembly methods for rapid construction of protein variant libraries [4].
- High-Throughput Screening: Express variants in cell-free systems coupled with robotic liquid handling and microfluidics to screen >100,000 reactions [4].
- Functional Assays: Implement colorimetric or fluorescent-based assays for high-throughput sequence-to-function mapping [4].
- Model Retraining: Incorporate experimental results to refine predictive models for subsequent design iterations [4].
Genomic Prediction and Disease Modeling
In genomics, machine learning approaches are advancing beyond traditional genome-wide association studies (GWAS) and polygenic risk scores (PRS) by capturing non-linear relationships and genetic interactions [59]. While PRS provides a single variable measuring genetic liability by aggregating genome-wide genotype data, ML methods can model complex interaction effects that contribute to disease risk [59]. For brain disorders specifically, ML has shown potential in identifying genetically homogenous subgroups and improving predictive accuracy beyond classical statistical methods [59].
Experimental Protocol: Disease Subtype Stratification
- Data Collection: Aggregate multi-omics data (genomic, transcriptomic, epigenomic) from patient cohorts and relevant preclinical models [61] [59].
- Feature Preprocessing: Conduct quality control, normalization, and batch effect correction across heterogeneous data sources [59].
- Dimensionality Reduction: Apply variational autoencoders or contrastive learning to compress high-dimensional data while preserving biological variance [61] [59].
- Stratification Modeling: Implement unsupervised or semi-supervised clustering algorithms (e.g., graph neural networks, multimodal transformers) to identify patient subtypes [61] [59].
- Biological Validation: Validate identified subtypes through differential expression analysis, pathway enrichment, and correlation with clinical outcomes [59].
Drug Discovery and Development
AI and ML are playing increasingly important roles throughout the drug development lifecycle, from target identification to clinical trial optimization [60] [62] [63]. The FDA has reported a significant increase in drug application submissions using AI components, with over 500 submissions incorporating AI across various development stages from 2016 to 2023 [62]. Digital twin technology—creating AI-driven models that predict individual patient disease progression—is being used to design clinical trials with fewer participants while maintaining statistical power [63].
Experimental Protocol: AI-Enhanced Clinical Trials
- Historical Data Aggregation: Compile longitudinal patient data from previous trials and real-world evidence [63].
- Digital Twin Development: Train generative models to simulate disease progression for individual patients based on their baseline characteristics [63].
- Trial Design Optimization: Use digital twins to create virtual control arms, reducing required trial participants [63].
- Prospective Validation: Conduct clinical trials comparing outcomes between treated patients and their digital twins [63].
- Regulatory Submission: Document model development, validation, and performance for regulatory review [60] [62].
Future Directions and Regulatory Considerations
Emerging Trends and Technologies
The field of machine learning in biology continues to evolve rapidly, with several emerging trends shaping its trajectory. Physics-informed machine learning represents a promising hybrid approach that incorporates known biological principles and physical laws into ML architectures, combining the flexibility of data-driven learning with the reliability of established biological knowledge [58]. Federated learning approaches enable model training across multiple institutions without sharing sensitive patient data, addressing privacy concerns while leveraging diverse datasets [61] [59]. As the capabilities of large language models expand, their application to biological sequences and structures is expected to drive further advances in zero-shot prediction and generative design [4] [63].
Regulatory Landscape and Validation Frameworks
The regulatory environment for AI/ML in drug development is evolving rapidly, with distinct approaches emerging across different jurisdictions. The U.S. Food and Drug Administration (FDA) has adopted a flexible, case-specific model, engaging with developers through its CDER AI Council and reviewing over 500 submissions incorporating AI components [62]. In contrast, the European Medicines Agency (EMA) has established a more structured, risk-tiered approach that explicitly addresses AI implementation across the entire drug development continuum [60]. Both agencies emphasize the importance of rigorous validation, documentation, and performance monitoring for AI systems used in regulatory decision-making [60] [62].
For researchers implementing ML in biological applications, key regulatory considerations include: (1) maintaining comprehensive documentation of data provenance and preprocessing steps; (2) implementing robust model validation using external datasets; (3) establishing protocols for ongoing performance monitoring and model maintenance; and (4) ensuring transparency and interpretability to the extent possible, particularly for "black box" models [60]. As regulatory frameworks continue to mature, early engagement with regulatory agencies through channels like the FDA's CDER AI Council or EMA's Innovation Task Force is recommended for high-impact applications [60] [62].
The integration of machine learning approaches to decipher complex biological data represents a paradigm shift in biological research and synthetic biology. By enhancing traditional DBTL cycles with predictive ML capabilities, researchers can accelerate the design of biological systems, from engineered proteins to optimized microbial cell factories. The synergistic combination of machine learning with high-throughput experimental technologies like cell-free systems and automated biofoundries is transforming biological engineering from an empirical, iterative process toward a more predictive discipline. As regulatory frameworks evolve to address the unique challenges of AI/ML in biological applications, and as computational methods continue to advance, the integration of machine learning promises to unlock new frontiers in understanding biological complexity and engineering biological systems for therapeutic and industrial applications.
Leveraging AI for Predictive Biodesign and Accelerated Learning
The integration of artificial intelligence (AI) into synthetic biology is transforming the traditional design-build-test-learn (DBTL) cycle from a sequential, time-consuming process into a rapid, iterative, and predictive framework. This paradigm shift is enabling researchers to move from manual trial-and-error approaches to algorithmic biodesign, dramatically accelerating the pace of biological innovation [64]. AI's capacity to process vast, multi-dimensional biological datasets and generate novel designs is compressing discovery timelines, reducing costs, and opening new frontiers in therapeutic development, sustainable chemistry, and materials science [65] [66].
The core of this transformation lies in the enhancement of each stage of the DBTL cycle. AI models, particularly machine learning (ML) and deep learning (DL), now assist in designing DNA sequences, predicting protein structures, optimizing metabolic pathways, and prioritizing the most promising constructs for experimental testing [65] [43]. This technical guide explores the specific methodologies and tools at the intersection of AI and biodesign, providing researchers and drug development professionals with a detailed roadmap for implementing these approaches to accelerate their own discovery pipelines.
AI-Enhanced Phases of the Biodesign Cycle
The Design Phase: In-Silico Prediction and Generation
In the design phase, AI shifts the paradigm from screening existing knowledge to generating novel, optimized biological parts and systems.
Protein Structure Prediction and Design: Tools like AlphaFold2 have demonstrated near-atomic accuracy in predicting protein structures from amino acid sequences, a breakthrough recognized by the 2024 Nobel Prize in Chemistry [64]. Subsequent models, such as RoseTTAFold and EvoDiff, have expanded this capability to the de novo design of novel proteins with desired functions [64]. These models use deep learning architectures trained on vast datasets of known protein sequences and structures to learn the fundamental principles linking sequence to structure and function.
Generative Biological Design: Generative AI algorithms can now propose novel DNA and protein sequences that meet specific functional criteria. For metabolic pathway optimization, this involves designing enzyme variants with improved catalytic activity or stability [65]. For instance, researchers used machine learning to predict specific mutations that led to the engineering of FAST-PETase, an enzyme that efficiently breaks down PET plastics at ambient temperatures [65].
Multi-Omics Integration: AI excels at integrating heterogeneous data types. Machine learning models can combine genomic, transcriptomic, and proteomic data to identify key regulatory nodes and promising targets for engineering [67]. Representation-learning techniques produce unified embeddings from these multi-omics inputs, enabling more comprehensive biomarker discovery and mechanistic inference [67].
The Build and Test Phases: Automation and High-Throughput Data Generation
The build and test phases are characterized by the rise of automation and sophisticated data collection, generating the high-quality datasets required to power AI models.
Automated Biofoundries: Automated "biofoundries" integrate robotic systems to execute the physical construction of genetic designs (e.g., via DNA synthesis and assembly) and testing (e.g., via culturing and assays) in a high-throughput manner [64]. This automation drastically shortens the cycle time and generates large, standardized datasets that are essential for training robust ML models [64].
In-Silico Simulation and Prototyping: AI enables extensive in-silico prototyping before any physical experiment. Mechanistic kinetic models, which use ordinary differential equations to describe metabolic networks, can simulate the behavior of thousands of strain designs [22]. This allows for the preliminary evaluation of designs, saving considerable resources. For example, simulations can model a batch bioreactor process to predict product titers based on perturbations to enzyme concentrations [22].
The Learn Phase: Machine Learning for Insight and Recommendation
The learn phase is where AI adds the most significant value, turning experimental data into actionable insights for the next cycle.
Machine Learning for Pathway Optimization: In combinatorial pathway optimization, the number of possible genetic designs often leads to a combinatorial explosion. ML models are used to learn from the data generated in the "test" phase and recommend the next set of promising strains to build [22]. As demonstrated in simulated DBTL cycles, algorithms like gradient boosting and random forest have proven particularly effective in the low-data regime typical of early-stage projects, showing robustness to training set biases and experimental noise [22].
The Automated Recommendation Algorithm: A key methodology is the implementation of automated recommendation tools. These systems use an ensemble of ML models to create a predictive distribution of strain performance. Based on this, they sample new designs for the next DBTL cycle, balancing the exploration of new regions of the design space with the exploitation of known high-performing areas [22]. The algorithm's performance can be optimized by tuning parameters based on the desired balance between risk and reward.
Table 1: Machine Learning Models and Their Applications in the DBTL Cycle
| ML Model | Primary Application in DBTL | Key Advantages | Example Use-Case |
|---|---|---|---|
| Random Forest / Gradient Boosting | Recommending strain designs in metabolic engineering [22] | Robust performance with small, noisy datasets; handles non-linear relationships [22] | Optimizing enzyme levels in a synthetic pathway to maximize product flux [22] |
| Convolutional Neural Networks (CNNs) | Screening compound libraries; predicting protein-ligand binding [65] [67] | Automates feature extraction from raw data (e.g., molecular structures) | Virtual screening of millions of compounds for drug discovery [65] [66] |
| Recurrent Neural Networks (RNNs) | Modeling biological sequences and time-series data [67] | Captures sequential dependencies and context | Predicting how genetic sequences evolve over time in continuous culture |
| Transformers/Large Language Models | Predicting biological structure and function from sequence [43] | Models long-range interactions in sequences; transfer learning from large corpora | Predicting regulatory elements or protein folding from DNA sequence [43] |
Experimental Protocols and Methodologies
Protocol: Simulated DBTL Cycles for Combinatorial Pathway Optimization
This protocol outlines a framework for using mechanistic kinetic models to simulate and optimize machine learning-guided DBTL cycles, as detailed in [22].
1. Define the Kinetic Model and Design Space:
- Model Construction: Develop or use a pre-existing mechanistic kinetic model of the host organism's core metabolism (e.g., an E. coli core kinetic model). Integrate the synthetic pathway of interest into this model [22].
- Define Library Components: Specify the genetic parts library (e.g., promoters, RBSs) for each gene in the pathway. Each part is associated with a specific effect on enzyme activity (Vmax) in the kinetic model [22].
- Map to Simulation: Each possible strain design is a unique combination of these library components, which is translated into a specific set of Vmax parameters for the kinetic model.
2. Execute the Initial DBTL Cycle:
- Design: Randomly select or use a design-of-experiments approach to choose an initial set of strain designs (e.g., 50-100 strains) from the full combinatorial space [22].
- Build & Test (In-Silico): Run simulations for each design. The model outputs the product flux (titer, yield, or rate) for each strain, simulating the experimental "test" phase [22].
- Learn: Train a machine learning model (e.g., Gradient Boosting Regressor) on the dataset of strain designs (input features) and their corresponding simulated product fluxes (target variable) [22].
3. Iterate with ML-Driven Recommendations:
- Recommend: Use the trained ML model to predict the performance of all untested strain designs in the combinatorial space. An algorithm (e.g., upper confidence bound sampling) selects the next set of strains to "build," balancing exploration of uncertain designs and exploitation of predicted high-performers [22].
- Repeat: Execute subsequent DBTL cycles by iterating through the Build-Test-Learn-Recommend steps. In each cycle, the ML model is re-trained on all data accumulated from previous cycles [22].
4. Validate and Analyze:
- Compare the performance of different ML recommendation algorithms and DBTL cycle strategies (e.g., large initial cycle vs. smaller, more frequent cycles) by tracking how quickly they converge to the optimal strain design as identified by the kinetic model [22].
Workflow Visualization: AI-Augmented DBTL Cycle
The following diagram illustrates the continuous, AI-enhanced DBTL cycle, showing the flow of information and the specific role of AI and automation at each stage.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Research Reagents and Platforms for AI-Driven Biodesign
| Item / Solution | Function in AI-Biodesign Workflow |
|---|---|
| Curated Multi-Omics Datasets | Provides the high-quality, annotated data required to train and validate machine learning models for predicting biological function [67]. |
| Mechanistic Kinetic Models (e.g., SKiMpy) | Serves as a simulated "ground truth" for benchmarking ML algorithms and optimizing DBTL cycle strategies before costly wet-lab experiments [22]. |
| Automated Biofoundry Infrastructure | Integrated robotics and software that automate the "Build" and "Test" phases, enabling high-throughput, reproducible data generation for learning [64]. |
| Cloud-Based AI/ML Platforms (e.g., TensorFlow, PyTorch) | Provides scalable computational frameworks and libraries for developing, training, and deploying custom machine learning models for biological data [67]. |
| Protein Structure Prediction Suites (e.g., AlphaFold, EvoDiff) | AI tools that accurately predict or generate protein 3D structures, bridging the sequence-structure-function gap in the design phase [64] [65]. |
| DNA Synthesis and Screening Services | Commercial services that provide physical DNA constructs from digital designs and offer functional screening, closing the loop between in-silico design and physical validation [65] [68]. |
Challenges, Risks, and Future Directions
Technical and Ethical Considerations
The adoption of AI in biodesign is not without significant challenges and risks that the scientific community must address.
Data Quality and Reproducibility: AI models are profoundly sensitive to the quality of their training data. Artifacts, biases, or simple errors in datasets can lead to inaccurate or non-reproducible models [69]. Ensuring data breadth, accuracy, and ethical sharing through open science practices is critical for generating reliable outcomes [69].
Dual-Use and Biosecurity Risks: Generative AI tools capable of designing proteins pose a potential biosecurity threat. AI can design novel harmful protein sequences with little homology to known pathogens, potentially evading current DNA-synthesis screening methods that rely on sequence similarity [64] [68]. There is an urgent need to develop function-based screening standards and update international biosecurity frameworks to address this gap [68].
Interpretability and Oversight: The "black box" nature of some complex AI models can make it difficult to understand the rationale behind their designs. This lack of interpretability challenges scientific validation and necessitates human oversight. A hybrid approach, combining AI with traditional mathematical modeling that incorporates known biological mechanisms, can provide greater specificity and transparency [69].
Visualization: The Biosecurity Screening Gap
The diagram below illustrates the emerging security challenge posed by AI-designed biological sequences and the proposed hybrid screening solution.
Future Outlook
The trajectory of AI in biodesign points toward increasingly autonomous systems. The development of "AI co-scientists"—multi-agent systems that can generate hypotheses, check them against existing knowledge, and propose experimental sequences—heralds a future of human-machine collaborative discovery at an unprecedented scale [65]. Furthermore, the market growth of AI in synthetic biology, projected to rise from USD 94.7 million in 2024 to USD 438.4 million by 2034, underscores the significant and sustained investment in this transformative convergence [64]. The focus for the research community will be to harness this power responsibly, developing the technical, ethical, and security frameworks that ensure these accelerated learning cycles lead to beneficial and safe outcomes for society.
The traditional path of drug development is notoriously protracted and costly, averaging 10-15 years and exceeding $2.5 billion from initial discovery to regulatory approval [70]. This timeline is primarily hampered by high failure rates at every stage, with approximately 90% of drug candidates failing during clinical development [70]. However, a transformative shift is underway. Artificial Intelligence (AI) is fundamentally restructuring this pipeline, offering a powerful strategy to compress development timelines and reduce attrition. This revolution is deeply intertwined with the engineering principles of synthetic biology, particularly the Design-Build-Test-Learn (DBTL) cycle, which provides a systematic, iterative framework for engineering biological systems [1]. AI is not merely accelerating this cycle; it is fundamentally reordering its components, enabling a leap from empirical iteration toward predictive, precision biological design [4] [25]. This whitepaper explores how AI-powered platforms, through specific case studies and new methodologies, are achieving unprecedented compression of development timelines, framed within the evolving context of the synthetic biology DBTL cycle.
Quantitative Impact: AI's Acceleration of Development Timelines
The integration of AI into biopharmaceutical research and development is generating substantial gains in speed and efficiency. The following table summarizes key quantitative evidence of this acceleration, drawn from recent industry analyses and scientific reports.
Table 1: Documented Impacts of AI on Drug Development Timelines and Efficiency
| Metric | Traditional Approach | AI-Accelerated Approach | Data Source & Context |
|---|---|---|---|
| Discovery to Clinical Trials | 5+ years | 12-30 months | Insilico Medicine (30 months); Exscientia (12 months for DSP-1181) [70] [71] |
| Clinical Trial Success Rate | ~40% (Phase I completion) | 80-90% (Phase I completion) | Analysis of 21 AI-developed drugs as of Dec 2023 [72] |
| Projected Annual Value for Pharma | N/A | $350 - $410 Billion | Projected annual value by 2025 from AI-driven innovations [71] |
| Reduction in Discovery Time/Cost | Baseline | Up to 40% time savings; 30% cost reduction | AI-enabled workflows for complex targets [71] |
| Candidate Drugs Entering Clinical Stages | Baseline | 3 (2016) → 67 (2023) | Exponential growth in AI-developed candidates [72] |
The Foundational Framework: The DBTL Cycle and Its AI-Driven Evolution
The Traditional DBTL Cycle in Synthetic Biology
Synthetic biology employs the Design-Build-Test-Learn (DBTL) cycle as its core engineering pipeline [1] [25]. This framework involves:
- Design: Researchers define objectives and use computational tools to design biological parts (e.g., DNA sequences, genetic circuits) [4].
- Build: The designed DNA constructs are synthesized and assembled into vectors, which are then introduced into a living chassis (like bacteria) or cell-free systems for characterization [1] [4].
- Test: The engineered constructs are experimentally measured for performance in functional assays, often using high-throughput automated screening [1] [25].
- Learn: Data from testing is analyzed to inform the next round of design, creating an iterative loop until the desired biological function is achieved [1].
Despite advancements in building and testing, the "Learn" phase has been a major bottleneck. The complexity of biological systems has made it difficult to extract definitive design principles from large datasets, often forcing researchers to rely on trial-and error [25].
The Paradigm Shift: From DBTL to LDBT
Recent advances in machine learning (ML) are prompting a radical rethinking of the DBTL cycle. With the advent of powerful models trained on vast biological datasets, the "Learning" phase can now precede the initial "Design". This is known as the "LDBT" paradigm—Learn-Design-Build-Test [4].
In LDBT, machine learning models that have been pre-trained on millions of protein sequences or structures are used to make zero-shot predictions for new designs with desired functions, without the need for multiple iterative cycles [4]. This approach leverages prior knowledge encoded in the models, potentially leading to functional solutions in a single pass and bringing synthetic biology closer to a "Design-Build-Work" model used in more mature engineering disciplines [4].
Diagram 1: The evolution from the traditional DBTL cycle to the AI-first LDBT paradigm.
Case Studies in Timeline Compression
Case Study 1: AI-Driven De Novo Drug Design for Idiopathic Pulmonary Fibrosis
Company: Insilico Medicine Achievement: Reduced the drug discovery and preclinical timeline from the industry standard of 5-6 years to just under 30 months [70].
Experimental Protocol & Workflow:
This case exemplifies the LDBT paradigm, leveraging a pre-trained AI platform for initial design.
- Learn (AI Foundation): The process began with AI models pre-trained on massive datasets of molecular structures, biological targets, and associated clinical data. This foundational "Learning" phase provided the knowledge base for all subsequent design.
- Design (Target Identification & Molecule Generation):
- Target Identification: Insilico's AI platform (PandaOmics) analyzed vast volumes of multi-omics data and scientific literature to identify a novel therapeutic target for idiopathic pulmonary fibrosis (IPF).
- Molecule Generation: Another AI system (Chemistry42) used generative adversarial networks (GANs) to design and optimize novel small molecule inhibitors specific to the identified target. The platform generated thousands of virtual candidate molecules, predicting their properties, efficacy, and synthesizability in silico.
- Build (Synthesis): The top-ranked AI-designed molecules were synthesized for testing. The AI's accurate prediction of synthesizable structures reduced the need for multiple, time-consuming chemical re-design rounds.
- Test (Preclinical Validation): The synthesized compounds underwent in vitro and in vivo testing. The lead candidate showed promising results in animal models of IPF, successfully progressing into human clinical trials.
Key AI Technologies: Generative Adversarial Networks (GANs) for molecular design; Deep Learning for target identification and validation.
Case Study 2: AI-Designed Enzyme for Plastic Depolymerization
Research Context: Engineering a hydrolase for improved depolymerization of polyethylene terephthalate (PET) plastic [4].
Experimental Protocol & Workflow:
This study showcases a hybrid DBTL cycle, where a structure-based machine learning model directly informed the design phase.
- Learn: A deep neural network model named MutCompute was trained on a large database of protein structures. The model learned to associate amino acids with their local chemical environments, allowing it to predict mutations that would improve protein stability and function [4].
- Design: Researchers used MutCompute to analyze the structure of a wild-type PET hydrolase. The model identified specific residue-level mutations predicted to stabilize the enzyme and enhance its activity.
- Build: DNA sequences encoding the AI-predicted enzyme variants were synthesized and assembled. The proteins were then expressed, typically in bacterial systems like E. coli, and purified for testing.
- Test: The engineered enzyme variants were experimentally characterized for their thermal stability and catalytic activity in depolymerizing PET. The AI-designed variants demonstrated increased stability and activity compared to the wild-type enzyme [4].
Key AI Technologies: MutCompute (structure-based deep neural network); ProteinMPNN for sequence design; AlphaFold and RoseTTAFold for structure prediction and assessment [4].
Diagram 2: Integrated AI-Cell-Free workflow for ultra-high-throughput protein engineering.
The Scientist's Toolkit: Essential Research Reagent Solutions
The acceleration of development timelines relies on a suite of enabling technologies and reagents that facilitate the high-throughput Build and Test phases of the (L)DBT cycle. The following table details key solutions used in the featured experiments and the broader field.
Table 2: Key Research Reagent Solutions for AI-Driven Biological Design
| Research Solution | Function in Workflow | Application in Case Studies |
|---|---|---|
| Cell-Free Gene Expression Systems | Crude lysates or purified cellular machinery that enable rapid in vitro transcription and translation of synthesized DNA templates without cloning. | Enables ultra-high-throughput testing of AI-designed protein variants (e.g., 100,000+ reactions), rapid pathway prototyping (iPROBE), and expression of potentially toxic proteins [4]. |
| DNA Synthesis & Assembly Kits | Commercial kits for the de novo chemical synthesis of DNA fragments (oligos) and their subsequent assembly into larger constructs (e.g., genes, pathways). | Essential for the "Build" phase, turning AI-designed digital sequences into physical DNA for testing in cell-free systems or living chassis [1] [4]. |
| Automated Biofoundries | Integrated facilities featuring robotic liquid handlers, automated incubators, and high-throughput analyzers that miniaturize and parallelize biological experiments. | Used for high-throughput molecular cloning, screening of large strain libraries, and generating the massive, standardized datasets required to train effective AI/ML models [13] [25]. |
| Protein Language Models (e.g., ESM, ProGen) | AI models trained on millions of natural protein sequences to learn evolutionary constraints and patterns, enabling zero-shot prediction of function and stability. | Used to design libraries of functional proteins, such as antimicrobial peptides (AMPs) and enzymes, directly from sequence data [4]. |
| Structure Prediction & Design Tools (e.g., AlphaFold, ProteinMPNN) | Deep learning systems that predict 3D protein structures from amino acid sequences (AlphaFold) or design sequences that fold into a specific structure (ProteinMPNN). | Critical for the de novo design of stable and active enzymes, as demonstrated in the engineering of PET hydrolases and TEV protease variants [72] [4]. |
The integration of AI-powered platforms into biopharmaceutical development is delivering on the promise of dramatically compressed timelines. The case studies of Insilico Medicine and AI-designed enzymes provide tangible evidence that AI can reduce years from the discovery process. This acceleration is not merely a matter of faster computing; it stems from a fundamental enhancement and reordering of the synthetic biology DBTL cycle into an LDBT paradigm. By placing Learning first through pre-trained models, AI enables more intelligent Design. When this is coupled with high-throughput Build and Test methodologies like cell-free systems and biofoundries, the entire path from concept to candidate becomes shorter, cheaper, and more predictable. As these technologies mature and regulatory frameworks adapt, the AI-driven compression of development timelines is poised to become the new standard, ushering in an era of more efficient and effective therapeutic and biomanufacturing innovation.
Validating the DBTL Impact: Comparative Analysis and Future Paradigms
In the fields of synthetic biology and drug development, the pursuit of efficiency has catalyzed a significant methodological evolution. The traditional trial-and-error approach, characterized by sequential, often intuitive experimentation, is increasingly being supplanted by the systematic, iterative framework of the Design-Build-Test-Learn (DBTL) cycle. The fundamental distinction between these paradigms lies in their core philosophy: trial-and-error operates as a linear, hypothesis-testing process, while DBTL embodies an integrated, data-driven engineering cycle where each phase systematically informs the next. This shift is particularly critical given the persistent challenges in biomedical research, where approximately 90% of clinical drug development fails despite extensive preclinical optimization efforts [73]. This analysis examines the comparative efficiency of these two approaches, quantifying their performance through empirical data, experimental protocols, and visual workflows to illustrate how DBTL principles are transforming biological engineering.
Core Conceptual Frameworks and Their Evolution
The Traditional Trial-and-Error Paradigm
The traditional trial-and-error approach has long been the default methodology in biological research and early-stage drug development. This paradigm typically follows a linear, sequential path where individual experiments are designed based on prior knowledge, executed, and then interpreted in isolation. The process lacks formalized feedback mechanisms to systematically inform subsequent design iterations, leading to extended development timelines and high failure rates. In clinical contexts, this manifests as high attrition rates where 40-50% of failures are attributed to lack of clinical efficacy and 30% to unmanageable toxicity, despite implementation of many successful strategies in preclinical development [73]. The approach is inherently reactive rather than proactive, with optimization occurring through discrete, often disconnected experiments rather than continuous, data-informed learning.
The Design-Build-Test-Learn (DBTL) Cycle
The DBTL framework represents a fundamental shift toward systematic biological engineering modeled after classical engineering disciplines. This iterative, closed-loop system comprises four integrated phases:
- Design: Researchers define objectives and create biological designs using computational tools, domain knowledge, and prior experimental data.
- Build: Genetic constructs are assembled and introduced into host systems using standardized biological parts and automated workflows.
- Test: Engineered systems are rigorously characterized through high-throughput analytical methods to quantify performance metrics.
- Learn: Data are analyzed using statistical methods and machine learning to extract insights that inform the next design iteration [12].
This framework creates a continuous improvement cycle where knowledge accumulates systematically with each iteration, enabling progressively refined designs and accelerated optimization.
The Emerging LDBT Paradigm
Recent advances in machine learning are catalyzing a further evolution toward Learning-Design-Build-Test (LDBT) frameworks. In this model, the "Learn" phase precedes "Design" through zero-shot predictions from pre-trained AI models on large biological datasets. This approach leverages protein language models (e.g., ESM, ProGen) and structural models (e.g., MutCompute, ProteinMPNN) that can directly generate functional biological designs without requiring multiple build-test iterations [4]. The paradigm shift enables researchers to begin with knowledge-rich computational predictions, effectively moving synthetic biology closer to a "Design-Build-Work" model akin to established engineering disciplines where first principles reliably guide development.
Quantitative Efficiency Comparison
The efficiency differential between DBTL and traditional approaches can be quantified across multiple performance dimensions, from development timelines to success rates and resource utilization.
Table 1: Efficiency Metrics Comparison Between Traditional and DBTL Approaches
| Performance Metric | Traditional Trial-and-Error | DBTL Approach | Efficiency Improvement |
|---|---|---|---|
| Pathway Optimization Time | Months to years for iterative testing | Weeks to months with automated cycling | 3-5x acceleration [12] |
| Experimental Throughput | Dozens of constructs manually | Thousands via automated workflows | >100x increase [4] |
| Success Rate | ~10% for clinical development | Competitive titers achieved in 2 cycles | 500-fold improvement demonstrated [12] |
| Data Generation Scale | Limited by manual processes | Megascale datasets via automation | >100,000 variants screenable [4] |
| Resource Efficiency | High reagent waste, personnel time | Optimized via statistical design | DoE achieves 162:1 compression [12] |
Table 2: Application-Specific Performance Gains with DBTL
| Application Domain | Traditional Results | DBTL Implementation | Documented Outcome |
|---|---|---|---|
| Flavonoid Production | Low or undetectable titers | Automated enzyme selection & expression tuning | 500-fold increase to 88 mg/L [12] |
| Protein Engineering | Multiple rounds of mutagenesis | Cell-free testing with ML guidance | Zero-shot prediction of functional variants [4] |
| Clinical Trial Success | 90% failure rate [73] | N/A (different application scope) | Limited direct impact on clinical failure causes |
| Biosensor Refactoring | Extensive manual optimization | Automated DBTL with modeling | Enhanced performance & circuit compatibility [74] |
Experimental Protocols and Implementation
Standardized DBTL Protocol for Metabolic Engineering
The implementation of DBTL cycles follows structured experimental protocols that enable reproducibility and scaling:
Design Phase Protocol:
- Pathway Identification: Use computational tools like RetroPath to identify potential biosynthetic routes from substrate to target compound [12].
- Enzyme Selection: Employ enzyme screening platforms like Selenzyme to select optimal enzyme candidates based on catalytic properties and host compatibility.
- Combinatorial Library Design: Utilize software such as PartsGenie to design regulatory elements and optimize codon usage.
- Library Compression: Apply Design of Experiments (DoE) methodologies with orthogonal arrays and Latin square designs to reduce combinatorial libraries to tractable sizes (e.g., 2592 to 16 constructs) [12].
Build Phase Protocol:
- Automated DNA Assembly: Implement ligase cycling reaction (LCR) or Golden Gate assembly on liquid handling robotics.
- Commercial Gene Synthesis: Outsource codon-optimized gene fragments for complex pathways.
- Quality Control: Perform high-throughput plasmid purification, restriction digest, and capillary electrophoresis for verification.
- Host Transformation: Introduce constructs into production chassis (e.g., E. coli, yeast) with standardized transformation protocols.
Test Phase Protocol:
- Cultivation: Execute automated 96-deepwell plate growth protocols with standardized media and induction conditions.
- Metabolite Extraction: Implement automated sample preparation using solid-phase extraction or liquid-liquid extraction.
- Analytical Screening: Employ UPLC-MS/MS with high mass resolution for quantitative analysis of target compounds and intermediates.
- Data Processing: Utilize custom R scripts for automated data extraction and peak integration.
Learn Phase Protocol:
- Statistical Analysis: Apply multivariate analysis (ANOVA) to identify significant factors influencing production titers.
- Machine Learning: Train predictive models on experimental datasets to identify non-linear relationships and interaction effects.
- Knowledge Integration: Feed insights back into the design phase for subsequent iterations with refined constraints [12].
Traditional Trial-and-Error Protocol
For comparative context, traditional approaches typically follow less standardized protocols:
- Hypothesis Generation: Design single constructs based on literature review and intuition.
- Manual Cloning: Use restriction enzyme-based cloning or PCR for construct assembly.
- Sequential Testing: Test constructs individually with limited replication.
- Qualitative Analysis: Interpret results through observation rather than statistical analysis.
- Intuitive Redesign: Modify subsequent designs based on researcher experience rather than systematic data modeling.
Workflow Visualization
The fundamental differences between these approaches become visually apparent when comparing their operational structures.
DBTL Cycle Workflow
Traditional Trial-and-Error Workflow
Enabling Technologies and Research Reagent Solutions
The implementation of efficient DBTL cycles relies on specialized technologies and reagents that enable automation, high-throughput processing, and data-driven analysis.
Table 3: Essential Research Reagent Solutions for DBTL Implementation
| Technology Category | Specific Tools/Reagents | Function in Workflow | Performance Benefit |
|---|---|---|---|
| DNA Assembly | Ligase Cycling Reaction (LCR) reagents | Modular pathway construction | Error-free assembly of multiple parts [12] |
| Cell-Free Systems | PURExpress, PANOx-SP | Rapid protein expression | >1 g/L protein in <4 hours [4] |
| Automation Platforms | Liquid handling robots, microfluidics | High-throughput screening | 100,000+ reactions screenable [4] |
| Analytical Instruments | UPLC-MS/MS systems | Metabolite quantification | High-resolution, quantitative data [12] |
| Machine Learning Tools | ESM, ProGen, ProteinMPNN | Zero-shot protein design | Reduced experimental cycles [4] |
| Design Software | RetroPath, Selenzyme, PartsGenie | Computational design | Automated part selection & optimization [12] |
Case Study: Flavonoid Production Optimization
A direct application comparing both approaches demonstrates the efficiency differential. In a project to optimize (2S)-pinocembrin production in E. coli:
DBTL Implementation:
- Round 1: A designed library of 16 representative constructs (from 2592 possible combinations) identified vector copy number and CHI promoter strength as most significant factors affecting production (P values = 2.00×10⁻⁸ and 1.07×10⁻⁷ respectively) [12].
- Round 2: Focused designs incorporating these insights achieved a 500-fold improvement in titers, reaching 88 mg/L competitive production levels.
- Total Timeline: Approximately 8 weeks for two complete DBTL cycles.
Projected Traditional Approach:
- Manual testing of limited construct combinations based on researcher intuition.
- Sequential optimization of individual factors rather than understanding interactions.
- Estimated timeline: 6-12 months to achieve similar optimization level.
- Lower final titers likely due to inability to detect complex factor interactions.
This case study exemplifies how the DBTL framework's systematic approach and statistical guidance dramatically accelerate the optimization process while generating fundamental insights into pathway rate-limiting steps.
The comparative analysis unequivocally demonstrates the superior efficiency of the DBTL framework over traditional trial-and-error approaches across multiple metrics. The structured iteration, statistical guidance, and integration of automation enable dramatic accelerations in development timelines, substantial improvements in success rates, and more efficient resource utilization. The emergence of LDBT paradigms with machine learning-forward approaches promises further efficiency gains through reduced experimental cycling.
However, successful implementation requires significant infrastructure investment in automation platforms, computational resources, and specialized expertise. The integration of cell-free systems with machine learning represents a particularly promising direction, enabling megascale data generation for model training while circumventing cellular complexity. As these technologies mature and become more accessible, the DBTL framework is positioned to fundamentally transform biological engineering from an empirical art to a predictive science, ultimately addressing the persistent efficiency challenges that have long constrained synthetic biology and drug development.
The Design-Build-Test-Learn (DBTL) cycle represents a foundational framework in synthetic biology, engineered to bring rigorous, iterative engineering principles to biological innovation. Within the biopharmaceutical industry, which currently has over 23,000 drug candidates in development, the traditional, linear research and development (R&D) model is proving increasingly challenging [75]. Declining R&D productivity, with phase 1 success rates falling to just 6.7%, coupled with an impending $350 billion patent cliff, has created an urgent need for more efficient R&D methodologies [75] [76]. The DBTL cycle, particularly when implemented within highly automated biofoundries, is emerging as a critical strategy to accelerate the development of new therapies, from gene editing and cell therapies to oligonucleotide-based drugs, by systematically reducing the time and cost associated with each experimental iteration [77] [55].
This whitepaper provides a quantitative analysis of the DBTL cycle's impact on biopharmaceutical innovation. It details the core components of the DBTL framework, presents measurable outcomes from industry implementations, and offers detailed experimental protocols that researchers can adapt to enhance their own drug development pipelines. The data and case studies presented herein demonstrate that DBTL is not merely a theoretical concept but a practical, powerful tool for addressing the sector's most pressing productivity challenges.
The Core DBTL Cycle: A Detailed Technical Breakdown
The DBTL cycle is a closed-loop system that transforms biological design from an art into a predictable engineering discipline. Its power lies in the rapid iteration of its four phases, with each cycle generating data that informs and improves the next.
Design
The Design phase involves the computational creation of genetic blueprints. This stage leverages generative AI and large language models (LLMs) to model biological systems and propose new DNA constructs, such as synthetic genes or plasmid vectors, intended to achieve a specific therapeutic function [78]. The shift towards more complex therapeutic modalities, including cell and gene therapies and oligonucleotide-based drugs, makes this computational design phase crucial for managing complexity and predicting biological behavior before moving to the lab [77] [76].
Build
In the Build phase, the digital designs are physically constructed into biological entities. This involves the synthesis of DNA/RNA and the creation of engineered cells or organisms. The market for this manufacturing is substantial and growing; the DNA manufacturing market, valued at USD 5.18 billion in 2024, is projected to reach USD 20.28 billion by 2032, driven by demand for plasmid and synthetic DNA used in these therapies [77]. This phase has evolved from manual, low-throughput cloning to automated, high-throughput DNA synthesis and assembly.
Test
The Test phase is where the constructed biological systems are rigorously evaluated against predefined performance criteria. This involves high-throughput analytical techniques such as next-generation sequencing (NGS), flow cytometry, and mass spectrometry to generate quantitative data on the design's performance [55]. The objective is to produce a high-fidelity dataset that captures the causal relationship between the genetic design (from Build) and the resulting phenotypic output.
Learn
The Learn phase is the engine of iterative improvement. Here, the data generated from the Test phase is analyzed, often using machine learning models, to extract insights into the underlying biological rules. For example, AI applications can be employed to "decipher the relationship between structure and function in enzyme production" [55]. These learned insights are then directly fed back into the next Design phase, creating a virtuous cycle where each iteration is smarter than the last, progressively optimizing the biological system toward the desired therapeutic outcome.
Table 1: Core Components of the DBTL Cycle in Biopharmaceutical R&D
| DBTL Phase | Key Activities | Enabling Technologies | Primary Output |
|---|---|---|---|
| Design | Target Identification, DNA Construct Design, In-silico Modeling | Generative AI, LLMs, CAD for Biology [78] | Digital Genetic Blueprint |
| Build | DNA/Gene Synthesis, Genome Editing, Plasmid Manufacture | DNA Synthesizers, CRISPR-Cas9, Automated Clone Picking [55] | Physical DNA/RNA/Engineered Cell |
| Test | High-Throughput Screening, Functional Assays, Omics Analysis | NGS, Flow Cytometry, HPLC, Automated Assays [55] | Quantitative Performance Dataset |
| Learn | Data Integration, Pattern Recognition, Model Refinement | Machine Learning, AI, Statistical Analysis [55] | Predictive Insights & New Hypotheses |
Diagram 1: The DBTL Cycle in Biopharma
Quantifying the Impact: Data and Case Studies
The implementation of DBTL cycles, particularly within automated biofoundries, is yielding measurable and dramatic improvements in R&D efficiency. The following data and case studies provide concrete evidence of its impact.
Market Growth and Efficiency Gains
The expansion of the DNA manufacturing market, a key enabler of the "Build" phase, is a direct indicator of DBTL's growing influence. Synthetic DNA alone dominated the market in 2024 with a 71.25% share, underscoring its utility in genetic engineering and pharmaceutical research [77]. The entire market is projected to grow at a CAGR of 18.65% from 2025 to 2032, far outpacing many traditional sectors, which reflects heavy investment in the infrastructure that supports iterative biological design [77].
Table 2: Quantitative Impact of DBTL and Biofoundry Implementation
| Metric | Traditional Workflow | DBTL/Biofoundry Workflow | Improvement | Source/Context |
|---|---|---|---|---|
| Strain Screening Capacity | 10,000 strains/year | 20,000 strains/day | ~500x increase | Lesaffre Biofoundry [55] |
| Project Timeline (Genetic Improvement) | 5 - 10 years | 6 - 12 months | ~90% reduction | Lesaffre Biofoundry [55] |
| DNA Manufacturing Market (2024) | USD 5.18 Billion | SNS Insider [77] | ||
| DNA Manufacturing Market (2032 Proj.) | USD 20.28 Billion | 18.65% CAGR | SNS Insider [77] | |
| Phase 1 Success Rate (Historical) | ~10% (10 years ago) | 6.7% (2024) | Industry Challenge | Evaluate [75] |
Case Study: The Lesaffre Biofoundry
A prominent example of DBTL acceleration comes from Lesaffre, a global provider of yeast and yeast-based products. The company invested in a private biofoundry consisting of over 100 interconnected programmable instruments [55]. This facility can perform 20,000 growth-based assays per day with automatic monitoring. The result was a staggering increase in screening capacity, from 10,000 strains per year to 20,000 per day [55]. This high-throughput "Test" capability directly compressed project timelines for genetic improvement from 5-10 years down to just 6-12 months [55]. This case demonstrates that DBTL is not confined to human therapeutics but is a versatile framework that accelerates biological engineering across multiple industries.
Application in Therapeutic Areas
The DBTL cycle is particularly critical for advanced therapies. The cell and gene therapy segment dominated the DNA manufacturing market's application share in 2024 at 46.20%, as the clinical and commercial demand for DNA constructs has "exponentially increased" [77]. Similarly, the oligonucleotide-based drugs segment is expected to be the fastest-growing, driven by an increasing focus on precision medicine and RNA-targeted drugs like those developed by Wave Life Sciences [77] [79]. These complex modalities require the precise, iterative optimization that the DBTL cycle provides.
Implementing DBTL: Experimental Protocols and Workflows
For research teams aiming to adopt this framework, below is a detailed protocol for a representative DBTL cycle aimed at optimizing a yeast strain for the production of a therapeutic protein.
Protocol: DBTL Cycle for Therapeutic Protein Strain Engineering
Aim: To increase the yield of a recombinant therapeutic protein in S. cerevisiae through two iterative DBTL cycles.
1. Design Phase (Computational Library Design)
- Step 1.1: Identify genetic targets (e.g., promoters, secretion signals, chaperones) known to influence protein expression and secretion. Use AI-based tools, like those from the Oxford Generative Biology Lab, to predict enzyme behavior and metabolic bottlenecks [80].
- Step 1.2: Design an oligonucleotide pool encoding a combinatorial library of these genetic variants. This can be sourced from synthetic DNA providers like Twist Bioscience, which manufactures DNA libraries for this purpose [79].
- Step 1.3: Use cloning design software to plan the assembly of these oligo pools into a yeast expression vector.
2. Build Phase (High-Throughput Library Construction)
- Step 2.1: Utilize an automated DNA assembly workstation (e.g., from Codex DNA or Opentrons) to perform Golden Gate assembly of the oligo pool into the linearized vector [77].
- Step 2.2: Transform the assembled library into competent S. cerevisiae cells using a high-throughput electroporator.
- Step 2.3: Plate transformed cells on selective agar in automated arraying systems. Use a high-throughput colony picker to inoculate thousands of individual clones into 96- or 384-well culture plates [55].
3. Test Phase (Automated Screening and Analytics)
- Step 3.1: Grow cultures in a controlled, automated incubator-shaker. The CULTIVATOR, a self-driving growth unit, is an example of technology that can monitor, harvest, and optimize this biomass [80].
- Step 3.2: After a set fermentation time, use liquid handling robots to separate cells from supernatant via centrifugation.
- Step 3.3: Quantify therapeutic protein titer in the supernatant using a high-throughput method like fluorescence-linked immunosorbent assay (FLISA) or HPLC. Integrate these instruments with a Laboratory Information Management System (LIMS) for automatic data capture [55].
4. Learn Phase (Data Analysis and Model Generation)
- Step 4.1: Correlate protein titer data (from Test) with the genetic sequence of each variant (from Build).
- Step 4.2: Employ machine learning (e.g., a random forest or neural network model) to identify which genetic combinations are most predictive of high yield.
- Step 4.3: Based on these insights, design a second, refined library that focuses on the most promising genetic space and incorporates new variations suggested by the model. This initiates the next DBTL cycle.
Diagram 2: Biofoundry Workflow Integration
The Scientist's Toolkit: Essential Research Reagents & Solutions
Table 3: Key Research Reagent Solutions for DBTL Workflows
| Reagent/Material | Function in DBTL Workflow | Example Use-Case |
|---|---|---|
| Synthetic Gene Fragments | "Build" phase; template for genetic constructs. | Assembling a library of promoter-gene fusions for expression testing. |
| Oligo Pools | "Build" phase; source of genetic diversity for libraries. | Creating a vast combinatorial library of protein variants. |
| GMP-Grade Plasmid DNA | "Build" phase; final vector for therapeutic development. | Manufacturing clinical-grade DNA for cell and gene therapies [77]. |
| Cloning & Assembly Kits | "Build" phase; enzymatic assembly of DNA parts. | High-throughput Golden Gate assembly of a yeast expression library. |
| Cell Culture Media | "Test" phase; supports growth of engineered organisms. | High-throughput screening of yeast clones in 384-well plates. |
| NGS Library Prep Kits | "Test" and "Learn" phases; enables genotyping of variants. | Sequencing the genomes of top-performing clones to identify causal mutations. |
| Antibodies & Detection Assays | "Test" phase; quantifies protein expression and function. | FLISA for measuring therapeutic protein titer in culture supernatant. |
The Future of DBTL: AI, Automation, and Advanced Analytics
The next stage of DBTL evolution is the deeper integration of physical and generative artificial intelligence, which promises to further compress cycle times and enhance predictive power.
- AI-Driven Design: AI is moving beyond analysis to become a generative partner. At Lesaffre, AI applications are being used to "improve high-throughput screening, troubleshoot robot performance, [and] decipher the relationship between structure and function" [55]. Furthermore, initiatives like the Global Biofoundry Alliance, a consortium of over 25 organizations, aim to create open-source models and share capabilities, broadening access to these powerful tools [55].
- The Rise of the Biofoundry: The highly automated biofoundry model is becoming the physical embodiment of the DBLC cycle. These facilities integrate advanced instrumentation for DNA sequencing, flow cytometry, and cell culturing to create a seamless, automated pipeline [55]. This integration is critical for achieving the 500-fold increases in screening throughput that define the new pace of biopharmaceutical R&D [55].
- Addressing Computational Demand: This AI-driven approach requires immense computational resources. The biotech industry's compute demand is rapidly outpacing supply, with forecasts projecting $2.8 trillion in AI-related infrastructure spending by 2029 across the tech sector [81]. Biopharma's share of this compute load is growing as it becomes one of AI's largest consumers, a trend exemplified by projects like DeepMind's AlphaFold and numerous AI-biotech partnerships [81].
The quantitative evidence is clear: the Design-Build-Test-Learn cycle is a transformative force in biopharmaceutical R&D. By applying an iterative, data-driven, and automated engineering framework to biology, the DBTL paradigm directly addresses the industry's core challenges of soaring costs, protracted timelines, and high attrition rates. The case of Lesaffre's biofoundry, which reduced decade-long projects to a matter of months, provides a powerful template for the entire sector [55]. As the industry navigates a significant patent cliff and increasing pipeline complexity, the widespread adoption and continuous refinement of the DBTL cycle will be a key determinant of success, enabling the efficient and accelerated delivery of the next generation of life-saving therapies.
The convergence of artificial intelligence (AI) and synthetic biology is fundamentally reshaping biological discovery and engineering. This fusion is revolutionizing the core synthetic biology pipeline—the Design-Build-Test-Learn (DBTL) cycle—by introducing unprecedented levels of speed, prediction accuracy, and automation [43]. AI-driven tools, particularly machine learning (ML) and large language models (LLMs), are accelerating bioengineering workflows, unlocking innovations in medicine, agriculture, and sustainability [43]. The integration of AI is so transformative that it is prompting a radical rethinking of the traditional DBTL sequence, potentially shifting towards a "Learning-Design-Build-Test" (LDBT) model where machine learning precedes and informs the initial design phase [4]. This technical guide examines the mechanisms of this convergence, detailing how AI optimizes each stage of the synthetic biology cycle, presents structured experimental data and protocols, and explores the emerging tools and computational frameworks that constitute the modern scientist's toolkit for next-generation biological design.
The AI-Enhanced Design-Build-Test-Learn (DBTL) Cycle
The DBTL cycle is a systematic, iterative framework used in synthetic biology to develop and optimize biological systems [1]. Even with rational design, the impact of introducing foreign DNA into a cell can be difficult to predict, creating the need to test multiple permutations to obtain a desired outcome [1]. AI and ML are now revolutionizing this cycle, enhancing efficiency and predictive power at every stage.
The "Design" Phase: From Manual Curation to AI-Driven Prediction
The Design phase involves defining objectives for a desired biological function and designing the biological parts or system to achieve it [4]. AI has dramatically expanded the capabilities of this stage.
- Protein Language Models: Tools like ESM (Evolutionary Scale Modeling) and ProGen are trained on evolutionary relationships between millions of protein sequences, enabling them to predict beneficial mutations, infer protein function, and perform zero-shot design of novel protein sequences with desired properties [4].
- Structure-Based Design Tools: ProteinMPNN uses deep learning to take an entire protein backbone structure as input and predict new sequences that fold into that structure. When combined with structure prediction tools like AlphaFold, it significantly increases design success rates [4].
- Functional Property Prediction: Specialized ML models now predict key protein properties such as thermostability (e.g., Prethermut, Stability Oracle) and solubility (e.g., DeepSol), allowing designers to filter and optimize constructs in silico before moving to physical assembly [4].
The "Build" Phase: Automation and Accelerated Assembly
In the Build phase, designed DNA constructs are synthesized, assembled into plasmids or other vectors, and introduced into a characterization system [4]. AI integration here focuses on automation and workflow optimization.
- Automated Biofoundries: Over 30 academic biofoundries worldwide (under the Global Biofoundry Alliance) use automation, high-throughput technologies, and computational design to accelerate the construction of biological systems [82] [25].
- LLM-Generated Code for Automation: ChatGPT-4 has been used to generate executable code for experimental design and liquid handling protocols without manual revision, dramatically reducing the time and specialized expertise required to automate build processes [82].
- Cell-Free Systems for Rapid Prototyping: Cell-free gene expression (CFPS) platforms leverage cellular transcription-translation machinery in lysates, allowing direct protein expression from DNA templates without time-intensive cloning. This enables high-throughput synthesis and testing [4] [82].
The "Test" Phase: High-Throughput Characterization and Data Generation
The Test phase involves experimentally measuring the performance of engineered biological constructs [4]. AI enables unprecedented scale and efficiency in testing.
- Ultra-High-Throughput Screening: Integration of cell-free systems with liquid handling robots and microfluidics allows screening of vast numbers of reactions. For example, DropAI used droplet microfluidics to screen over 100,000 picoliter-scale reactions [4].
- Megascale Data Generation: These platforms generate massive, high-quality datasets on protein stability and function, which are essential for training and refining ML models [4].
The "Learn" Phase: From Data to Predictive Insight
The Learn phase involves analyzing test data to inform the next design iteration [25]. This has traditionally been a bottleneck due to biological complexity.
- Active Learning (AL) Frameworks: AL is a machine learning approach where the model selectively queries the most informative data points for experimental testing, efficiently improving its predictive performance while minimizing required experiments [82].
- Closed-Loop Optimization: Fully automated DBTL pipelines incorporate AL to select diverse and informative experimental conditions, creating a continuous loop where learning directly drives the next design cycle without human intervention [82].
The figure below illustrates the AI-enhanced DBTL cycle and the proposed LDBT paradigm.
A Paradigm Shift: From DBTL to LDBT
Recent advances are prompting a fundamental rethinking of the traditional DBTL sequence. The increasing success of zero-shot predictions—where models can accurately design functional biological parts without additional training—suggests that "Learning" can now precede "Design" [4]. This new LDBT (Learn-Design-Build-Test) paradigm leverages pre-trained models on vast biological datasets to generate initial designs that are highly likely to work, potentially reducing or eliminating the need for multiple iterative cycles [4].
- Zero-Shot Prediction Success: Protein language models and structure-based tools can now design functional proteins without iterative experimental training data. For instance, researchers used a deep-learning model to survey over 500,000 antimicrobial peptide sequences and select optimal variants for testing, resulting in six promising designs with minimal experimental iteration [4].
- Foundation Models in Biology: Large-scale models trained on millions of protein sequences and structures encapsulate fundamental biological principles, allowing them to generate viable designs from the outset [4].
- Towards a "Design-Build-Work" Model: The ultimate goal is a workflow resembling established engineering disciplines like civil engineering, where designs based on first principles work correctly on the first implementation, drastically reducing development timelines [4].
Quantitative Impact of AI-Synthetic Biology Convergence
The integration of AI into synthetic biology workflows delivers measurable improvements in efficiency, yield, and success rates. The table below summarizes key quantitative findings from recent implementations.
Table 1: Quantitative Performance of AI-Enhanced Synthetic Biology Workflows
| AI Technology | Application Context | Key Performance Metrics | Result |
|---|---|---|---|
| Active Learning (Cluster Margin) [82] | Optimization of colicin M and E1 production in E. coli and HeLa CFPS systems | Yield improvement over baseline in 4 DBTL cycles | 2- to 9-fold increase in protein yield [82] |
| ProteinMPNN + AlphaFold [4] | Design of TEV protease variants | Increase in design success rate | Nearly 10-fold increase in design success rates [4] |
| Cell-Free Screening [4] | Ultra-high-throughput protein stability mapping | Scale of variants characterized | ∆G calculation for 776,000 protein variants in one dataset [4] |
| AI-Guided DBTL [82] | Fully automated pipeline implementation | Reduction in coding time for experimental design | ChatGPT-4 generated code without manual revisions [82] |
| DropAI Microfluidics [4] | High-throughput screening of reactions | Number of parallel reactions | Screening of >100,000 picoliter-scale reactions [4] |
Detailed Experimental Protocol: An AI-Driven DBTL Workflow for Protein Optimization
The following protocol details a specific implementation of a fully automated, AI-driven DBTL pipeline for optimizing protein production in cell-free systems, as demonstrated in a recent study [82].
This protocol describes a modular, fully automated DBTL workflow for optimizing cell-free protein synthesis (CFPS) in both bacterial (E. coli) and mammalian (HeLa) systems. The pipeline integrates experimental design, microplate layout generation, liquid handling execution, readout calibration, and data-driven candidate selection within the Galaxy platform, ensuring FAIR (Findable, Accessible, Interoperable, Reusable) compliance and accessibility for non-programmers [82].
Materials and Equipment
Table 2: Research Reagent Solutions and Essential Materials
| Item | Function/Description | Application in Protocol |
|---|---|---|
| Cell-Free System | Crude cell lysate or purified components containing transcription-translation machinery [4] | Core reaction environment for protein synthesis without living cells. |
| DNA Template | Synthesized DNA encoding the target protein(s). | Direct template for in vitro transcription and translation. |
| ChatGPT-4 | Large Language Model (LLM) for natural language processing and code generation [82] | Automates generation of Python scripts for experimental design and plate layout without manual coding. |
| Active Learning Model | Machine learning model with Cluster Margin (CM) sampling strategy [82] | Selects the most informative and diverse experimental conditions for each subsequent DBTL cycle. |
| Liquid Handler | Automated robotic liquid handling system. | Executes reagent dispensing, plate setup, and reaction assembly with high precision and throughput. |
| Microplate Reader | Instrument for measuring optical density, fluorescence, or luminescence. | Quantifies protein yield or activity in high-throughput format. |
Step-by-Step Methodology
Design Phase (Automated)
- Objective Definition: Define the goal (e.g., "optimize CFPS component concentrations to maximize yield of colicin M").
- LLM-Powered Script Generation: Use ChatGPT-4 with natural language prompts to generate Python scripts that create the experimental design and microplate layouts. These scripts define the initial set of conditions to test (e.g., varying concentrations of magnesium, nucleotides, and energy sources) [82].
- Output: A ready-to-execute experimental plan in machine-readable format.
Build Phase (Automated)
- Reagent Preparation: Prepare stock solutions of the CFPS components and DNA templates.
- Automated Liquid Handling: The liquid handler is programmed using the LLM-generated code to dispense reagents according to the designed plate layout, assembling the CFPS reactions in a microplate [82].
Test Phase (Automated)
- Incubation: Incubate the reaction plate under optimal conditions for protein synthesis (e.g., specific temperature and duration).
- Yield Quantification: Use the microplate reader to measure protein yield, for example via a colorimetric assay (e.g., Bradford) or a functional assay specific to the target protein (e.g., antimicrobial activity assay for colicins) [82].
- Data Collection: Results are automatically collected and formatted for analysis.
Learn Phase (Automated)
- Active Learning with Cluster Margin: The AL model, using the Cluster Margin strategy, analyzes all collected experimental data. It identifies the most "informative" conditions to test next—those the model is most uncertain about but that also represent diverse areas of the experimental space [82].
- Iteration: The outputs of this phase automatically become the new experimental designs for the next DBTL cycle, creating a closed-loop optimization system.
Expected Outcomes and Analysis
- In a typical application optimizing colicin production, this protocol achieved a 2- to 9-fold increase in protein yield after only four DBTL cycles [82].
- The Cluster Margin AL strategy significantly reduces the number of experiments required to find optimal conditions compared to traditional grid searches or one-variable-at-a-time approaches [82].
- The entire workflow, from design to learning, operates with minimal human intervention, demonstrating a robust framework for routine synthetic biology optimization tasks [82].
The figure below visualizes this automated, closed-loop workflow.
The Scientist's Toolkit: Key Computational and Biological Platforms
The modern AI-synthetic biology workflow relies on a suite of specialized computational tools and biological platforms. The table below catalogs the essential components of the integrated research toolkit.
Table 3: Essential Tools for AI-Driven Synthetic Biology
| Tool Category | Representative Examples | Primary Function |
|---|---|---|
| Protein Language Models | ESM (Evolutionary Scale Modeling), ProGen [4] | Predict protein structure and function from sequence; enable zero-shot design of novel proteins. |
| Structure-Based Design Tools | ProteinMPNN, MutCompute [4] | Design protein sequences that fold into a specific backbone structure or optimize local residue environments. |
| Stability & Solubility Predictors | Prethermut, Stability Oracle, DeepSol [4] | Predict the effects of mutations on protein thermodynamic stability (ΔΔG) and solubility. |
| Active Learning Frameworks | Cluster Margin Sampling [82] | Intelligently select the most informative experiments to perform, minimizing the number of cycles needed for optimization. |
| Cell-Free Expression Systems | E. coli lysates, HeLa lysates [4] [82] | Enable rapid, high-throughput protein synthesis and testing without the constraints of living cells. |
| Automation & Integration Platforms | Galaxy Platform, Biofoundries [82] [25] | Provide integrated, FAIR-compliant environments for executing and reproducing automated DBTL workflows. |
| Large Language Models (LLMs) | ChatGPT-4 [82] | Generate executable code for experimental design and automation from natural language prompts, democratizing access. |
The convergence of AI and synthetic biology is ushering in a new era of precision biological design. By deeply integrating machine learning, large language models, and automated experimental platforms into the DBTL cycle, researchers are overcoming traditional bottlenecks and accelerating the pace of bioinnovation. The emergence of paradigms like LDBT, powered by zero-shot predictive models, points toward a future where biological engineering is more predictable, efficient, and accessible. As these tools continue to evolve, they promise to unlock transformative applications across medicine, manufacturing, and environmental sustainability, fundamentally changing our approach to designing and programming biological systems.
The integration of in silico models into the synthetic biology and drug development pipeline represents a paradigm shift in how researchers approach biological design. Framed within the classic Design-Build-Test-Learn (DBTL) cycle, advanced computational techniques are accelerating the path from conceptual design to clinical application. This technical guide explores the validation frameworks, computational architectures, and experimental methodologies that enable researchers to bridge the gap between computational predictions and clinical success, with particular emphasis on how machine learning is transforming traditional workflows into more efficient Learn-Design-Build-Test (LDBT) approaches [4]. We examine how large-scale perturbation models, cell-free testing systems, and computational validation protocols are creating a new foundation for predictive biological engineering that reduces development timelines while increasing success rates.
The Design-Build-Test-Learn cycle has served as the fundamental engineering framework for synthetic biology, providing a systematic, iterative approach to biological system design [1]. In traditional implementation, researchers first Design biological components based on existing knowledge, Build DNA constructs and introduce them into biological systems, Test the resulting systems experimentally, and finally Learn from the results to inform the next design cycle [4]. While effective, this approach often requires multiple iterations to achieve desired functionality, creating bottlenecks in development timelines, particularly in the Build and Test phases.
Recent advances in machine learning and computational modeling are fundamentally transforming this paradigm. The emergence of the LDBT framework, where Learning precedes Design, leverages large datasets and pre-trained models to generate more effective initial designs [4]. This approach utilizes zero-shot predictions from protein language models and structural bioinformatics to create functional designs without requiring multiple experimental iterations, potentially reducing the number of DBTL cycles needed to achieve target functionality.
Table 1: Evolution of the DBTL Cycle in Synthetic Biology
| Framework | Sequence | Key Features | Advantages |
|---|---|---|---|
| Traditional DBTL | Design → Build → Test → Learn | Domain knowledge-driven design, experimental iteration | Systematic approach, proven effectiveness |
| Machine Learning-Enhanced DBTL | Design → Build → Test → Learn | ML-guided design, predictive modeling | Improved initial designs, reduced iterations |
| LDBT | Learn → Design → Build → Test | Zero-shot prediction, foundational models | Potential for single-cycle success, reduced experimental burden |
Computational Foundations: Large-Scale Perturbation Models and AI-Assisted Discovery
Large Perturbation Models (LPMs) for Biological Discovery
The Large Perturbation Model (LPM) represents a breakthrough in computational biology for integrating heterogeneous perturbation data. As described in recent literature, LPMs are deep-learning models designed to integrate multiple, heterogeneous perturbation experiments by representing perturbation (P), readout (R), and context (C) as disentangled dimensions [83]. This PRC-conditioned architecture enables learning from diverse experimental data across different readouts (transcriptomics, viability), perturbations (CRISPR, chemical), and biological contexts (single-cell, bulk) without loss of generality.
A key innovation of LPM architecture is its decoder-only design, which avoids the limitations of encoder-based models that struggle with low signal-to-noise ratios in high-throughput screens [83]. By explicitly conditioning on representations of experimental context, LPMs learn perturbation-response rules disentangled from the specific context in which readouts were observed. This approach has demonstrated state-of-the-art predictive performance in forecasting post-perturbation transcriptomes of unseen experiments, outperforming established methods including CPA, GEARS, and foundation models like Geneformer and scGPT [83].
Mapping Compound-CRISPR Shared Perturbation Space
LPMs enable the integration of genetic and pharmacological perturbations within a unified latent space, facilitating the study of drug-target interactions. When visualizing the perturbation embedding space learned by LPMs, pharmacological inhibitors consistently cluster in close proximity to genetic CRISPR interventions targeting the same genes [83]. For example, genetic perturbations targeting MTOR cluster closely with compounds inhibiting MTOR, similarly, genetic perturbations targeting related pathway genes (PSMB1 and PSMB2, HDAC2 and HDAC3) show tight clustering.
This unified representation enables important discoveries, such as identifying off-target activities and novel mechanisms. For instance, LPMs autonomously positioned pravastatin closer to nonsteroidal anti-inflammatory drugs targeting PTGS1 rather than with other statins, suggesting an anti-inflammatory mechanism that aligns with clinical observations of pravastatin's pleiotropic effects [83]. This demonstrates how in silico models can generate clinically relevant hypotheses about drug mechanisms.
AI-Assisted In Silico Validation
In silico validation has become a critical component of early drug development, using computational approaches to predict efficacy, safety, and mechanisms of action before experimental testing [84]. AI-assisted drug discovery leverages large datasets from biological, chemical, and clinical sources to train models capable of predicting therapeutic efficacy, toxicity profiles, and off-target interactions.
These approaches combine AI algorithms with molecular modeling, docking simulations, and machine learning to simulate drug-target interactions, allowing promising candidates to be prioritized for experimental testing [84]. The integration of in silico validation reduces both time and cost associated with traditional drug development approaches while increasing the accuracy and reliability of outcomes. However, challenges remain in model generalization and the need for extensive clinical validation to ensure translational success.
Table 2: Computational Models for Biological Discovery
| Model Type | Key Applications | Architecture | Performance Advantages |
|---|---|---|---|
| Large Perturbation Models (LPM) | Perturbation outcome prediction, mechanism identification, gene interaction modeling | PRC-disentangled dimensions, decoder-only | Outperforms CPA, GEARS, Geneformer, scGPT in predicting unseen perturbation effects [83] |
| Protein Language Models (ESM, ProGen) | Protein design, mutation effect prediction, function inference | Transformer-based, trained on evolutionary sequences | Zero-shot prediction of beneficial mutations, antibody sequences [4] |
| Structure-Based Models (ProteinMPNN, MutCompute) | Protein sequence design, stability optimization | Neural networks trained on protein structures | 10-fold increase in design success rates when combined with AlphaFold [4] |
| Stability Prediction Models (Prethermut, Stability Oracle) | Thermodynamic stability prediction of mutants | Machine learning trained on stability data | Predicts ΔΔG of proteins, identifies stabilizing mutations [4] |
Experimental Validation: From Silicon to Reality
Cell-Free Platforms for Rapid Testing
Cell-free gene expression systems have emerged as a powerful platform for accelerating the Test phase of synthetic biology cycles. These systems leverage protein biosynthesis machinery from cell lysates or purified components to activate in vitro transcription and translation without intermediate cloning steps [4]. The advantages of cell-free systems include:
- Rapid expression (>1 g/L protein in <4 hours)
- Scalability from picoliter to kiloliter scales
- Production of toxic products that would be untenable in live cells
- Compatibility with automation and high-throughput screening
Recent advances have demonstrated how cell-free systems can be paired with droplet microfluidics and multi-channel fluorescent imaging to screen upwards of 100,000 picoliter-scale reactions in parallel [4]. This massive throughput provides the large-scale datasets necessary for training and validating machine learning models, creating a virtuous cycle of improvement for in silico predictions.
Ultra-High-Throughput Protein Stability Mapping
The integration of cell-free protein synthesis with cDNA display has enabled unprecedented throughput in stability mapping, allowing ΔG calculations for 776,000 protein variants in a single experiment [4]. This massive dataset has been instrumental for benchmarking zero-shot predictors and improving model predictability. Similar approaches have been applied to enzyme engineering campaigns, where linear supervised models trained on over 10,000 reactions from iterative site saturation mutagenesis have accelerated identification of enzyme candidates with favorable properties [4].
For antimicrobial peptide discovery, researchers have paired deep-learning sequence generation with cell-free expression to computationally survey over 500,000 potential variants, select 500 optimal candidates for experimental validation, and identify 6 promising designs [4]. This demonstrates the power of combining in silico screening with rapid experimental validation.
Methodologies and Protocols
LPM Training and Implementation Protocol
Large Perturbation Models require specific training methodologies to achieve state-of-the-art performance:
Data Integration and Preprocessing
- Collect heterogeneous perturbation data from diverse sources (LINCS, etc.)
- Standardize perturbation, readout, and context representations
- Implement quality control metrics to ensure data integrity
Model Architecture Specification
- Implement PRC-disentangled architecture with separate conditioning variables
- Configure decoder-only transformer architecture with attention mechanisms
- Set optimization parameters (learning rate, batch size, regularization)
Training Procedure
- Train on pooled perturbation experiments with diverse configurations
- Use transfer learning from related biological domains when available
- Implement early stopping based on validation set performance
Validation and Benchmarking
- Compare against established baselines (CPA, GEARS, Geneformer, scGPT)
- Evaluate on held-out experiments with unseen perturbations
- Assess performance across multiple biological discovery tasks
Cell-Free Testing Protocol for Validation
Rapid experimental validation of in silico predictions using cell-free systems:
DNA Template Preparation
- Design and synthesize DNA templates encoding predicted variants
- Use high-throughput DNA assembly methods for library construction
- Verify sequence fidelity through next-generation sequencing
Cell-Free Reaction Setup
- Prepare cell-free transcription-translation machinery
- Dispense reactions using liquid handling robots or microfluidics
- Implement controls for background subtraction and normalization
Functional Assay Implementation
- Configure colorimetric, fluorescent, or other reporter assays
- Set up high-throughput screening platforms
- Establish quantitative readouts for function and stability
Data Integration and Model Refinement
- Correlate experimental results with computational predictions
- Identify patterns of prediction success and failure
- Refine models based on experimental feedback
Visualization of Workflows and Pathways
LDBT Cycle for Predictive Design
LPM Architecture for Perturbation Modeling
The Scientist's Toolkit: Essential Research Reagents and Solutions
Table 3: Research Reagent Solutions for In Silico Validation
| Reagent/Solution | Function | Application Context |
|---|---|---|
| Cell-Free Transcription-Translation Systems | Provides protein biosynthesis machinery for rapid expression without living cells | High-throughput testing of protein variants, pathway prototyping [4] |
| DNA Assembly Kits | Enables modular construction of genetic circuits and variant libraries | Build phase for synthetic biology constructs, library preparation [1] |
| Fluorescent Reporters | Quantifies gene expression and protein production through measurable signals | Test phase for functional assessment, high-throughput screening [4] |
| Next-Generation Sequencing Kits | Verifies DNA sequence fidelity and analyzes genetic composition | Quality control for Build phase, validation of designed sequences [1] |
| Microfluidic Devices | Enables picoliter-scale reactions for ultra-high-throughput screening | Test phase automation, massive parallelization of assays [4] |
| Stability Assay Reagents | Measures protein thermodynamic stability and folding efficiency | Functional validation of computationally designed proteins [4] |
The integration of advanced in silico models with rapid experimental validation represents a fundamental shift in synthetic biology and drug discovery. The transformation from traditional DBTL cycles to LDBT approaches demonstrates how machine learning and computational forecasting are reducing dependence on iterative experimental optimization. Large Perturbation Models, protein language models, and structure-based prediction tools are creating new opportunities for accurate first-pass design of biological systems.
As these technologies mature, the field moves closer to a Design-Build-Work paradigm where biological systems can be engineered with reliability approaching that of traditional engineering disciplines. However, challenges remain in model generalization, data standardization, and clinical translation. Future developments will likely focus on integrating multi-omic data, improving model interpretability, and establishing robust validation frameworks that bridge the gap between computational prediction and clinical application. Through continued refinement of both in silico and experimental methods, researchers are building a foundation for more efficient, predictive, and successful biological design.
Synthetic biology employs the Design-Build-Test-Learn (DBTL) cycle as its core development pipeline for engineering biological systems [25]. While advancements in DNA sequencing and synthesis have dramatically accelerated the 'Build' and 'Test' phases, the 'Learn' stage has become a critical bottleneck [25]. The inherent complexity, non-linearity, and high-dimensional interactions within biological systems generate vast amounts of data that are difficult to decipher using traditional analytical methods [18]. This often forces the engineering process away from rational design and into a regime of ad-hoc tinkering [18]. Explainable Artificial Intelligence (XAI) is emerging as a transformative solution to this challenge, offering both predictive power and interpretability. When combined with standardized data generation, XAI is poised to debottleneck the DBTL cycle, enabling a shift from iterative, empirical testing to predictive biological design [25] [18].
Quantitative Landscape of XAI in Drug and Biological Research
The adoption of XAI in life sciences is growing rapidly. A 2025 bibliometric analysis provides a snapshot of this trend, highlighting the application of XAI in drug research and revealing key geographical leaders and research foci [85].
Table 1: Top Countries in XAI for Drug Research (2002-2024)
| Rank | Country | Total Publications (TP) | Total Citations (TC) | TC/TP Ratio | Publication Start Year |
|---|---|---|---|---|---|
| 1 | China | 212 | 2949 | 13.91 | 2013 |
| 2 | USA | 145 | 2920 | 20.14 | 2006 |
| 3 | Germany | 48 | 1491 | 31.06 | 2002 |
| 4 | UK | 42 | 680 | 16.19 | 2007 |
| 5 | South Korea | 31 | 334 | 10.77 | 2009 |
| 6 | India | 27 | 219 | 8.11 | 2017 |
| 7 | Japan | 24 | 295 | 12.29 | 2018 |
| 8 | Canada | 20 | 291 | 14.55 | 2016 |
| 9 | Switzerland | 19 | 645 | 33.95 | 2006 |
| 10 | Thailand | 19 | 508 | 26.74 | 2015 |
The data shows that while China and the USA lead in publication volume, Switzerland, Germany, and Thailand produce research with the highest academic impact, as measured by citations per paper [85]. This reflects distinct and mature research niches: Switzerland excels in molecular property prediction and drug safety [85]; Germany has a long-standing focus on multi-target compounds and drug response prediction [85]; and Thailand shows rapid development in biologics for infections and cancer [85].
Table 2: Key XAI Techniques and Their Applications in Biological Research
| XAI Technique | Primary Function | Application Example in Drug Discovery |
|---|---|---|
| SHAP (Shapley Additive Explanations) [85] [86] | Quantifies the contribution of each input feature to a model's prediction for a specific instance. | Explaining which molecular descriptors most influenced a toxicity prediction for a novel compound. |
| LIME (Local Interpretable Model-agnostic Explanations) [86] | Approximates a complex "black box" model locally with an interpretable model to explain individual predictions. | Highlighting the chemical substructures in a molecule that led a model to classify it as bioactive. |
| Similarity Maps [86] | Visualizes the similarity of a molecule to known active compounds based on its fingerprint. | Assessing the novelty of a de novo-designed chemical entity and its relationship to existing chemical space. |
| Counterfactual Explanations [86] | Generates examples of minimal changes to the input that would alter the model's prediction. | Proposing specific, minimal structural changes to a drug candidate to reduce its predicted hepatotoxicity. |
Technical Protocols for Integrating XAI into the DBTL Cycle
Integrating XAI effectively requires structured methodologies. The following protocols outline how to incorporate XAI at the 'Learn' stage to accelerate biological design.
Protocol for Model Interpretation using SHAP
Objective: To interpret a machine learning model that predicts protein expression levels from genetic part sequences (e.g., promoters, RBS) in a microbial host [25] [18].
Materials:
- Trained Predictive Model: A random forest or neural network model trained on sequence-expression data.
- Dataset: A curated test set of genetic sequences and corresponding measured expression levels.
- SHAP Library: The Python
shaplibrary (or equivalent in R).
Method:
- Compute SHAP Values: Using the trained model and the test set, calculate SHAP values for each sequence feature in the test set. For sequence data, features can be one-hot-encoded nucleotides or k-mers.
- Global Interpretation: Generate a summary plot that ranks features by their mean absolute SHAP value. This identifies the sequence motifs or positions that have the largest average impact on expression levels globally across the dataset [86].
- Local Interpretation: For a specific genetic construct of interest, create a force plot or waterfall plot. This visualizes how each feature of that specific sequence contributed to shifting the model's prediction from the base value to the final predicted expression level [86].
- Dependence Analysis: Select a top-contributing feature from the global analysis and create a SHAP dependence plot. This scatter plot shows the effect of varying that single feature on the model's output, which can reveal interaction effects with other features [86].
Protocol for Knowledge Extraction via Counterfactual Generation
Objective: To understand the sequence-function relationship of an enzyme and guide rational engineering for improved activity [1].
Materials:
- Trained Classifier: A model that predicts "high" or "low" catalytic activity from protein sequence or structure features.
- Generative Model: A variational autoencoder (VAE) or generative adversarial network (GAN) capable of generating realistic protein sequence variants.
Method:
- Identify a Target Instance: Select a protein variant from your dataset that the model correctly classified as having "low" activity.
- Generate Counterfactuals: Use the generative model to create a set of novel protein sequences that are minimally different from the target sequence but are predicted by the classifier to have "high" activity. This often involves optimizing in the model's latent space [18].
- Analyze Sequence Differences: Align the counterfactual sequences with the original target sequence. Identify the conserved and mutated residues.
- Formulate Hypotheses: The pattern of mutations required to flip the prediction constitutes a testable hypothesis. It suggests that those specific residues are critical for the desired function. This hypothesis can then be validated experimentally in the next 'Build-Test' cycle [86] [18].
The following diagram illustrates the synergistic workflow of the augmented DBTL cycle, highlighting how XAI directly informs subsequent design iterations.
The Scientist's Toolkit: Essential Research Reagents and Solutions
The implementation of the AI-augmented DBTL cycle relies on a suite of wet-lab and dry-lab tools.
Table 3: Key Research Reagent Solutions for an AI-Driven Workflow
| Item / Solution | Function in the Workflow |
|---|---|
| Biofoundry Services [25] [13] | Automated facilities for high-throughput DNA assembly, genome editing, and strain cultivation, essential for generating the large, standardized datasets required for ML. |
| Gibson Assembly / DNA Synthesis Kits [1] [25] | Molecular tools for the seamless and rapid construction of genetic variants as designed by the AI. |
| CRISPR-Cas9 Genome Editing Systems [18] | Enables precise, targeted modifications to host chassis genomes to implement designed genetic circuits or pathways. |
| Next-Generation Sequencing (NGS) [1] [18] | Provides high-throughput genotypic verification of built constructs and can be used for transcriptomic analysis in the 'Test' phase. |
| Mass Spectrometry [13] | Critical for proteomic and metabolomic profiling in the 'Test' phase, quantifying the output of engineered systems (e.g., metabolite titers, protein expression). |
| SHAP (shap Python library) [85] [86] | The primary XAI library for interpreting ML model outputs and generating feature importance scores. |
| LIME (lime Python library) [86] | A model-agnostic library for creating local, interpretable explanations of complex model predictions. |
The 'Learn' bottleneck has long constrained the pace of innovation in synthetic biology and drug discovery. The integration of Explainable AI directly addresses this challenge by transforming complex, high-dimensional data into actionable biological insights. This moves the field beyond black-box predictions towards a deeper, causal understanding of biological design principles [86] [25]. The synergistic combination of standardized data generation from automated biofoundries and the interpretative power of XAI creates a virtuous cycle of learning, dramatically accelerating the DBTL cycle [18]. As these technologies mature and become more accessible, they will underpin a new paradigm of predictive, precision biological engineering, fundamentally reshaping our approach to developing therapeutics, sustainable materials, and bio-based solutions to global challenges [87] [18].
Conclusion
The DBTL cycle has firmly established itself as the foundational paradigm for rational biological design, proving indispensable in accelerating drug development and biomanufacturing. The integration of AI and machine learning is fundamentally reshaping this cycle, transforming it from an iterative, empirical process into a more predictive and efficient engineering discipline. This convergence is key to unlocking high-precision biological design, from engineering robust cell factories for therapeutic protein production to developing sophisticated diagnostic and delivery systems like engineered vesicles and biosensing tattoos. For biomedical and clinical research, the future lies in fully automated, AI-driven DBTL pipelines that can rapidly generate and validate novel therapeutic candidates, personalize treatments, and ultimately democratize the ability to engineer biology. Success in this new era will depend on continued advancements in computational tools, the establishment of robust data standards, and the development of proactive governance frameworks to ensure responsible innovation.
References
- 1. DBTL Approach in Synthetic Biology [https://www.moleculardevices.com/applications/synthetic-biology/dbtl-approach]
- 2. Engineering | ESSENTIAL KOREA - iGEM 2025 [https://2025.igem.wiki/essential-korea/engineering]
- 3. The knowledge driven DBTL cycle provides mechanistic ... [https://microbialcellfactories.biomedcentral.com/articles/10.1186/s12934-025-02729-6]
- 4. combining machine learning and rapid cell-free testing [https://www.nature.com/articles/s41467-025-65281-2]
- 5. DBTL bioengineering cycle for part characterization and ... [https://www.sciencedirect.com/science/article/pii/S2405896324017488]
- 6. A machine learning Automated Recommendation Tool for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7519645/]
- 7. The automated Galaxy-SynBioCAD pipeline for synthetic ... [https://www.nature.com/articles/s41467-022-32661-x]
- 8. Abstraction hierarchy to define biofoundry workflows and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12214803/]
- 9. A generalized platform for artificial intelligence-powered ... [https://www.nature.com/articles/s41467-025-61209-y]
- 10. - Design - Build - Test (DBTL) | EBRC Learn Roadmap Research [https://roadmap.ebrc.org/dbtl/]
- 11. Life: The Transformative Potential of Designing ... Synthetic Biology [https://brite.ikeinstitute.org/brite_innovation_review_issue_35_may_2025/synthetic_biology]
- 12. An automated Design-Build-Test-Learn pipeline for ... [https://www.nature.com/articles/s42003-018-0076-9]
- 13. Automating the design-build-test-learn cycle towards next- ... [https://www.sciencedirect.com/science/article/pii/S187167842300002X]
- 14. Evolutionary tinkering vs. rational engineering in the times of ... [https://lsspjournal.biomedcentral.com/articles/10.1186/s40504-018-0086-x]
- 15. Rational design: best route for synthetic biology? - Wyss Institute [https://wyss.harvard.edu/news/rational-design-best-route-for-synthetic-biology/]
- 16. Engineering is evolution: a perspective on design processes ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11059173/]
- 17. 14 Comparing quantitative data between individuals [https://bookdown.org/pkaldunn/SRM-Textbook/BetweenQuantData.html]
- 18. Digital to Biological Translation: How the Algorithmic Data ... [https://www.mdpi.com/2674-0583/3/4/17]
- 19. a solution to involution of design–build–test–learn cycle [https://www.sciencedirect.com/science/article/abs/pii/S0958166922000453]
- 20. Engineering development: From the repressilator and ... [https://www.sciencedirect.com/science/article/pii/S0012160625001800]
- 21. Recent advances in the Design-Build-Test-Learn (DBTL ... [https://pubmed.ncbi.nlm.nih.gov/40195836/]
- 22. Simulated Design–Build–Test–Learn Cycles for Consistent ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10510747/]
- 23. Quantifying microbial interactions based on compositional ... [https://pubmed.ncbi.nlm.nih.gov/41202104/]
- 24. Design-Build-Test-Learn: Impact of AI on the Synthetic Biology ... [https://www.ncbi.nlm.nih.gov/books/NBK614601/]
- 25. Learning the way toward high-precision biological design [https://pmc.ncbi.nlm.nih.gov/articles/PMC10231942/]
- 26. Synthetic Genomes: Rewriting the Blueprint of Life [https://www.the-scientist.com/synthetic-genomes-rewriting-the-blueprint-of-life-72010]
- 27. Automated high-throughput DNA synthesis and assembly [https://www.sciencedirect.com/science/article/pii/S2405844024029980]
- 28. Synthetic DNA Synthesis and Assembly - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC5204324/]
- 29. Recent advances in DNA assembly technologies - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4257898/]
- 30. High Throughput Screening Assays for Drug Discovery [https://bellbrooklabs.com/high-throughput-screening-assays-drug-discovery/?srsltid=AfmBOoq9cPQFdV8T1mFzBkDUH_wurAY6NELuT6Rb53ZdE_ALdqaxUtwH]
- 31. Gene-based calibration of high-throughput functional ... [https://pubmed.ncbi.nlm.nih.gov/40654914/]
- 32. 2025 Trends: Multiomics [https://www.genengnews.com/topics/omics/2025-trends-multiomics/]
- 33. Data Analysis Methods: 7 Essential Techniques for 2025 [https://atlan.com/data-analysis-methods/]
- 34. Presenting Quantitative Data Graphically [https://courses.lumenlearning.com/waymakermath4libarts/chapter/presenting-quantitative-data-graphically/]
- 35. Presentation of Quantitative Data [https://ihatepsm.com/blog/presentation-quantitative-data]
- 36. Text has enhanced contrast | ACT Rule | WAI [https://www.w3.org/WAI/standards-guidelines/act/rules/09o5cg/]
- 37. Text has enhanced contrast - Rules - Siteimprove [https://alfa.siteimprove.com/rules/sia-r66]
- 38. Engineering bacteria to modulate host metabolism - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10909415/]
- 39. Developing a new class of engineered live bacterial ... [https://www.nature.com/articles/s41467-020-15508-1]
- 40. Combining enzyme and metabolic engineering for ... [https://www.sciencedirect.com/science/article/pii/S0958166924000466]
- 41. Synthetic Biology Market worth $31.52 billion in 2029 [https://www.marketsandmarkets.com/PressReleases/synthetic-biology.asp]
- 42. Global Synthetic Biology Market Size and Industry Analysis [https://www.bccresearch.com/market-research/biotechnology/synthetic-biology-global-markets.html?srsltid=AfmBOoq1lifdBJbY-GGL7bV-wUuPRIQUFIcEcavZ3NTIbpQ4FlLyDq-g]
- 43. The convergence of AI and synthetic biology: the looming ... [https://www.nature.com/articles/s44385-025-00021-1]
- 44. Natural products, synthetic biology and artificial ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11761351/]
- 45. Scientists will eliminate bottlenecks to breakthroughs with a ... [https://bioengineering.ucsb.edu/news/all/2023/scientists-will-eliminate-bottlenecks-breakthroughs-newly-acquired-synthetic-biology]
- 46. Breaking the Bottleneck in AI-Driven Protein Engineering [https://www.ailurus.bio/post/breaking-the-bottleneck-in-ai-driven-protein-engineering]
- 47. SynBioBeta 2025: Key Industry Trends and Challenges in… [https://www.biocatalysts.com/media-resources/trends-synthetic-biology]
- 48. Breaking the Bottleneck: On-Demand Biology in the Age of AI [https://www.genengnews.com/topics/bioprocessing/breaking-the-bottleneck-on-demand-biology-in-the-age-of-ai/]
- 49. Building a biofoundry - PMC - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC7998708/]
- 50. Frontiers in biofoundry: opportunities and challenges [https://www.frontiersin.org/journals/synthetic-biology/articles/10.3389/fsybi.2025.1630026/full]
- 51. Architectures of emerging biofoundry platforms for synthetic ... [https://www.sciencedirect.com/science/article/abs/pii/S0958166925001235]
- 52. Engineering biological systems using automated biofoundries [https://pmc.ncbi.nlm.nih.gov/articles/PMC5544601/]
- 53. Biofoundries: a revolution for research and global economy [https://bioconvs.org/biofoundries-a-revolution-for-research-and-global-economy/]
- 54. Abstraction hierarchy to define biofoundry workflows and ... [https://www.nature.com/articles/s41467-025-61263-6]
- 55. How Biofoundries are Accelerating Life-Enhancing ... [https://www.synbiobeta.com/read/how-biofoundries-are-accelerating-life-enhancing-innovations-in-biomanufacturing]
- 56. The Life Scientist's Guide to Building a Successful Biofoundry [https://www.scisure.com/blog/guide-to-building-a-successful-biofoundry]
- 57. Machine learning in biological research: key algorithms ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12574268/]
- 58. Machine Learning in Computational Biology: Bridging Data ... [https://medium.com/techloop/machine-learning-in-computational-biology-bridging-data-and-discovery-for-next-generation-416e07b3e8c6]
- 59. unravelling brain disorders through data integration and ... [https://www.nature.com/articles/s41380-025-03330-4]
- 60. The future of AI regulation in drug development: a comparative ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12598624/]
- 61. Artificial Intelligence, Machine Learning and Omic Data ... [https://www.sciencedirect.com/science/article/pii/S1063458425011951]
- 62. Artificial Intelligence for Drug Development [https://www.fda.gov/about-fda/center-drug-evaluation-and-research-cder/artificial-intelligence-drug-development]
- 63. How AI will reshape pharma in 2025 [https://www.drugtargetreview.com/article/154981/how-ai-will-reshape-pharma-by-2025/]
- 64. AI and Synthetic Biology: The New Frontier of Promise and ... [https://bisi.org.uk/reports/ai-and-synthetic-biology-the-new-frontier-of-promise-and-power]
- 65. History and Scope of AI in Biodesign [https://www.biodesign.academy/p/history-and-scope-of-ai-in-biodesign]
- 66. AI & Machine Learning Driving Drug Discovery in 2025 [https://biopharmainfo.net/drug-discovery-ai-2025]
- 67. AI vs Traditional Methods in Bioinformatics Data Analysis [https://riveraxe.com/ai-vs-traditional-methods-in-bioinformatics-data-analysis/]
- 68. Closing the Biosecurity Gap in Synthetic Biology [https://globalbiodefense.com/2025/10/07/closing-the-biosecurity-gap-in-synthetic-biology/]
- 69. While AI Could Be the Game Changer in Predicting Health ... [https://www.medschool.umaryland.edu/news/2025/while-ai-could-be-the-game-changer-in-predicting-health-outcomes-it-should-not-be-the-only-method.html]
- 70. AI in Drug Development: Use-cases and Trends [https://www.scilife.io/blog/ai-in-drug-development]
- 71. AI in Pharma and Biotech: Market Trends 2025 and Beyond [https://www.coherentsolutions.com/insights/artificial-intelligence-in-pharmaceuticals-and-biotechnology-current-trends-and-innovations]
- 72. AI In Action: Redefining Drug Discovery and Development [https://pmc.ncbi.nlm.nih.gov/articles/PMC11800368/]
- 73. Why 90% of clinical drug development fails and how to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9293739/]
- 74. DBTL bioengineering cycle for part characterization and ... [https://www.sciencedirect.com/science/article/abs/pii/S2405896324017488]
- 75. Biopharma RD Faces Productivity And Attrition Challenges ... [https://www.clinicalleader.com/doc/biopharma-r-d-faces-productivity-and-attrition-challenges-in-2025-0001]
- 76. Biopharma's Patent Cliff Puts Costs Front and Center [https://www.bcg.com/publications/2025/patent-cliff-threatens-biopharmaceutical-revenue]
- 77. DNA Manufacturing Market Size to Hit USD 20.28 Billion by [https://www.globenewswire.com/news-release/2025/11/07/3183624/0/en/DNA-Manufacturing-Market-Size-to-Hit-USD-20-28-Billion-by-2032-Fueled-by-Gene-Therapy-and-Synthetic-Biology-Growth-SNS-Insider.html]
- 78. Research report: The state of synthetic research in 2025 [https://christophersilvestri.com/research-reports/state-of-synthetic-research-in-2025/]
- 79. 3 Promising Genomics & Synthetic Biology Stocks in ... [https://finance.yahoo.com/news/3-promising-genomics-synthetic-biology-153500286.html]
- 80. iGEM Jamboree: leap into the synthetic biology future [https://www.labiotech.eu/trends-news/igem-2025-synthetic-biology-highlights/]
- 81. AI Compute Demand in Biotech: 2025 Report & Statistics [https://intuitionlabs.ai/articles/ai-compute-demand-biotech]
- 82. An AI-driven workflow for the accelerated optimization of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12529354/]
- 83. In silico biological discovery with large perturbation models [https://www.nature.com/articles/s43588-025-00870-1]
- 84. In Silico Validation of AI-Assisted Drugs in Healthcare [https://pubmed.ncbi.nlm.nih.gov/40553347/]
- 85. Explainable Artificial Intelligence in the Field of Drug Research [https://pmc.ncbi.nlm.nih.gov/articles/PMC12129466/]
- 86. Drug discovery with explainable artificial intelligence [https://www.nature.com/articles/s42256-020-00236-4]
- 87. Generative AI agents are transforming biology research [https://pmc.ncbi.nlm.nih.gov/articles/PMC11583719/]